Institute of Oceanology, Chinese Academy of Sciences
Article Information
- HU Zhangxi, SONG Xiaoying, WANG Jinxiu, TAO Zhe, SUN Yuanyuan, LI Yuhang, LIU Yuyang, DENG Yunyan, SHANG Lixia, CHAI Zhaoyang, TANG Yingzhong
- Reviving and characterizing three species of dinoflagellate cysts dormant for about 70 years in the East China Sea: Biecheleria brevisulcata, Biecheleriopsis adriatica, and Scrippsiella donghaienis
- Journal of Oceanology and Limnology, 40(6): 2292-2311
- http://dx.doi.org/10.1007/s00343-022-2122-1
Article History
- Received Mar. 14, 2022
- accepted in principle Apr. 18, 2022
- accepted for publication Jun. 4, 2022
2 CAS Key Laboratory of Marine Ecology and Environmental Sciences, Institute of Oceanology, Chinese Academy of Sciences, Qingdao 266071, China;
3 Laboratory of Marine Ecology and Environmental Science, Pilot National Laboratory for Marine Science and Technology(Qingdao), Qingdao 266237, China;
4 CAS Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, Qingdao 266071, China;
5 Laboratory of Marine Organism Taxonomy and Phylogeny, Institute of Oceanology, Chinese Academy of Sciences, Qingdao 266071, China;
6 Center for Ocean Mega-Science, Chinese Academy of Sciences, Qingdao 266071, China;
7 University of Chinese Academy of Sciences, Beijing 100049, China
Dinoflagellates constitute one of the main groups of marine phytoplankton in terms of their important contribution to the primary production, and comprise about 2 400 species belonging to 259 genera (Gómez, 2012), which is still growing as more new taxa are to be described (Gómez et al., 2015; Takahashi et al., 2015, 2017, 2019; Boutrup et al., 2017; Luo et al., 2018; Hu et al., 2020b, 2021; Ok et al., 2021; Gu et al., 2022). The appearance of dinoflagellates dates back more than 400 million years in the fossil record, and this group of protists has evolved diverse features to adapt to their environments, including diverse morphologies, multiple-membrane cell walls, different modes of nutrition, pigments, toxins, asexual and sexual reproduction, and other characters (Steidinger and Meave del Castillo, 2018). Among these features, the ability of about 10% of dinoflagellate species to produce resting cysts as a part of their life cycle is a vital one (Head, 1996; Bravo and Figueroa, 2014; Tang et al., 2016, 2021). It is now widely accepted that resting cysts play important roles in the biology and ecology of dinoflagellates (Anderson and Wall, 1978; Dale, 2001; Bravo and Figueroa, 2014; Tang et al., 2016, 2021; Ellegaard and Ribeiro, 2018; Figueroa et al., 2018).
In the past, field studies on dinoflagellate cysts are generally focused on the surface sediments for mapping the distribution and abundance of cysts of important species, investigating species diversity of cyst assemblage in a region of concern, and confirming the cyst presence for some species with particular importance (Luo et al., 2018; Limoges et al., 2020; Liu et al., 2020a, b, 2021; Mertens et al., 2020; Van Nieuwenhove et al., 2020; Hu et al., 2021, 2022), whereas sediment cores representing a valuable archive of phytoplankton communities are usually used to reconstruct past environmental changes (Keafer et al., 1992; Dai et al., 2012; Ellegaard et al., 2013, 2020; Bringué et al., 2016; García-Moreiras et al., 2018; Kim et al., 2018; Price et al., 2018; de Freitas et al., 2020; Li et al., 2021; Siano et al., 2021). Cultures established from reviving dinoflagellate resting cysts collected from sediment cores have been used to evaluate the impact of environmental changes on the physiology, genetic structure, and diversity in various species (Ribeiro et al., 2011; Klouch et al., 2016; Lundholm et al., 2017; Kremp et al., 2018; Delebecq et al., 2020; Ellegaard et al., 2020; Girault et al., 2021). However, the numbers of dinoflagellate cysts remaining viable in long-buried sediments and the mechanisms behind it still need to be explored. Therefore, mining cyst records of dinoflagellate species buried in sediment cores is important in many aspects of the ecology of dinoflagellates and in reconstruction of the history of marine environmental changes.
Recently, we successfully established seven clonal cultures of Biecheleria brevisulcata, Biecheleriopsis adriatica, and Scrippsiella donghaienis via cystgermination from the depth dated back to 1941±18 AD of a sediment core collected from the East China Sea, and further characterized their morphologies, pigment compositions, and the genetic diversity in their LSU rRNA gene and ITS sequences.
2 MATERIAL AND METHOD 2.1 Sediment core and surface sediment sampling and datingOne sediment core (S06-2, 120.417°E, 26.122°N) was collected from the East China Sea in September 19, 2018 during the public cruise of R/V Xiang Yang Hong 18 organized by the National NaturalScience Foundation of China and the First Institute of Oceanography, Ministry of Natural Resources, China, and one surface sediment sample (0‒2 cm; S01-1, 122.997°E, 31.000°N) was also collected from the East China Sea in September 17, 2019 during the public cruise of R/V Xiang Yang Hong 18. The sediment core was sliced into 2-cm layers (the top 20 cm) and 4-cm layers (20 cm to the bottom) by caution. A total of 16 subsamples were collected for the 210Pb and 137Cs measurements. The detailed measurement and age determination was reported in Liu et al. (2021), and the standard error of age was produced in regression uncertainties.
2.2 Culture establishmentCyst assemblage in the subsample (42‒44 cm) of core sediment (S06-2) and surface sediment sample (S01-1) was concentrated using sodium polytungstate solution (SPT) (Bolch, 1997). For subsample of core sediment, single cysts were washed at least three times using sterile seawater (with a salinity of 31) enriched with f/2-Si medium (Guillard, 1975), and then micropipetted to a 24-well culture plate with each well containing 2.5-mL fresh medium and 2% antibiotic solution (a mixture of 10 000-IU penicillin and 10 000-μg/mL streptomycin; Solarbio, Beijing, China). The rest of cyst assemblage was transferred to a 6-well culture plate with each well containing 10-mL fresh medium and 2% antibiotic solution. The plates were incubated at 21 ℃, 12-h꞉12-h light꞉dark cycle, and ~100 μmol photons/(m2·s). Cysts and new germlings were observed every day or every other day with an inverted microscope (IX73, Olympus, Japan) and photographed by a DP80 digital camera (Olympus, Japan). Five strains of B. brevisulcata (S1, S2, S3, S4, and S5) were established from the cyst assemblage germination experiment, but the resting and empty cysts of B. brevisulcata were not observed. One strain of Bps. adriatica (S21) and one strain of S. donghaienis (S23) were established from singlecysts germination experiments, their resting and empty cysts were clearly recorded using an inverted microscope (IX73, Olympus, Japan) equipped with a DP80 digital camera (Olympus, Japan). All cultures were routinely maintained in the same condition mentioned above.
2.3 Light microscopic observationLive cells of B. brevisulcata (strain S1), Bps. adriatica (strain S21), and S. donghaienis (strain S23) were observed and photographed using a Zeiss Imager Z2 (Carl Zeiss, Gottingen, Germany) equipped with differential interference contrast (DIC), or an inverted microscope (IX73, Olympus, Japan) equipped with a digital camera (DP80, Olympus, Japan). For observation of thecal plates of S. donghaienis, live cells were stained with Calcofluor White (Sigma-Aldrich, St. Louis, MO, USA) and examined using an epifluorescence microscope (BX53, Olympus, Japan) with a UV filter set (Fritz and Triemer, 1985). Cells sizes of B. brevisulcata, Bps. adriatica, and S. donghaienis for 50 live cells at the mid-exponential growth phase were measured at ×400 (for B. brevisulcata and Bps. adriatica), and ×200 (for S. donghaienis) magnification using a DP80 digitalcamera (Olympus, Tokyo, Japan).
