Institute of Oceanology, Chinese Academy of Sciences
Article Information
- GAO Dahai, ZHANG Qingchun, SUN Zhongmin
- Development of chloroplast marker for identification of Ulva species
- Journal of Oceanology and Limnology, 40(6): 2364-2371
- http://dx.doi.org/10.1007/s00343-022-2154-6
Article History
- Received Apr. 6, 2022
- accepted in principle Jun. 8, 2022
- accepted for publication Jul. 26, 2022
2 Shanghai Engineering Research Center of Aquaculture, Shanghai Ocean University, Shanghai 201306, China;
3 Department of Marine Organism Taxonomy and Phylogeny, Institute of Oceanology, Chinese Academy of Sciences(IOCAS), Qingdao 266071, China;
4 Key Laboratory of Marine Ecology and Environmental Sciences, Institute of Oceanology, Chinese Academy of Sciences, Qingdao 266071, China
As a type of harmful algal boom, green tide caused by Ulva spp. is a source of marine pollution (Blomster et al., 2002). Recently, large-scale green tides occur frequently in coastal areas worldwide (Smetacek and Zingone, 2013). In China, Ulva green tides occurred in the Bohai Sea, Yellow Sea, and South China Sea in last decade, which damaged local marine environments and ecological systems (Zhang et al., 2019; Xie et al., 2020b). Specifically, the world's largest green tide in the Yellow Sea have lasted for 15 consecutive years since 2007, whose causal species were identified as U. prolifera (Pang et al., 2010; Zhang et al., 2011, 2018, 2019; Zhao et al., 2012; Qi et al., 2016). Based on laboratory culturing analysis, it showed that the multiple life cycle modes were exhibited for U. prolifera, involving sexual reproduction with biflagellate gametes and quadriflagellate meiospores, and asexual reproduction from biflagellate spores, quadriflagellate spores, and unfertilized gametes (Liu et al., 2015; Zhang et al., 2017). Although U. prolifera can be recognized from its related taxa in adult stage, it is hard to morphologically discriminate each another in the young stage even for the experts. Phylogenetically, U. prolifera, U. procera, and U. linza could not be distinct and formed a clade called linza-procera-prolifera (LPP) clade based on solo ITS or rbcL markers (Shimada et al., 2008, 2016). And the hybridization studies indicated that U. prolifera and U. linza was the closely related Ulva species with complex speciation history, as the marine strains of U. prolifera were complete reproductive isolated with U. linza exhibiting by gamete incompatibility (Hiraoka et al., 2011), whereas some brackish strains of U. prolifera showed partial gamete compatibility with U. linza (Xie et al., 2020a).
To solve the relationships among closely related species in Ulva, several molecular approaches were reported. Besides the combination of ITS or rbcL markers, Shimada et al. (2008) firstly introduced the 5S rDNA spacer sequences to analyze the genetic diversity within LPP complex, and this marker was widely used for studying origin of floating U. prolifera under intra-specific level (Zhao et al., 2019). However, the using of 5S rDNA marker has some deficiencies as the PCR products of 5S rDNA marker were not specific as several bands were observed during electrophoretic analysis (Liu et al., 2022). On the other hand, as 5S rDNA marker was from nuclear genome, it will confuse the analysis if specimens were heterozygous. In another study, a PCR-based marker of sequence characterized amplified region (SCAR) was designed for identifying the U. prolifera populations (Zhao et al., 2015), but the application of SCAR marker is limited as only floating U. prolifera could be specifically amplified. In addition, the ribosomal large subunit (LSU) and intergenic spacer (IGS) sequences in U. prolifera were investigated, but their ability to delimitate closely related species remains uncertain (Shen et al., 2019). Therefore, specific markers should be exploited to identify the closely related species in a stable manner.
In land plants, the molecular markers have developed from chloroplast genome, including gene-coding genes of rbcL and matK, as well as intergenic regions of trnH-psbA and trnL-F, which were widely used for species delimitation and phylogenetic analysis (Mishra et al., 2016). However, in algae, few studies were involved in the application of molecular markers from chloroplast (cp) genome. In this study, by investigating the intergenic regions of cp genome, we tried to develop novel marker with purpose to discrimination of Ulva species.