2.4 SEM observationFor SEM observation, vegetative cells of B. brevisulcata and Bps. adriatica at mid-exponentialgrowth stage were fixed with OsO4 (2% final concentration), and S.donghaieniswith glutaraldehyde (2.5% final concentration) for 40–50 min. Fixed cells were gently filtered onto 5-μm (B. brevisulcata and Bps. adriatica) and 11-μm (S. donghaienis) pore sizeMillipore nylon membranes, dehydrated in an acetone series (10%, 30%, 50%, 70%, 90%, and three times in 100%, 15 min for each step), and critical point-dried with liquid CO2 (EM CPD300, Leica, Austria). They were sputter-coated with platinum-palladium (EM ACE200, Leica, Austria), and observed using an S-3400N SEM (Hitachi, Japan).
2.5 Pigment analysesFifty milliliter of each culture of B. brevisulcata (strain S1), Bps. adriatica (strain S21), and S. donghaienis (strain S23) in exponential growth(in cell densities of ca. 102 150, 95 900, and 980 cells/mL, respectively) were filtered through 25-mm diameter glass fiber filter (Whatman, Maidstone, UK) and immediately frozen at -80 ℃ for later analyses. Pigments were analyzed on an Alliance HPLC (e2695, Waters, Milford, Massachusetts, USA) using a 100-μL sample injection according to Kong et al. (2012) and Hu et al. (2020a). Pigments were identified and quantified using Shimadzu Class-VP software and by comparing pigment spectra and retention times with those of 26 standard pigments (DHI Water and Environment, H rsholm, Denmark; Kong et al., 2012; Hu et al., 2020a).
2.6 DNA extraction, PCR amplification, and rDNA-based phylogenetic analysesGenomic DNA of B. brevisulcata, Bps. adriatica, and S. donghaienis were extracted using a plant DNA extraction kit (Tiangen, Beijing, China) according to the manufacturer's protocol. For B. brevisulcata, Bps. adriatica, and S. donghaienis, about 1 400 bpof LSU rDNA were amplified using primers of D1R (forward, 5′-ACCCGCTGAATTTAAGCATA-3′) (Scholin et al., 1994) and 28-1483R (reverse, 5′-GCTACTACCACCAAGATCTGC-3′) (Daugbjerg et al., 2000), and for S. donghaienis, about 660 bp of ITS was amplified using primers of ITS1 (forward, 5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (reverse, 5′-GCATATCAATAAGCGGAGGA-3′) (White et al., 1990). Polymerase chain reaction (PCR) reactions were conducted using a PCR Master Cycler nexus gradient (Eppendorf, Hamburg, Germany), and performed with a final volume of 25 μL, containing 9.5-μL ddH2O, 12.5-μL 2×Taq PCR MasterMix, 1 μL of each PCR primer (10 mmol/L), and 1 μL of the DNA template. The following cycling conditions were used: an initial denaturation at 94 ℃ for 5 min, 35 cycles at 94 ℃ for 20 s, 55 ℃ for 30 s, and 72 ℃ for 2 min, and a final elongation step of 10 min at 72 ℃. The PCR products were confirmed using 1% agarose gel electrophoresis and visualized with ultraviolet light. Targeted bands were purified using an agarose gel DNA fragment recovery kit (GENEray Biotechnology, Shanghai, China), ligated with pMD18-T cloning vector (TaKaRa, Tokyo, Japan), and then sequenced (Sangon, Shanghai, China). Sequences were deposited in GenBank with accession numbers OL355144‒OL355148 (LSU, B. brevisulcata), OL355142 (LSU, Bps. adriatica), OL314541 (LSU, S. donghaienis), and OL314542 (ITS, S. donghaienis).
For surface sediment sample (S01-1), the single cysts were individually micropipetted onto a glass slide, then photographed with an inverted microscope (IX73, Olympus, Japan) equipped with a DP80 digital camera (Olympus, Japan). Subsequently, individual cyst was micropipetted and transferred onto a sterile slide, then broken by another coverslip. The crushed cyst and the coverslip pieces were transferred into a 250-μL centrifuge tube, as the template to amplify about 1 400 bp of the LSU rDNA using the primer set, D1R (forward, 5′-ACCCGCTGAATTTAAGCATA-3′) (Scholin et al., 1994) and 28-1483R (reverse, 5′-GCTACTACCACCAAGATCTGC-3′) (Daugbjerg et al., 2000). The following procedure was according to Shang et al. (2019).
For phylogenetic analyses of the LSU rDNA regions of B. brevisulcata and Bps. adriatica, and LSU and ITS rDNA regions of S. donghaienis, newly obtained LSU rDNA and ITS sequences were incorporated into those of closely related species available in the GenBank and that of outgroup taxa were first aligned using MAFFT v7.475 (Katoh et al., 2002) online program (http://mafft.cbrc.jp/alignment/server/) with default settings, and alignments were manually checked with BioEdit v7.2.5 (Hall, 1999). The final alignments of the LSU and ITS rDNA sequences of S. donghaienis consisted of 97 and 41 taxa and contained 750 and 600 positions (including gaps introduced from alignment), and the sequences of Cryptoperidiniopsis brodyi (DQ991374) and Pentapharsodinium dalei (JX262496) were used asthe outgroup, respectively. LSU rDNA sequences of Bps. adriatica and B. brevisulcata consisted of 66taxa with 1 590 positions (including gaps introduced from alignment), and the sequence of Alexandrium margalefii (AY154957) was used as outgroup. Theprogram jModelTest 2.1.4 was used to select the most appropriate model of molecular evolution with Akaike information criterion (AICc) (Posada, 2008), models GTR+G+I and TrN+I+G were selected as the best-fit model for the LSU rDNA and ITS datasets of S. donghaienis, and TrN+I+G for the LSU rDNA dataset of Bps. adriatica and B. brevisulcata. Phylogenetic trees were constructed using Bayesian inference (BI) and maximum likelihood (ML) analyses. Bayesian inference (BI) was performed with MrBayes 3.2.6 (Ronquist and Huelsenbeck, 2003) with the best-fitting substitution models (GTR+I+G for LSU rDNA dataset of S. donghaienis, TrN+I+G for ITS dataset of S. donghaienis, and TrN+I+G for LSU rDNA dataset of Bps. adriatica and B. brevisulcata). Four independent Markov chain Monte Carlo simulations were run simultaneously for 5 000 000 generations and trees were sampled every 1 000 generations. The first 10% trees were discarded as burn-in. The convergence was judged based on the average standard deviation of split frequencies (all less than 0.01). The remaining trees were used to generate a consensus tree and calculate the posterior probabilities of all branches using a majority-rule consensus approach. Maximum likelihood (ML) analyses were conducted with raxmlGUI v1.3.1 (Silvestro and Michalak, 2012; Stamatakis, 2014) using the models GTR+I+G (for LSU rDNA dataset of S. donghaienis), GTR+G (for ITS dataset of S. donghaienis; the model GTR+G ranked the third, andthe score of this model was close to model TrN+I+G), and GTR+I+G (for LSU rDNA dataset of Bps. adriatica and B. brevisulcata; the model GTR+I+Granked the second, and the score of this model was close to model TrN+I+G). Node support was assessed with 1 000 bootstrap replicates. FigTree (v1.4.4) was used to view and edit trees for publication.