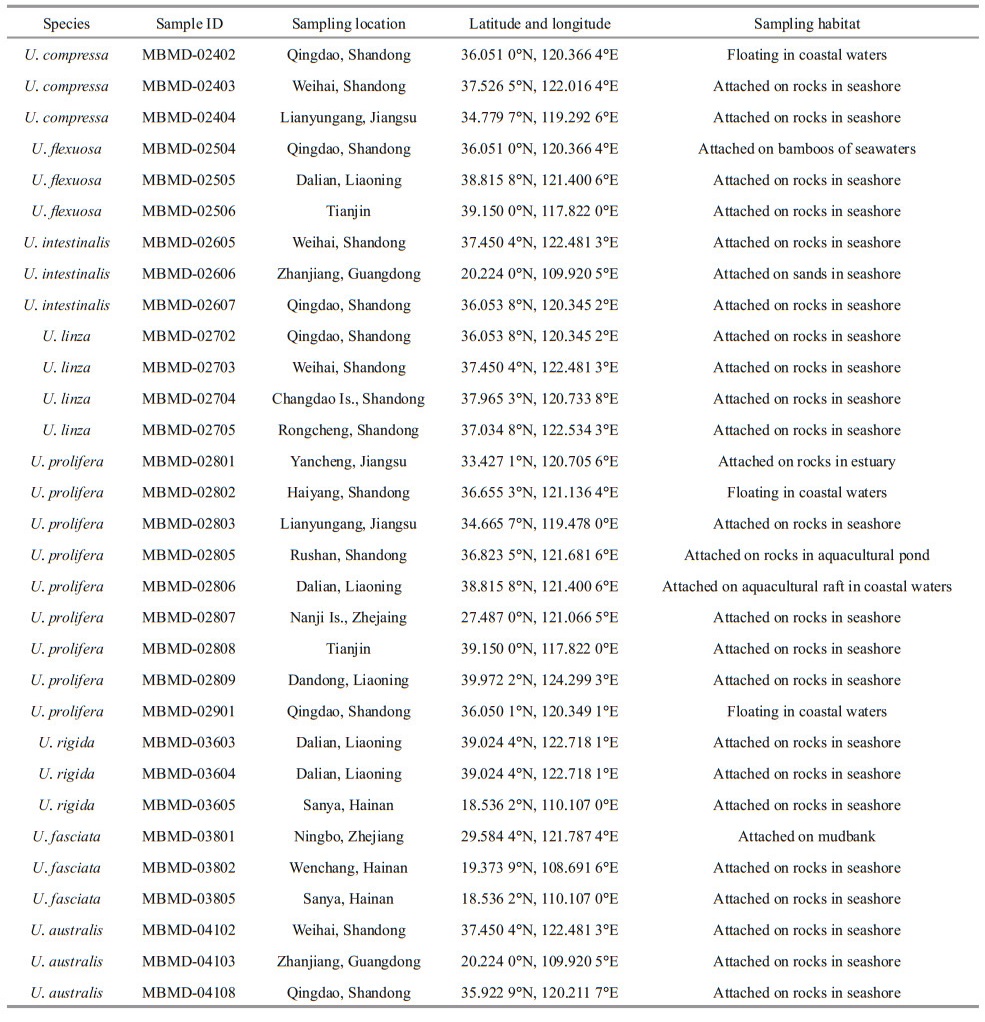
2 MATERIAL AND METHOD 2.1 Sampling and species identificationThe specimens of Ulva spp. used in this study were collected from the coast and offshore areas of China, which distributed in ecologically diverse locations (Fig. 1). For morphologically identification, the healthy adult individuals were selected and washed several times with sterile seawater to remove surface attachments, and then herbarium comparation were carried out according to the taxonomic features of Ulva taxa. Voucher herbarium specimens were deposited in the Marine Biological Museum of the Chinese Academy of Sciences (MBMCAS) (Fig. 2). For molecular identification, the genome DNA of each specimen were extracted and ITS barcoding were characterized via PCR amplification, sequencing, and Blast analysis.