2.7 Genetic diversity analysesThe pairwise distances were computed among all sequences that were newly obtained in the present work for B. brevisulcata, Bps. adriatica, and S. donghaienis and that retrieved from the NCBIdatabase for these three species together with other reference sequences. Sequences were aligned using the MAFFT v7.475 with the default settings (Katoh et al., 2002) (http://mafft.cbrc.jp/alignment/server/) and modified manually using BioEdit v7.2.5 (Hall, 1999). Pairwise evolutionary distances were then computed using Jukes and Cantor algorithm implemented in the MEGA X (Tamura et al., 2004; Kumar et al., 2018).
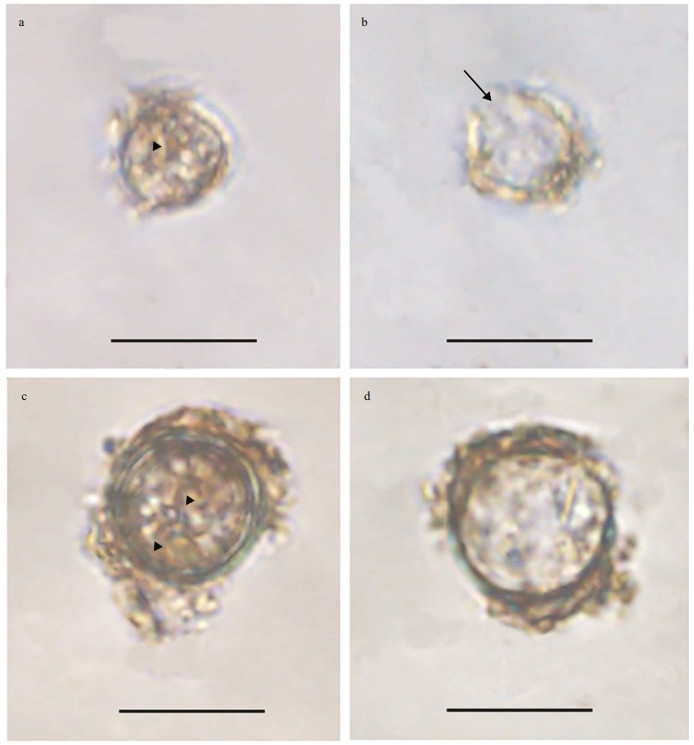
3 RESULT 3.1 Morphological observations of the resting cystsResting cyst of Bps. adriatica was sub-spherical to spherical and light brown, with a diameter of ~7.3 μm, full of small granules and had 2–3 red accumulation bodies (Fig. 1a). After four days' incubation, the cyst was germinated. The cyst wall was thick and smooth, and the archeopyle was tremic (Fig. 1b). Resting cyst of S. donghaienis was noncalcareous, spherical and brown, full of different sizes of granules (Fig. 1c). The diameter of the cyst was ~10.7 μm, and it contained several red accumulation bodies (Fig. 1c). After four days' incubation, the cyst was germinated. The cyst wall was thick, and the archeopyle was unclear (Fig. 1d). As the five clonal cultures of B. brevisulcata were established from cyst assemblage germination, the resting and empty cysts of this species were not observed.
|
| Fig.1 Light microscopy photographs of Bps. adriatica and S. donghaienis cysts a, b. resting cyst and empty cyst of Bps. adriatica (arrowhead and arrow indicate the accumulation body and archeopyle); c, d. resting cyst and empty cyst of S. donghaienis (arrowheads indicate the accumulation bodies). Scale bars=10 μm. |
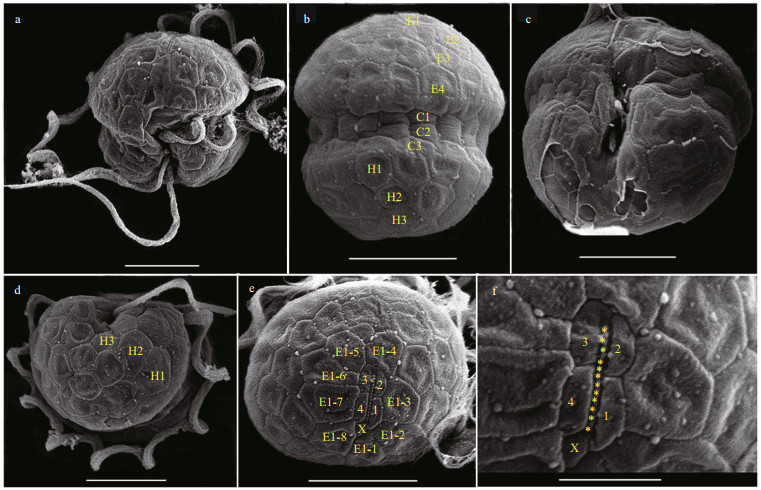
The vegetative cells of B. brevisulcata were spherical to ellipsoidal, 7.2–9.2 μm long (average 8.2±0.6 μm; n=50) and 5.8–8.9 μm wide (average 7.3±0.7 μm; n=50) (Figs. 2–3). The epicone was slightly wider and longer than the hypocone, and mushroom-shaped (Figs. 2–3). The hypocone was bilobed (Figs. 2–3). The descending cingulum was deep and wide, and displaced by 1.5 times its own width (Figs. 2a–b, 3a–b). The sulcus was in the form of a sigmoid curve (Figs. 2a, b, d, & 3a). The nucleus was round and located in the middle or slightly upper part of the hypocone (Fig. 2f). Numerous yellow-brownish and reticulated or granulated chloroplasts were distributed peripherally (Fig. 2). The arrangement of polygonal amphiesmal vesicles (AVs) was shown in Fig. 3. Latitudinal rows of AVs were in four series on the epicone (E1‒E4), three series on the hypocone (H1‒H3), and three series in the cingular area (C1‒ C3; Fig. 3b, d & e). A narrow elongate apical vesicle (EAV) was present in the apical area (Fig. 3e–f), which was surrounded by five AVs (a small four-sided AV (X) and four elongated quadrangular AVs) and eight irregular pentagonal AVs (E1-1‒E1-8; Fig. 3e–f).