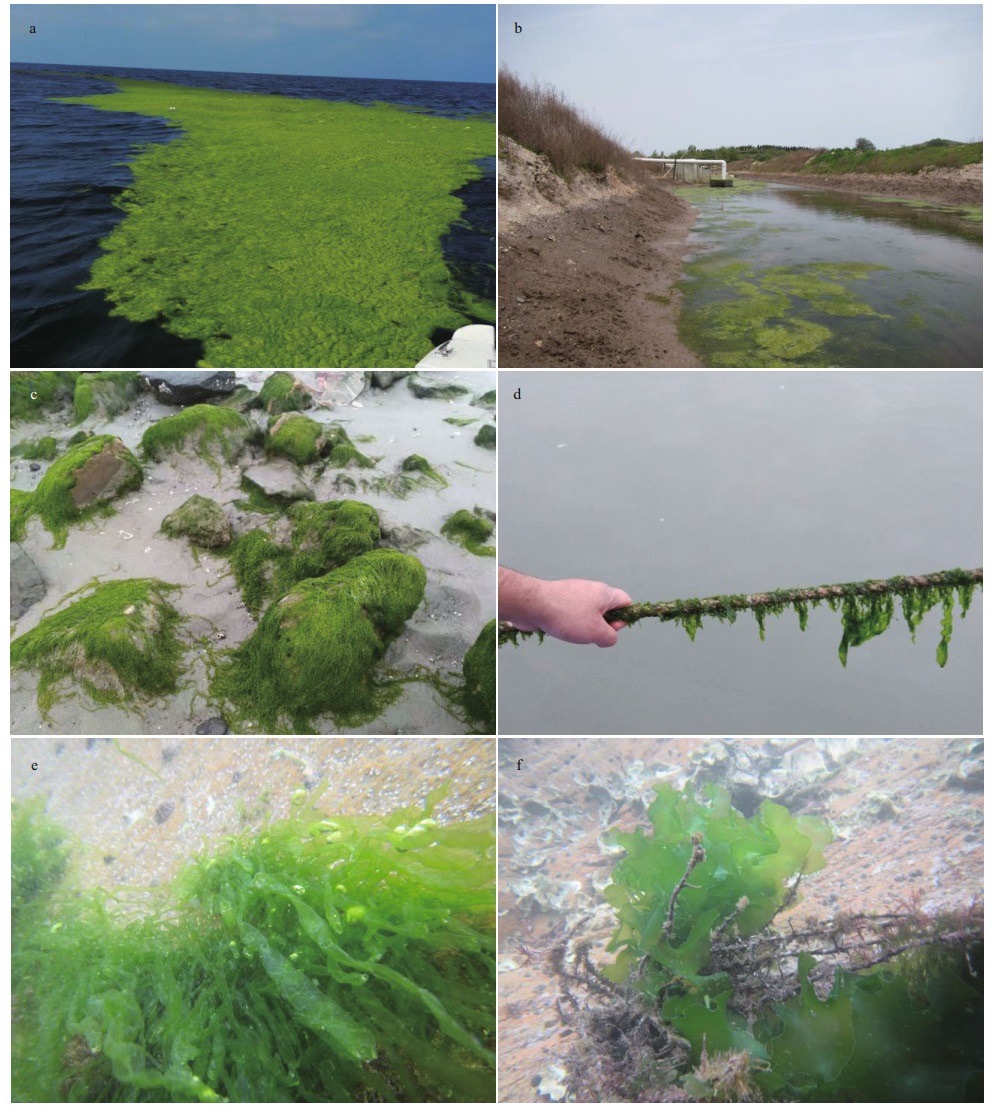

|
| Fig.1 Sampling of Ulva species a. floating on the sea; b. floating on aquaculture ponds; c. attaching on rocks in intertidal zone; d. attaching on aquaculture ropes; e and f. attaching on rocks in subtidal zone. |
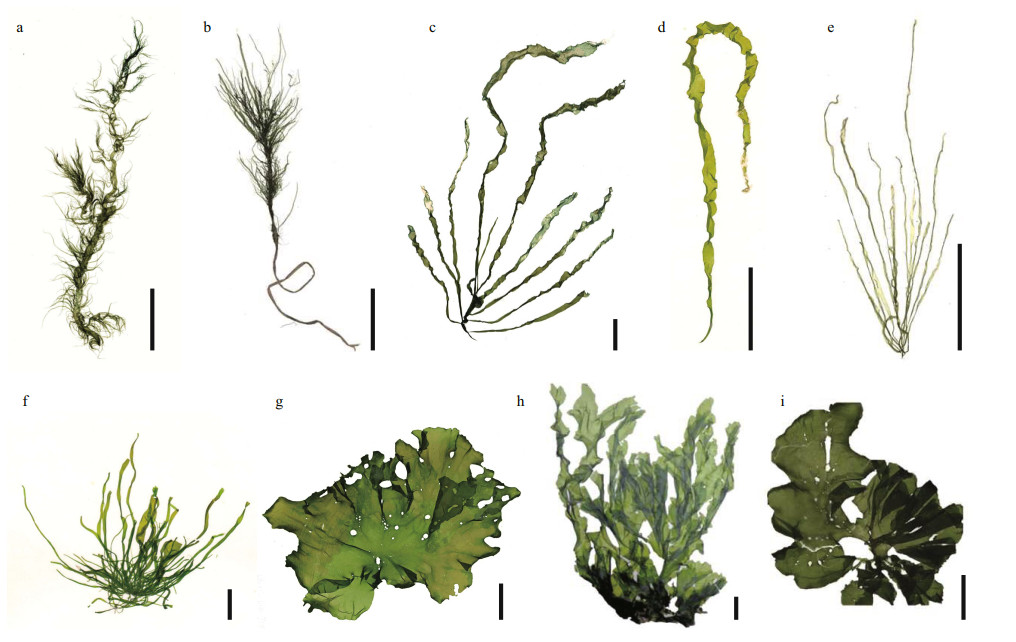

|
| Fig.2 The specimens of representative species in the genus of Ulva a. U. prolifera (floating); b. U. prolifera (attaching); c. U. compressa; d. U. linza; e. U. intestinalis; f. U. flexuosa; g. U. australis; h. U. fasciata; i. U. rigida. Scale bars: 2 cm. |
Total genomic DNA was extracted from fresh specimens with a DNeasy Plant kit (Tiangen Biotech. Co. Ltd., Beijing, China) following manufacturer's instructions. The ITS region was amplified by primers designed for Ulva taxa by Hayden et al. (2003) (F: 5′-TCTTTGAAACCGTATCGTGA-3′; R: 5′-GCTT-ATTGATATGCTTAAGTTCAGCGGGT-3′), and the c15 region of cp genomes were designed in this study (F: 5′-CTGACATACCATCACGATAGT-3′; R: 5′-CGTATTATTTCACCAGACCCTT-3′). The PCR reaction were carried out by TaKaRa Ex Taq enzyme in 25-L reaction column (TaKaRa, Shiga, Japan), with condition of an initial denaturation at 94 ℃ for 5 min, 36 cycles at 94 ℃ for 50 s, 54 ℃ for 50 s, and 72 ℃ for 1 min, and a final elongation step of 10 min at 72 ℃. All PCR products were examined by electrophoresis on a 1% agarose gel and then sequenced using autosequencer (ABI, 3730) according to manufacturer's instructions (ABI, BigDye® Terminator v3.1 Cycle Sequencing Kit).
2.3 Data analysisTo develop high-resolution markers, the cp genomes of U. prolifera (KX342867), U. linza (KX058323), and U. flexuosa (KX579943) were recruited for comparative genomic analysis. The sequences alignment of cp genomes was analyzed and the conserved regions were identified by mVISTA online software (https://genome.lbl.gov/vista/mvista/submit.shtml). Thus, several less conserved regions were selected for primers design and evaluated via routine PCR and sequencing analysis. For phylogenetic analysis, the sequences were aligned using Clustal module in MEGA software v.11 (Tamura et al., 2021) and then manually adjusted. The phylogenetic trees based on ITS and ycf3-rps7 sequences were constructed with Neighbor-joining (NJ), maximum likelihood (ML), and Bayesian inference (BI) methods, respectively. NJ analysis was carried out by MEGA software v.11 with 1 000 bootstrap replicates, whereas ML and BI analysis were carried out by PhyloSuite software (Zhang et al., 2020) following corresponding instructions. Trees were visualized using FigTree v.1.4.2 (http://tree.bio.ed.ac.uk/software/figtree/).
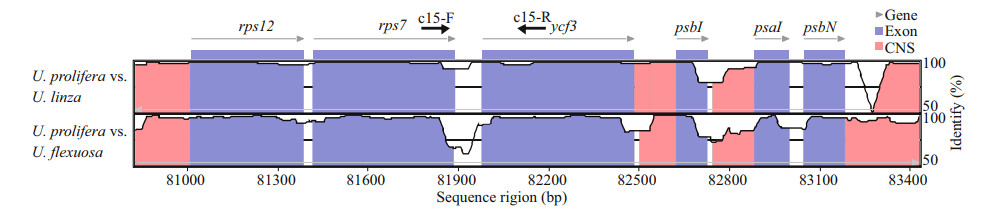
3 RESULT 3.1 Chloroplast genomes comparison and marker developmentBased on the cp genomes alignment of Ulva prolifera, U. linza, and U. fl exuosa, a high degree of conservation of synteny was exhibited, indicting the evolutionary relationships of these species was closely related (Fig. 3). It is shown that most coding regions were highly conserved, reflecting the evolutionary conservation of these genes to the function of chloroplast. In addition, there were conserved no-coding sequences (CNS) scattered in the alignment, suggesting these sequences might have conserved regulatory roles to the genes. Therefore, the less conserved sequences located in intergenic region could be theoretically used to develop molecular markers. Accordingly, to PCR-amplify these intergenic regions, dozens of primers were designed based on U. prolifera genomic sequences. After evaluation by gel electrophoresis and sequencing, it is showed that majority of primer pairs were failed to obtain a specific and unique PCR product. Nevertheless, a specific PCR product corresponding to the intergenic region of rps7 and ycf3 genes could be stabilized amplified by one primer pair labelled c15, exhibiting a potential role as novel molecular marker.