|
| Fig.2 Light microscopy photographs of B. brevisulcata strain S1 germinated from resting cyst a. surface of ventral view showing sulcus and cingulum; b. deeper focus of ventral view showing cingulum; c. surface of dorsal view showing cingulum; d. dorsal-antapical view showing sulcus; e. antapical view; f. ventral view of a planozygote showing sulcus, nucleus (N), and two longitudinal flagella (arrows).Scale bars=10 μm. |
|
| Fig.3 Scanning electron micrographs (SEM) of B. brevisulcata strain S1 germinated from resting cyst a. ventral view showing sulcus, cingulum, transverse, and longitudinal flagella; b. dorsal view showing four latitudinal amphiesmal vesicle series on the epicone (E1–E4), three vesicle series in the cingulum (C1–C3), and three latitudinal vesicle series on the hypocone (H1–H3); c. antapic-ventral view showing surface area lacking the outer amphiesmal membrane; d. antapical view showing three latitudinal vesicle series on the hypocone (H1–H3); e. apical view showing an apical furrow composed of an elongate apical vesicle, surrounded by four amphiesmal vesicles (1-4) and a small vesicle (X). The apical furrow area is surrounded by eight vesicles (E1-1–E1-8); f. apical view showing knobs (yellow asterisks) on the single elongate apical vesicle, surrounded by four amphiesmal vesicles (1-4) and a small vesicle (X). Scale bars=5 μm (a–e), 2 μm (f). |
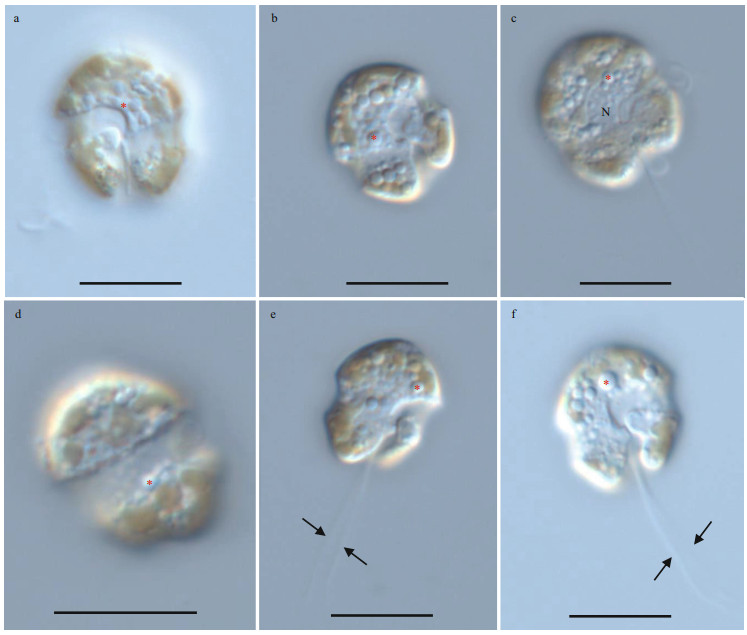
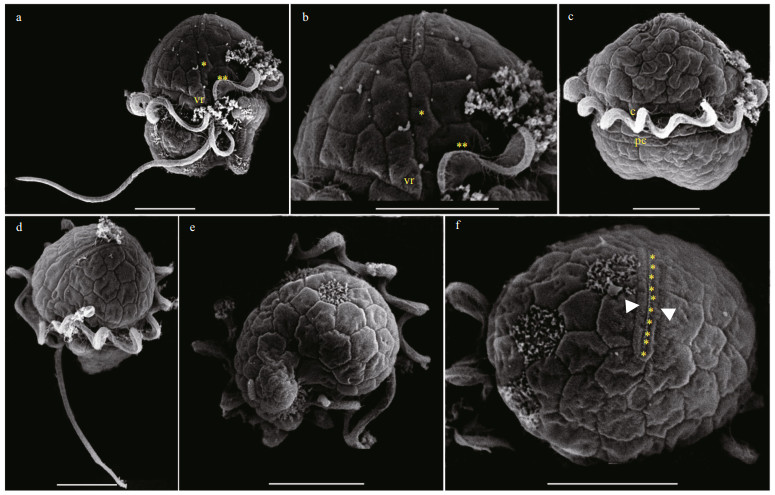
The vegetative cells of Bps. adriatica was spherical to ellipsoid, 5.8–10.1 μm in length (average 7.8±1.2 μm; n=50) and 4.1–8.0 μm in width (average 6.0±0.9 μm; n=50). The epicone was rounded, and almost equal to the hypocone (Figs. 4–5). The cingulum was deeper and wider than the sulcus (Figs. 4–5). The cingulum was median and displaced by 1.5 times its own width (Figs. 4–5). The sulcus was slightly sigmoid (Figs. 4a–b, e–f, & 5a). Chloroplasts were yellowish-brown, and reticulated or granulated, which were distributed peripherally (Fig. 4). The round refractive bodies were commonly observed (Fig. 4). The nucleus was located in the center or slightly upper part of the hypocone (Fig. 4c). Many pentagonal or hexagonal AVs were observed on the cell surface (Fig. 5). Four epiconal, three cingular, and four hypoconal AV series were formed the latitudinal series (Fig. 5a & c). The apical furrow was composed of an EAV and several surrounding AVs (Fig. 5a–b, d, & f).
|
| Fig.4 Light microscopy photographs of Bps. adriatica strain S21 germinated from resting cyst a. surface of ventral view showing sulcus and cingulum; b. deeper focus of ventral view showing sulcus and cingulum; c. dorsal view showing nucleus (N); d. deeper focus of dorsal view showing wide cingulum; e–f. ventral view of two planozygotes showing sulcus, cingulum, and two longitudinal flagella (arrows). The red asterisks indicate round refractive bodies. Scale bars=10 μm. |
|
| Fig.5 Scanning electron micrographs (SEM) of Bps. adriatica strain S21 germinated from resting cyst a, b. ventral view showing sulcus, cingulum, ventral ridge area (vr), a large vesicle at the upper end of the sulcus (two asterisks), an elongate vesicle (single asterisk) in contact with the elongate apical vesicle, transverse, and longitudinal flagella; c. dorsal view showing cingulum, transverse flagellum, and postcingular plates (pc); d. apical-lateral view showing apical vesicles and the elongate apical vesicle; e. antapical view showing latitudinal vesicle series on the hypocone; f. apical view showing knobs (yellow asterisks) on the single elongate apical vesicle, bordered on each side by an elongate vesicle (white arrowheads). Scale bars=5 μm. |
The vegetative cells of S. donghaienis was 12.8–20.2 μm long (average 16.4±2.1 μm; n=50) and 9.6–16.0 μm wide (average 12.9±1.5 μm; n=50). The epitheca was conical and longer than the hypotheca, and the hypotheca was rounded and bilobed (Figs. 6–7). The plate formula is Po, x, 4′, 3a, 7″, 6c, 6s, 5‴, 2 (Fig. 7). The cingulum was wide and deep (Fig. 7a–c). The upper part of sulcus was narrower than its lower part (Figs. 6a–b & 7a). The apical pore complex (APC) comprised a round apical pore plate and a long canal plate (Fig. 7a & e–f). The nucleus was rounded and located centrally (Fig. 6d). The yellowish-brown, and granulated chloroplasts were distributed peripherally (Fig. 6d–e).

|
| Fig.6 Light microscopy photographs of S. donghaienis strain S23 germinated from resting cyst a, b, c, f. the Calcofluor White stained cells showing plates; d. centrally located nucleus (N); e. epifluorescence image of vegetative cells showing numerous ellipsoid to elongated, rod-like or irregular shaped chloroplasts. Scale bars=10 μm. |

|
| Fig.7 Scanning electron micrographs (SEM) of S. donghaienis strain S23 germinated from resting cyst a. ventral view of epitheca, cingulum, sulcus, and hypotheca; b. dorsal view of epitheca, cingulum, and hypotheca; c. ventral-lateral view of epitheca, cingulum, and hypotheca; d. antapical view of hypotheca and posterior sulcal plates (s.p.); e. apical-lateral view of epitheca and cingulum; f. apical view of cover plate (cp), pore plate (Po), and canal plate (x). cp. cover plate; Po: pore plate; x: canal plate; s.a.: anterior sulcal plate; s.p., posterior sulcal plate. Scale bars=10 μm. |
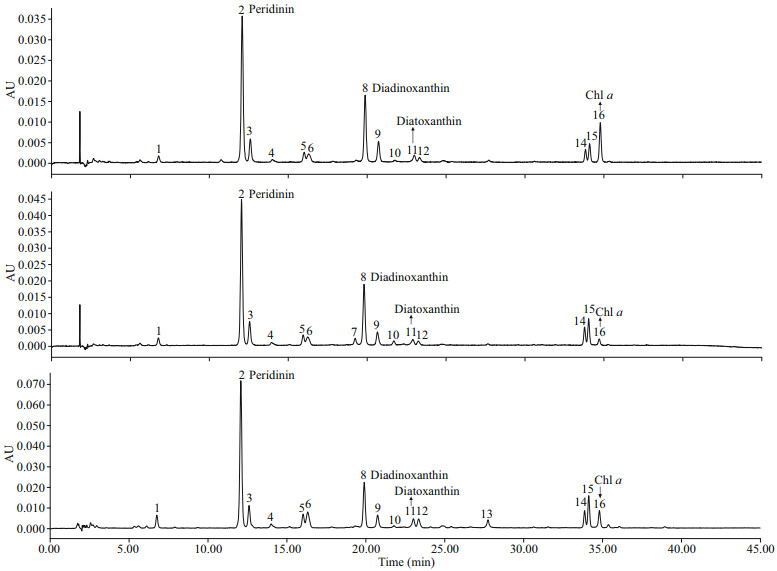
Based on available standards, four photosynthetic pigments were identified in B. brevisulcata (strain S1), Bps. adriatica (strain S21), and S. donghaienis (strain 23), including one kind of chlorophyll (Chl a) and three carotenoids (peridinin, diadinoxanthin, and diatoxanthin; Fig. 8). Chl-a contents of B. brevisulcata (strain S1), Bps. adriatica (strain S21), and S. donghaienis (S23) were 1.02, 0.25, and 10.89 pg/cell, respectively. Peridinin (3.51, 4.91, and 79.11 pg/ cell) was the most abundant carotenoid for the three species, then diadinoxanthin, and diatoxanthin. There were 10, 11, and 11 unidentified small peaks (either new pigments or known pigments but without standards) for B. brevisulcata strain S1, Bps. adriatica strain S21, and S. donghaienis strain S23.