|
| Fig.3 Comparative analysis of the chloroplast genome of three Ulva species The arrows show the position of designed primers. |
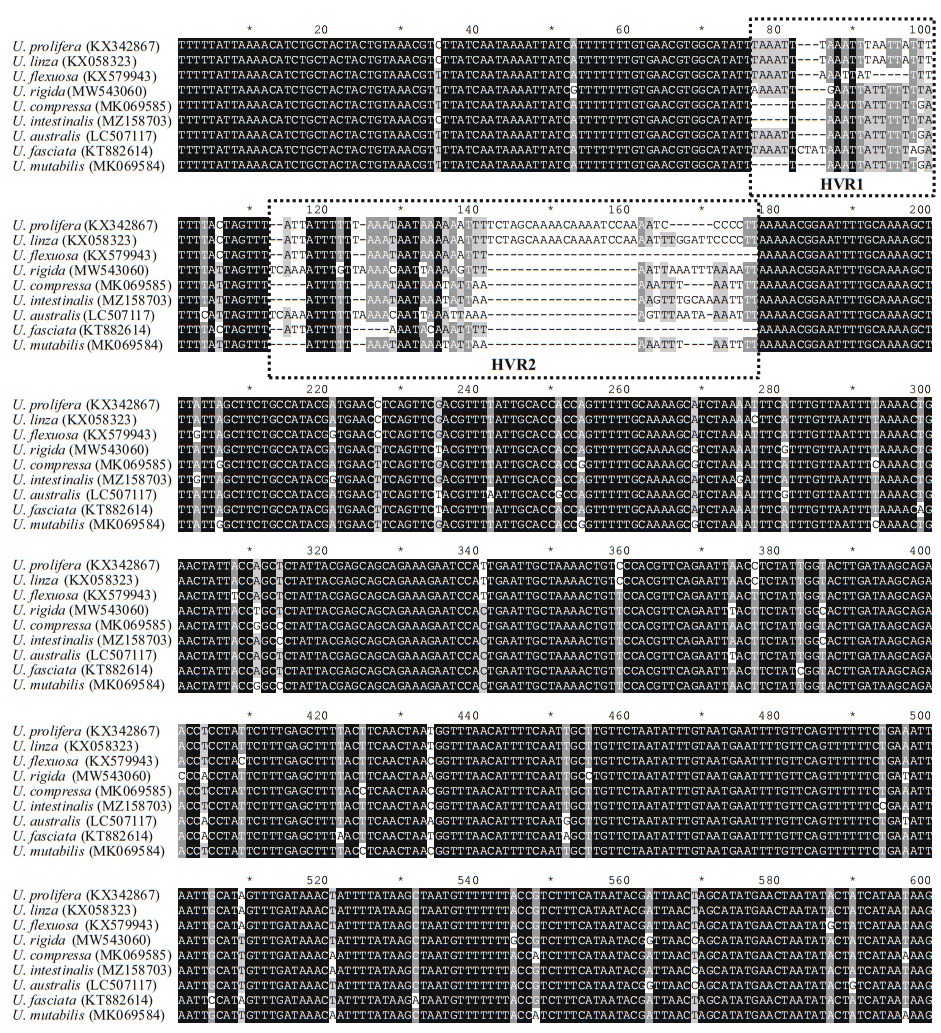
To evaluate the usability of rps7-ycf3 region, the sequence divergence between species was analyzed. The corresponding amplification regions of primer pair c15, which include intergenic sequences of rps7-ycf3, were retrieved from 9 published Ulva cp genomes. According to the sequence alignment results, the coding region were highly conserved (Fig. 4). Specifically, there were two highly variable regions (HVR1 and HVR2) located in the intergenic regions, suggesting these sequences divergence could be used for markers to distinguish species.