|
| Fig.8 Absorption chromatograms (440 nm) of the pigment extracts of B. brevisulcata strain S1, Bps. adriatica strain S21, and S. donghaienis strain 23 from the East China Sea, China a. B. brevisulcata strain S1; b. Bps. adriatica strain S21; c. S. donghaienis strain 23. The peak numbers and corresponding pigments are as follows: 1, 3, 4, 5, 6, 7, 9, 10, 12, 13, 14, 15: unknown (either new pigments or known pigments but without standards); 2: peridinin; 8: diadinoxanthin; 11: diatoxanthin; 16: Chl a. |
Seven partial LSU rRNA gene sequences of B. brevisulcata (strains S1, S2, S3, S4, and S5;OL355144‒OL355148), Bps. adriatica (strain S21; OL355142), and S. donghaienis (strain 23; OL314541), and one ITS sequence of S. donghaienis (strain 23; OL314542) were obtained from the clonal cultures. One partial LSU rRNA gene sequence of Bps. adriatica (ON350794) was obtained using single-cell PCR sequencing for the cyst from surface sediment sample of S01-1.
The partial LSU rRNA gene sequence of B. brevisulcata strain S1 (1 374 bp; OL355144) was 99.93% (1 373 bp/1 374 bp) identical to the entity of the type material (AB858351), 99.88% (852 bp/853 bp) to 99.93% (1 359 bp/1 360 bp) identical to four entities (AB858352, AB858353, LC068842, and OL699922) deposited as B. brevisulcata, and 98.60% (845 bp/857 bp), 99.41%(509 bp/512 bp), 99.30% (853 bp/859 bp), and 99.32% (877 bp/883 bp) identical to B. pseudopalustris (syn. Woloszynskia pseudopalustris; AF260402), B. baltica (syn. Woloszynskia halophila sensu Kremp et al. (2005); AY628430), B. cincta (syn. Woloszynskia cincta; FJ024705), and B. tirezensis (LT601379).
The partial LSU rRNA gene sequence of Bps. adriatica strain S21 (1 425 bp; OL355142) was 99.69% (1 267 bp/1 271 bp) identical to the entity of the type material deposited at GenBank as Gymnodinium pygmaeum strain K-0968, 99.24% (1 172 bp/1 181 bp)to 99.79% (1 422 bp/1 425 bp) identical to 12 entities (AB858354‒AB858356, LC068843, LC413947‒ LC413950, LM992904‒LM992906, and OL691545) deposited as Bps. adriatica, 99.78% identical to 11 entities (KM603188‒KM603198) deposited as Bps. cf. adriatica, and 99.13% (794 bp/801 bp) to99.30% (1 415 bp/1 425 bp) identical to four entities (JN558103‒JN558105, KM603185) deposited as Protodinium simplex. Among all these entities inGenBank, Bps. adriatica also corresponded to the cultures established from vegetative cells isolated from the same area where the cultures from cysts were established (i.e., East China Sea; Luo et al., 2015).
The partial LSU rRNA gene sequence of S. donghaienis strain 23 (1 433 bp; OL314541) was 98.51% (796 bp/808 bp) to 99.51% (812 bp/816 bp) identical to 69 entities deposited as S. donghaienis in GenBank. The ITS sequence of S. donghaienis strain 23 (663 bp; OL314542) was 93.80% (454 bp/484 bp) to 99.64% (558 bp/560 bp) identical to five entities (AY685008, HQ729492, HQ729502, JN982374, and MG914024) deposited as S. donghaienis, and 99.28% (550 bp/554 bp) to 99.68% (617 bp/619 bp) identical to seven entities (AY499533, AY676151, AY67615, AY788357, and FJ823594‒FJ823596) deposited as Scrippsiella sp. in GenBank.
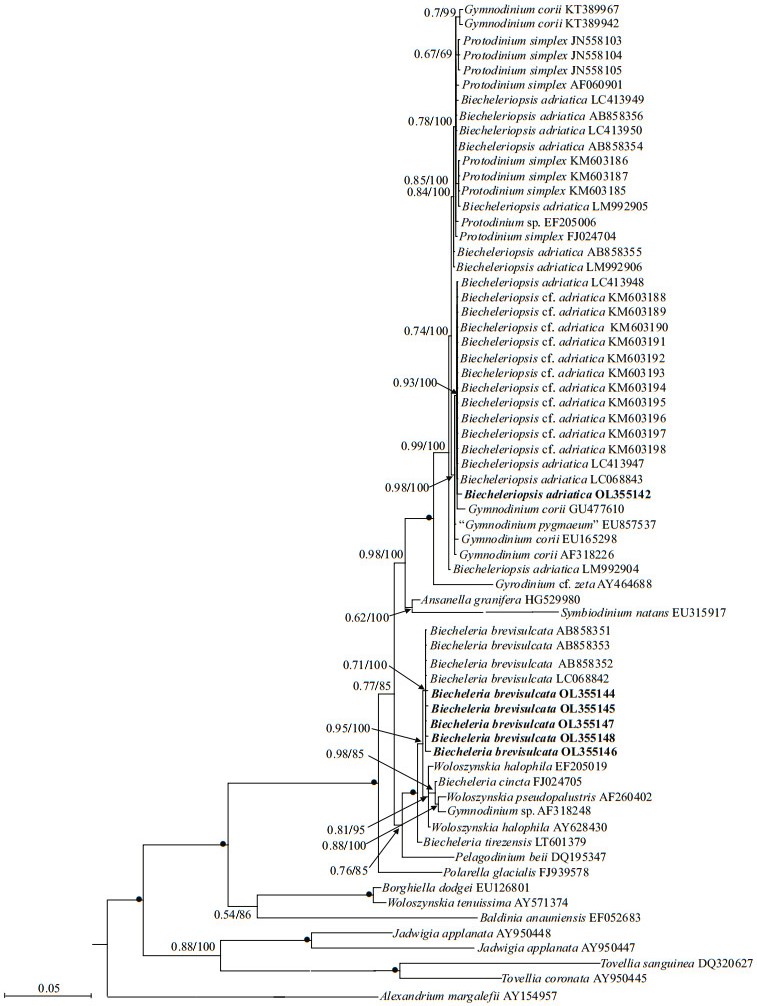
Phylogenetic analyses of B. brevisulcata, Bps. adriatica, and S. donghaienis using maximumlikelihood (ML) and Bayesian inference (BI) generated similar trees based on LSU rRNA gene and ITS sequences but differed at a few internal nodes (Figs. 9‒11). For B. brevisulcata, our sequences and other sequences (AB858351‒AB858351, LC068842) formed a coherent clade with strong support (0.71/100), which is sistering to the clade including Woloszynskia halophila (EF205019, AY628430), B. cincta (FJ024705), Woloszynskia pseudopalustris (AF260402), and Gymnodinium sp. (AY318248) with strong support (0.95/100; Fig. 9). For Bps. adriatica, our sequence and other sequences (LC068843, LC413947, and LC413948), Bps. cf. adriatica (KM603188‒KM603198), and G. corii (GU477610) formed a coherent clade withstrong support (0.93/100; Fig. 9), and formed a well-supported sister clade (0.98/100; Fig. 9) including the type material of Bps. adriatica ("G. pygmaeum", EU857537), and G. corii (EU165298, AF318226). For the phylogenetic analysis of S. donghaienis based on LSU rRNA gene sequences, our sequence (OL31454), 69 sequences deposited as S.donghaienis, and Scrippsiella sp. (AY685011) formed a coherent clade with maximal support (1/100; Fig. 10), and formed sister groups with other Scrippsiella species (Fig. 10). For the phylogenetic analysis of S. donghaienis based on ITS sequences, our sequence (OL314542), four sequences deposited as S. donghaienis (HQ729492, HQ729502, JN982374, and MG914024), and seven sequences deposited as Scrippsiella sp. (AY499533, AY676151, AY676155, AY788357, FJ823594‒FJ823596) formed a coherent clade with medium support (0.51/88; Fig. 11), and formed a sister group with one sequence deposited as S. donghaienis (AY685008) with maximal support (1/100; Fig. 11).