|
| Fig.4 Sequence alignment of rps7-ycf3 regions of representative Ulva cp genomes The dotted rectangle is two highly variable regions. HVR: highly variable region. |
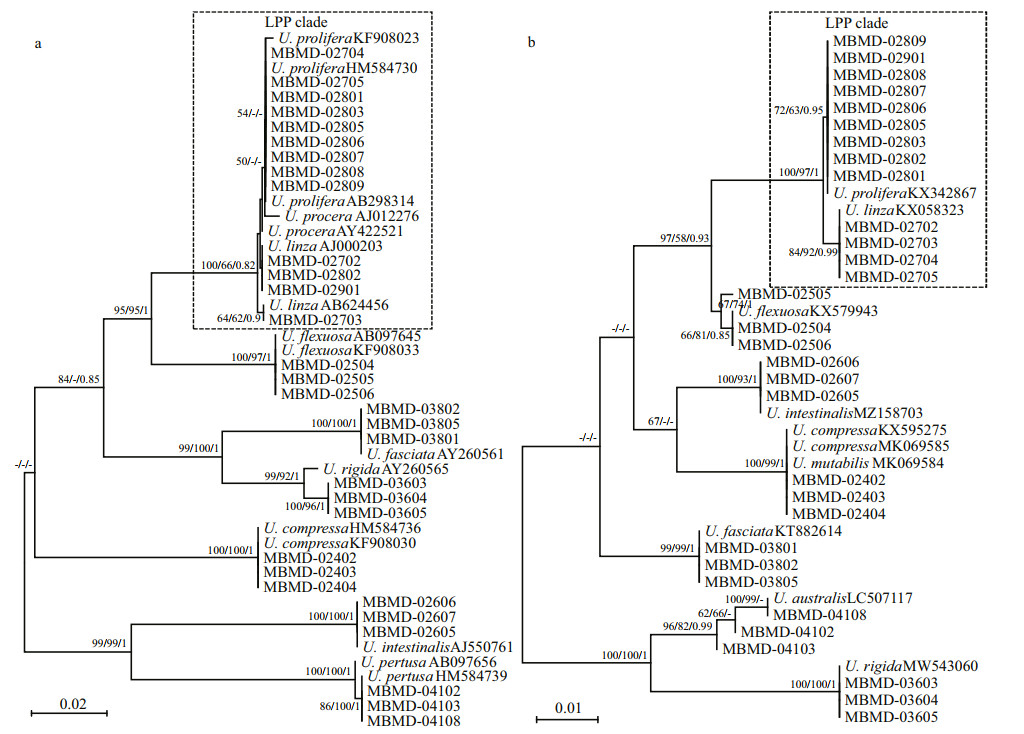
To validate the resolution of the c15 marker, the specimens collected from the coastal area of China were used for phylogenetic analysis (Table 1). These specimens could be identified as eight Ulva species based on their distinguished morphological characteristics (Fig. 2). U. linza and U. prolifera could not separate as monophyly in the ITS tree, which clustered into LPP complex as previous studies (Fig. 5a). However, in the c15 tree, U. linza and U. prolifera split into two separated clades clearly, suggesting the high resolution of this marker (Fig. 5b). Furthermore, the monophyly of other six Ulva species were unaffected, which was consistent with ITS or rbcL markers.
|
| Fig.5 Phylogenetic trees based on ITS (a) and rps7-ycf3 (b) sequences Bootstrap values and posterior probabilities are indicated in the branches (NJ/ML/BI). A bootstrap value or posterior probabilities below 50% or 0.50 is indicated by "-". The LPP clade is highlighted with dotted rectangle for each tree. |
The key to control the Ulva green tides is to identify the causal species accurately, which is important to prevent the occurrence of bloom. Several studies have demonstrated that the U. prolifera on bamboos, nets and ropes of aquaculture infrastructure were the seed source to green tides in the Yellow Sea (Liu et al., 2013, Zhang et al., 2017). As there were four major species of U. linza, U. compressa, U. flexuosa and U. prolifera been identified in Neopyropia cultivation area (Zhang et al., 2011), it is difficult to morphologically discriminate each another in the young germlings. Thus, the early interference by recognizing the U. prolifera specifically will increase the efficiency of controlling green tides blooms.
In this study, a novel marker was developed from cp genome for phylogenetically discriminating of Ulva species in a high-resolution manner. Specifically, comparing with tradition markers, c15 could precisely resolve the species relationships of U. linza and U. prolifera in LPP complex (Fig. 5b). Although the nuclear marker 5S rDNA were applied to distinct the species in LPP complex (Shimada et al., 2008, 2016), the specificity of this marker was doubted as incomplete concerted evolution (Liu et al., 2020). Moreover, the new c15 marker could be used for other frequently distributed Ulva species, indicating its potential to work as universal barcode to investigate the cryptic biodiversity of this taxa.
The identification of closely related species by molecular method was challengeable as the events of hybridization and gene flow between species occur frequently. As a result, different strategies are selected to solve problematic genera when resolution is desired. In most cases, the molecular markers developed from organelle genomes are uniparental inherited and single copy, providing resolution up to inter or intra-species levels (Lahaye et al., 2008; Matiz-Ceron et al., 2022). For example, the markers of atpF-atpH and trnH-psbA can simultaneously discriminate all the 46 representative medicinal plant species of 28 families (Thakur et al., 2019). However, given the different evolutionary pattern and reproductive mode between land plant and Ulva species, whether the cp genome of Ulva was inherited uniparentally need to be investigated.
In our analysis, the interspecific relationships were not completely identical between ITS and c15 phylogenies. The reasons for this inconsistent could be the different evolutionary rate and patterns of distinct markers (Wang et al., 2017). As both ITS and c15 are from no-coding intergenic region, it is suitable to identify taxa but not analyze phylogenetic relationships. Therefore, emerging phylogenomic analysis combining protein-coding markers could expected to better understanding the phylogenetic relationships for Ulva species. Besides, the intraspecific diversification in Ulva species were not well-solved, which need novel markers been developed.
5 CONCLUSIONIn this study, by investigating the intergenic regions of chloroplast (cp) genome, a novel marker of c15 (based on ycf3-rps7 region) were reliable for discrimination of Ulva species. Notably, the resolution of c15 was suitable for resolve the closely related species of U. linza and U. prolifera, which may provide tools for deciphering the blooming mechanisms of green tides.
6 DATA AVAILABILITY STATEMENTThe datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Blomster J, Bäck S, Fewer D P, et al. 2002. Novel morphology in Enteromorpha (Ulvophyceae) forming green tides. American Journal of Botany, 89(11): 1756-1763.
DOI:10.3732/ajb.89.11.1756 |
Hayden H S, Blomster J, Maggs C A, et al. 2003. Linnaeus was right all along: Ulva and Enteromorpha are not distinct genera. European Journal of Phycology, 38(3): 277-294.
DOI:10.1080/1364253031000136321 |
Hiraoka M, Ichihara K, Zhu W R, et al. 2011. Culture and hybridization experiments on an Ulva clade including the Qingdao strain blooming in the Yellow Sea. PLoS One, 6(5): e19371.
DOI:10.1371/journal.pone.0019371 |
Lahaye R, Savolainen V, Duthoit S et al. 2008. A test of psbK-psbI and atpF-atpH as potential plant DNA barcodes using the flora of the Kruger National Park (South Africa) as a model system. Nature Precedings, https://doi.org/10.1038/npre.2008.1896.1.
|
Liu D Y, Keesing J K, He P M, et al. 2013. The world's largest macroalgal bloom in the Yellow Sea, China: Formation and implications. Estuarine, Coastal and Shelf Science, 129: 2-10.
DOI:10.1016/j.ecss.2013.05.021 |
Liu J L, Tong Y C, Xia J, et al. 2022. Ulva macroalgae within local aquaculture ponds along the estuary of Dagu River, Jiaozhou Bay, Qingdao. Marine Pollution Bulletin, 174: 113243.
DOI:10.1016/j.marpolbul.2021.113243 |
Liu J L, Zhao X H, Kang X Y, et al. 2020. Good news: we can identify Ulva species erupted in the Yellow Sea more easily and cheaply now. Conservation Genetics Resources, 12(3): 447-449.
DOI:10.1007/s12686-019-01114-x |
Liu Q, Yu R C, Yan T, et al. 2015. Laboratory study on the life history of bloom-forming Ulva prolifera in the Yellow Sea. Estuarine, Coastal and Shelf Science, 163: 82-88.
DOI:10.1016/j.ecss.2014.08.011 |
Matiz-Ceron L, Reyes A, Anzola J. 2022. Taxonomical evaluation of plant chloroplastic markers by bayesian classifier. Frontiers in Plant Science, 12: 782663.
DOI:10.3389/fpls.2021.782663 |
Mishra P, Kumar A, Nagireddy A, et al. 2016. DNA barcoding: an efficient tool to overcome authentication challenges in the herbal market. Plant Biotechnology Journal, 14(1): 8-21.
DOI:10.1111/pbi.12419 |
Pang S J, Liu F, Shan T F, et al. 2010. Tracking the algal origin of the Ulva bloom in the Yellow Sea by a combination of molecular, morphological and physiological analyses. Marine Environmental Research, 69(4): 207-215.
DOI:10.1016/j.marenvres.2009.10.007 |
Qi L, Hu C M, Xing Q G, et al. 2016. Long-term trend of Ulva prolifera blooms in the western Yellow Sea. Harmful Algae, 58: 35-44.