|
| Fig.9 Molecular phylogeny of B. brevisulcata, Bps. adriatica, and diverse assemblage of other dinoflagellates inferred from the partial LSU rRNA gene sequences based on Bayesian inference (BI) with A. margalefii (AY154957) as outgroup Species names and accession numbers in bold were obtained from our cultures. Branch lengths are drawn to the scale, with scale bar indicating the number of substitutions per site. Numbers on branches are statistical support values for clusters on the right (Bayesian posterior probability/ML bootstrap support). Bootstrap values > 50% and posterior probabilities > 0.5 are shown. Solid dots indicate maximum support (Bayesian posterior probability: 1.00/ML bootstrap support: 100). |

|
| Fig.10 Molecular phylogeny of S. donghaienis and diverse assemblage of other dinoflagellates inferred from the partial LSU rRNA gene sequences based on Bayesian inference (BI) with C. brodyi (DQ991374) as outgroup Species name and accession number in bold were obtained from our culture. Branch lengths are drawn to scale, with scale bar indicating the number of substitutions per site. Numbers on branches are statistical support values for clusters on the right (Bayesian posterior probability /ML bootstrap support). Bootstrap values > 50% and posterior probabilities > 0.5 are shown. Solid dots indicate maximum support (Bayesian posterior probability: 1.00/ML bootstrap support: 100). |

|
| Fig.11 Molecular phylogeny of S. donghaienis and diverse assemblage of other dinoflagellates inferred from the internal transcribed spacer (ITS) region sequences based on Bayesian inference (BI) with P. dalei (DQ991374) as outgroup Species name and accession number in bold were obtained from our culture. Branch lengths are drawn to scale, with scale bar indicating the number of substitutions per site. Numbers on branches are statistical support values for clusters on the right (Bayesian posterior probability / ML bootstrap support). Bootstrap values > 50% and posterior probabilities > 0.5 are shown. Solid dots indicate maximum support (Bayesian posterior probability: 1.00/ML bootstrap support: 100). |
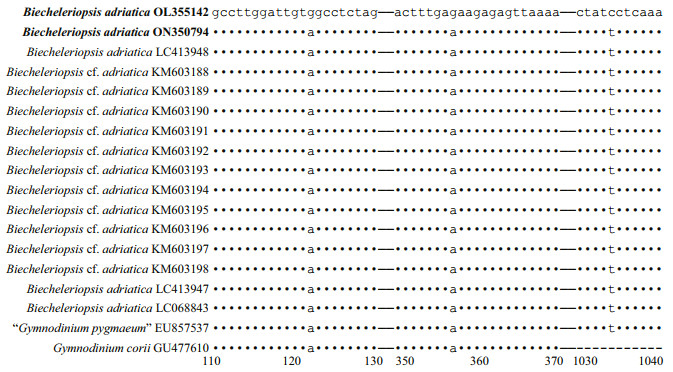
The pairwise distances computed using Jukes-Cantor model showed that the sequence divergence among the LSU rRNA gene sequences of Bps. adriatica from the present study, type material (deposited in GenBank as G. pygmaeum with the accession No. EU857537), and sequences deposited as Bps. adriatica (AB858354‒AB858356, LC068843, LC413947‒LC413950, and LM992904‒LM992906) ranged from 0.000 to 0.006, the distances between our sequence and Bps. cf. adriatica (KM603188‒ KM603198) were 0.002, and among our sequence, and the sequences deposited as P. simplex (AF060901), Gymnodinium sp. (EF205006), G. corii (AF318226, EU165298, GU477610, KT389967, KT389942), P. simplex (FJ024704, JN558103‒JN558105, KM603185‒KM603187) ranged from 0.000 to 0.007 (Supplementary Table S1). However, the distances between our sequence and B. brevisulcata (AB858351), Ansanella granifera (HG529980), and A. margalefii (AY154957) were 0.039, 0.029, and 0.244, respectively (Supplementary Table S1). Sequence differences among our strain (S21), the other stains from water samples or surface sediments formed a well-supported clade with ours (Fig. 9), and the one we found in the surface sediment sample in the same sea area were found mainly at three stable positions (3 bp/1 438 bp (D1–D6 regions); Fig. 12).
|
| Fig.12 Sequence variation in the LSU rDNA sequences among Bps. adriatica strain 21, the one cyst of this species found in the surface sediment sample in the same sea area, and other strains of Bps. adriatica formed a well-supported clade with ours in Fig. 9 |
The genetic distance based on LSU rRNA gene of B. brevisulcata obtained in the present study and previous works, and other related species were compared (Supplementary Table S2). The sequence divergence among the LSU rRNA gene sequences of B. brevisulcata obtained in our work (OL355144‒OL355148) and other strains previously deposited in GenBank (AB858351‒AB858353, LC068842) ranged within 0.000‒0.007 (Supplementary Table S2), the sequence divergence among B. brevisulcata (OL355144‒OL355148) and other Biecheleria species (B. baltica, B. cincta, B. pseudopalustris, and B. tirezensis) ranged 0.002‒0.015 (SupplementaryTable S2), but the sequence divergence among B. brevisulcata (OL355144‒OL355148) and otherdistant species Pelagodinium beii (DQ195347), Polarella glacialis (FJ939578), and A. margalefii (AY154957) were 0.055‒0.057, 0.077‒0.079, and 0.808‒0.875, respectively (Supplementary Table S2).