DOI:10.1016/j.hal.2016.07.004 |
Shen W J, He Y, Shen S D. 2019. A new molecular label applied to the study of the Yellow Sea green tide. Journal of Ocean University of China, 18(6): 1507-1514.
DOI:10.1007/s11802-019-3988-1 |
Shimada S, Ichihara K, Masakiyo Y et al. 2016. Phylogeography of macroalgal species distributed in brackish water: Ulva prolifera (Ulvophyceae) and Pyropia tenera (Bangiophyceae). In: Hu Z M, Fraser C eds. Seaweed Phylogeography. Springer, Dordrecht. p. 345-360.
|
Shimada S, Yokoyama N, Arai S, et al. 2008. Phylogeography of the genus Ulva (Ulvophyceae, Chlorophyta), with special reference to the Japanese freshwater and brackish taxa. Journal of Applied Phycology, 20(5): 979-989.
DOI:10.1007/s10811-007-9296-y |
Smetacek V, Zingone A. 2013. Green and golden seaweed tides on the rise. Nature, 504(7478): 84-88.
DOI:10.1038/nature12860 |
Tamura K, Stecher G, Kumar S. 2021. MEGA11: molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution, 38(7): 3022-3027.
DOI:10.1093/molbev/msab120 |
Thakur V V, Tiwari S, Tripathi N, et al. 2019. Molecular identification of medicinal plants with amplicon length polymorphism using universal DNA barcodes of the atpF-atpH, trnL and trnH-psbA regions. 3 Biotech, 9(5): 188.
DOI:10.1007/s13205-019-1724-6 |
Wang A, Gopurenko D, Wu H, Lepschi B. 2017. Evaluation of six candidate DNA barcode loci for identification of five important invasive grasses in eastern Australia. PLoS One, 12(4): e0175338.
DOI:10.1371/journal.pone.0175338 |
Xie E Y, Xu R S, Zhang J H, et al. 2020a. Growth characteristics of hybrids produced by closely related Ulva species. Aquaculture, 519: 734902.
DOI:10.1016/j.aquaculture.2019.734902 |
Xie W F, Wu C H, Zhao J, et al. 2020b. New records of Ulva spp.(Ulvophyceae, Chlorophyta) in China, with special reference to an unusual morphology of U. meridionalis forming green tides. European Journal of Phycology, 55(4): 412-425.
DOI:10.1080/09670262.2020.1740946 |
Zhang D, Gao F L, Jakovlić I, et al. 2020. PhyloSuite: an integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Molecular Ecology Resources, 20(1): 348-355.
DOI:10.1111/1755-0998.13096 |
Zhang J H, Zhao P, Huo Y Z, et al. 2017. The fast expansion of Pyropia aquaculture in "Sansha" regions should be mainly responsible for the Ulva blooms in Yellow Sea. Estuarine, Coastal and Shelf Science, 189: 58-65.
DOI:10.1016/j.ecss.2017.03.011 |
Zhang Q C, Yu R C, Chen Z F, et al. 2018. Genetic evidence in tracking the origin of Ulva prolifera blooms in the Yellow Sea, China. Harmful Algae, 78: 86-94.
DOI:10.1016/j.hal.2018.08.002 |
Zhang X W, Xu D, Mao Y Z, et al. 2011. Settlement of vegetative fragments of Ulva prolifera confirmed as an important seed source for succession of a large-scale green tide bloom. Limnology and Oceanography, 56(1): 233-242.
DOI:10.4319/lo.2011.56.1.0233 |
Zhang Y Y, He P M, Li H M, et al. 2019. Ulva prolifera green-tide outbreaks and their environmental impact in the Yellow Sea, China. National Science Review, 6(4): 825-838.
DOI:10.1093/nsr/nwz026 |
Zhao J, Jiang P, Liu Z Y, et al. 2012. The Yellow Sea green tides were dominated by one species, Ulva (Enteromorpha) prolifera, from 2007 to 2011. Chinese Science Bulletin, 58(19): 2298-2302.
|
Zhao J, Jiang P, Qin S, et al. 2015. Genetic analyses of floating Ulva prolifera in the Yellow Sea suggest a unique ecotype. Estuarine, Coastal and Shelf Science, 163: 96-102.
DOI:10.1016/j.ecss.2015.05.027 |
Zhao X H, Cui J J, Zhang J H, et al. 2019. Reproductive strategy of the floating alga Ulva prolifera in blooms in the Yellow Sea based on a combination of zoid and chromosome analysis. Marine Pollution Bulletin, 146: 584-590.
DOI:10.1016/j.marpolbul.2019.07.018 |
 2022, Vol. 40
2022, Vol. 40