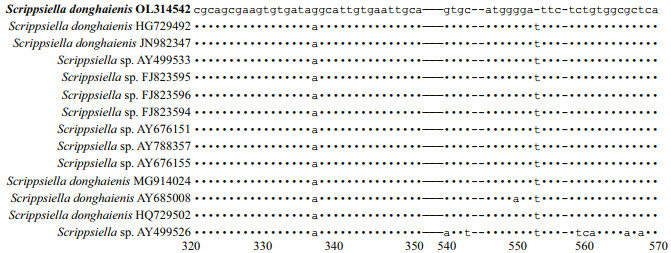
The genetic distance based on LSU rRNA gene and ITS of S. donghaienis obtained in the present study and previous work, and other related species were compared (Supplementary Tables S3 & S4). The sequence divergence among the LSU rRNA gene sequence of S. donghaienis obtained in our work (OL314541) andother strains previously deposited in GenBank ranged 0.000‒0.009 (Supplementary Table S3), 0.013‒0.139, and the sequence divergence among S. donghaienis (OL314541) and other Scrippsiella species (S. acuminate, S. erinaceus, S. sweeneyae, S. spinifera, S. plana, S. bicarinata, S. kirschiae, S. trifida, S. infula, S. rotunda, S. lachrymose, S. enormis, S. masanensis, S. precaria, and S. ramonii) ranged 0.002‒0.015(Supplementary Table S3). The pairwise distances computed using Jukes-Cantor model showed that the sequence divergence between the ITS sequences of S. donghaienis obtained in the present study, and sequences deposited as S. donghaienis (AY685008, HQ729492, HQ729502, JN982374, and MG914024) ranged from 0.000 to 0.015 (Supplementary Table S4), the sequence divergence between the ITS sequences of S. donghaienis obtained in the present study, and the other Scrippsiella species ranged 0.175–0.360 (Supplementary Table S4). Sequence differences between our strain (S23) and the other stains from water samples or surface sediments formed a well-supported clade with ours (Fig. 11) were found mainly at two stable positions (2 bp/599 bp; Fig. 13).

|
| Fig.13 Sequence variation in the ITS sequences between S. donghaienis strain 23 and the other strains formed a well clade with ours in Fig. 11 |
Biecheleriopsis adriatica was described by Moestrup et al. (2009) who differentiated it from the species of Biecheleria based on the presence of a nuclear connector and a 51-bases long fragment of D2 domain of LSU rRNA gene. Moestrup et al. (2009) also found resting cysts in the culture of Bps. adriatica, described as having ellipsoidal to ovoid shapes and numerous spines, a size of 7‒8-μm length and 5‒6-μm width. Benico et al. (2019) found resting cyst-like cells in unialgal culture being morphologically different from vegetative cell, and they called them as resting cyst-like cells, which were ~16 μm, spherical, covered by a transparent thick wall, and contained a red accumulation body. Kang et al. (2009), Wang et al. (2013), and Kang and Wang (2018) germinated Bps. adriatica from cyst assemblages (identified as G. corii in Kang et al. (2009) and Wang et al. (2013))from Southern Chinese coastal sediments, but did not observe the genuine resting cyst of this organism. Luo et al. (2015) also germinated Bps. cf. adriatica (very possible conspecific with our strain S21) from the sediment collected from the Yellow Sea and the South China Sea, but also without observation on the morphology of Bps. cf. adriatica cyst. The above-mentioned works on Bps. adriatica have confirmed that it could form resting cyst, but none of them reported the morphology of cyst from the field. In this work, we germinated Bps. adriatica from a single cyst collected from the East China Sea, and the species was identified with morphological and molecular characterization allowing us to confidently confirm the cyst-motile stage relationship of this small-sized species. Resting cyst of Bps. adriatica was spherical and sub-spherical, which is similar to the so-called "resting cyst-like cells" as observed by Benico et al. (2019), but different from the cyst from culture having an ellipsoidal to ovoid morphology (Moestrup et al., 2009). The diameter of Bps. adriatica in our work (~7.3 μm) is very close to the cyst (7‒8 μm long and 5‒6 μm wide) observed by Moestrup et al. (2009), but much smaller than the resting cyst-like cells (~16 μm) observed by Benico et al. (2019). The sieves with the pore size larger than 20 μm were routinely used to concentrate cysts, and then followed by cyst assemblage germination, the cysts with smaller size (< 20 μm) would be lost during processing, therefore, the efforts in discovering more small sized cysts should focus on the cyst assemblage being smaller than 20 μm.
4.2 Viability of dinoflagellates cysts stored in long-buried sedimentsIn the past, changes of species composition in sediments have been used to assess environmental changes including eutrophication, changes in salinity, or oxygen concentration (Dale et al., 1999; Ellegaard et al., 2013, 2020; Bringué et al., 2016; García-Moreiras et al., 2018; Li et al., 2021; Siano et al., 2021). The adaptive responses of dinoflagellate species might be influenced by anthropogenic activities or climate changes, and revealing the evolutionary processes in a species is very important to understanding the genetic structure of populations. However, the length of live resting cyst preserved in the sediment cores (e.g., decades to centuries) or the molecular information (rRNA genes or other gene markers) for the buried cysts or residual fragments have not been well explored before 2010. After that, the impacts of environmental changes on the physiology, genetic structure, and diversity in various dinoflagellate species (Ribeiro et al., 2011; Klouch et al., 2016; Lundholm et al., 2017; Kremp et al., 2018; Delebecq et al., 2020; Ellegaard et al., 2020; Girault et al., 2021) have been investigated. In this study, we successfully revived B. brevisulcata, Bps. adriatica, and S. donghaienis from a sediment coredated back to 1941±18 AD from the East China Sea, which indicates that these cysts are viable for at least 70 years. Klouch et al. (2016) detected molecular signal of S. donghaienis in a sediment core sample dated back to 1866±7 AD from the Bay of Brest, France, and successfully germinated this organism only from the layers of 2–17 cm corresponding to the years of 2010±1 AD to 1978±2 AD, much more recent than 1941±18 AD. In older sediments (70‒100 years), Protoceratium reticulatum, Lingulodinium polyedrum, and P. dalei were successfully germinated(Lundholm et al., 2011; Ribeiro et al., 2011). Girault et al. (2021) successfully germinated A. minutum and S. acuminata from the sediment corresponding to1947±11 AD. Kremp et al. (2018) found Apocalathium malmogiense is viable in the sediment layers of 106-year old. Recently, Delebecq et al. (2020) germinated A. minutum, Heterocapsa minima, Margalefidinium polykrikoides, Protoperidinium spp., and Diplopsalis group in sediments dated back to 117±21 years ago, and S. acuminata beyond 156±27 years ago. Our work increases the diversity of species (B. brevisulcata and Bps. adriatica) potentially revivable from more long-buried sediments, which will promote studies in the field of resurrection ecology.
4.3 Genetic diversity among dinoflagellates revived from cysts stored in long-buried sediments and current conditionGenetic diversity is an important aspect of biodiversity, which is defined as measurements that determine the changes of genetic variability within any level of a taxon but more often within a species or even population, e.g., allelic diversity or richness, mutational diversity, and effective population size (Hughes et al., 2008; Ebenezer et al., 2012; Ellegren and Galtier, 2016). Previous phylogeographic studies have revealed high levels of genetic diversity in a number of dinoflagellate species, e.g., A. fundyense (Erdner et al., 2011), Amphidinium spp. (Murray et al., 2012), Gambierdiscus spp. (Nishimura et al., 2013), Margalefidinium fulvescens (Lin et al., 2020), Ostreopsis spp. (Lee and Park, 2020), and Pseudocochlodinium profundisulcus (Hu et al., 2021).However, all above-mentioned works examined the genetic diversity among contemporary populations or even within a population rather than that between historical and contemporary populations as done in the present work, although the history here is less than a century (~70 years). From our phylogenetic tree, our five strains of B. brevisulcata branched together with the other strains of this species, but their evolution distance varied. Our strain of Bps. adriatica and three strains of Bps. adriatica, eleven strains of Bps. cf. adriatica, and one strain of G. corii formeda coherent clade, and a sister group with the type material of this species, but differed at three stable positions. Our strain of S. donghaienis, the other strains of this species, and one strain of Scrippsiella sp. formed a coherent clade. Within this clade, there were four branches, our strain grouped together with those historical and contemporary populations. We also compared the rRNA gene based genetic distances of the seven strains of B. brevisulcata, Bps. adriatica, and S. donghaienis revived from a long-buried sediment dated back to 1941±18 AD and those from water samples and surface sediments (the contemporary age), and found the genetic distance between our strain of B. brevisulcata and other strains range from 0.002 to 0.006, but among other strains, the genetic distances are 0.000‒0.005, most 0.001, and similar trends were also found in Bps. adriatica, and S. donghaienis. We also found differences in three stable positions of LSU rRNA gene sequences between the population of Bps. adriatica we found 70 years ago and the contemporary or present-day population (including the cyst of this species we found in the surface sediment and vegetative cells found by Luo et al. (2015) in the same sea area), and ITS sequences in two stable positions between the population of S. donghaienis we found 70 years ago and the contemporary or present-day population (including the cyst of this species found by Gu et al. (2008) in the surface sediment in the same sea area). It seemed these current populations are genetically different from those that existed in the area 70 years ago, which suggests that there has been a shift in the populations of Bps. adriatica and S. donghaienis. Due to a limited size of dataset for the present work, the possible historical succession of populations of Bps. adriatica, S. donghaienis and even other speciesin the area requires a more intensive and extensive investigation, as these possible shifts may be highly indicative of the environmental changes and anthropological activities that occurred in the area during the past.
5 DATA AVAILABILITY STATEMENTAll data generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
6 ACKNOWLEDGMENTWe are highly grateful of the two anonymous reviewers for their patience, critical comments, and generous suggestions, which helped greatly the improvement of the manuscript. Sediment samples' collections were supported by NSFC Open Research Cruise, funded by Shiptime Sharing Project of NSFC. We appreciate for the cruises conducted by R/V Xiang Yang Hong 18 and staff from First Institute ofOceanography, Ministry of Natural Resources, China.
Electronic supplementary material
Supplementary material (Supplementary Tables S1–S4) is available in the online version of this article at https://doi.org/10.1007/s00343-022-2122-1.
Anderson D M, Wall D. 1978. Potential importance of benthic cysts of Gonyaulax tamarensis and G. excavata in initiating toxic dinoflagellate blooms. Journal of Phycology, 14(2): 224-234.
DOI:10.1111/j.1529-8817.1978.tb02452.x |
Benico G A, Takahashi K, Lum W M, et al. 2019. First report of Biecheleriopsis adriatica in Bolinao, northwestern Philippines and its wide distribution in Southeast Asia and adjacent waters. Philippine Journal of Natural Sciences, 24: 34-41.
|
Bolch C J S. 1997. The use of sodium polytungstate for the separation and concentration of living dinoflagellate cysts from marine sediments. Phycologia, 36(6): 472-478.
DOI:10.2216/i0031-8884-36-6-472.1 |
Boutrup P V, Moestrup Ø, Tillmann U, et al. 2017. Ultrastructure and phylogeny of Kirithra asteri gen. et sp. nov. (Ceratoperidiniaceae, Dinophyceae) -a freeliving, thin-walled marine photosynthetic dinoflagellate from Argentina. Protist, 168(5): 586-611.
DOI:10.1016/j.protis.2017.08.001 |
Bravo I, Figueroa R I. 2014. Towards an ecological understanding of dinoflagellate cyst functions. Microorganisms, 2(1): 11-32.
DOI:10.3390/microorganisms2010011 |
Bringué M, Pospelova V, Calvert S E, et al. 2016. High resolution dinoflagellate cyst record of environmental change in Effingham Inlet (British Columbia, Canada) over the last millennium. Palaeogeography, Palaeoclimatology, Palaeoecology, 441: 787-810.
DOI:10.1016/j.palaeo.2015.10.026 |
Dai X F, Lu D D, Xia P, et al. 2012. A 50-year temporal record of dinoflagellate cysts in sediments from the Changjiang Estuary, East China Sea, in relation to climate and catchment changes. Estuarine, Coastal and Shelf Science, 112: 192-197.
DOI:10.1016/j.ecss.2012.07.016 |
Dale B, Thorsen T A, Fjellsa A. 1999. Dinoflagellate cysts as indicators of cultural eutrophication in the Oslofjord, Norway. Estuarine, Coastal and Shelf Science, 48(3): 371-382.
DOI:10.1006/ecss.1999.0427 |
Dale B. 2001. The sedimentary record of dinoflagellate cysts: looking back into the future of phytoplankton blooms. Scientia Marina, 65(S2): 257-272.
DOI:10.3989/scimar.2001.65s2257 |
Daugbjerg N, Hansen G, Larsen J, et al. 2000. Phylogeny of some of the major genera of dinoflagellates based on ultrastructure and partial LSU rDNA sequence data, including the erection of three new genera of unarmoured dinoflagellates. Phycologia, 39(4): 302-317.
DOI:10.2216/i0031-8884-39-4-302.1 |
de Freitas A D S, Escamilla J H, Barreto C F, et al. 2020. Dinocysts as a tool for palaeoenvironmental reconstruction in Vitória Bay, Brazil. Radiocarbon, 62(2): 289-311.
DOI:10.1017/RDC.2020.4 |
Delebecq G, Schmidt S, Ehrhold A, et al. 2020. Revival of ancient marine dinoflagellates using molecular biostimulation. Journal of Phycology, 56(4): 1077-1089.
DOI:10.1111/jpy.13010 |
Ebenezer V, Medlin L K, Ki J S. 2012. Molecular detection, quantification, and diversity evaluation of microalgae. Marine Biotechnology, 14(2): 129-142.
DOI:10.1007/s10126-011-9427-y |
Ellegaard M, Clokie M R J, Czypionka T, et al. 2020. Dead or alive: sediment DNA archives as tools for tracking aquatic evolution and adaptation. Communications Biology, 3(1): 169.
DOI:10.1038/s42003-020-0899-z |
Ellegaard M, Ribeiro S, Lundholm N et al. 2013. Using the sediment archive of living dinoflagellate cysts and other protist resting stages to study temporal population dynamics. In: Lewis J M, Marret F, Bradley L R eds. Biological and Geological Perspectives of Dinoflagellates. Geological Society, London. p. 149-153.
|
Ellegaard M, Ribeiro S. 2018. The long-term persistence of phytoplankton resting stages in aquatic 'seed banks'. Biological Reviews, 93(1): 166-183.
DOI:10.1111/brv.12338 |
Ellegren H, Galtier N. 2016. Determinants of genetic diversity. Nature Reviews Genetics, 17(7): 422-433.
DOI:10.1038/nrg.2016.58 |
Erdner D L, Richlen M, McCauley L A R, et al. 2011. Diversity and dynamics of a widespread bloom of the toxic dinoflagellate Alexandrium fundyense. PLoS One, 6(7): e22965.
DOI:10.1371/journal.pone.0022965 |
Figueroa R I, Estrada M, Garcés E. 2018. Life histories of microalgal species causing harmful blooms: haploids, diploids and the relevance of benthic stages. Harmful Algae, 73: 44-57.
DOI:10.1016/j.hal.2018.01.006 |
Fritz L, Triemer R E. 1985. A rapid simple technique utilizing calcofluor white M2R for the visualization of dinoflagellate thecal plates. Journal of Phycology, 21(4): 662-664.
|
García-Moreiras I, Pospelova V, García-Gil S, et al. 2018. Climatic and anthropogenic impacts on the Ría de Vigo(NW Iberia) over the last two centuries: a high-resolution dinoflagellate cyst sedimentary record. Palaeogeography, Palaeoclimatology, Palaeoecology, 504: 201-218.
DOI:10.1016/j.palaeo.2018.05.032 |
Girault M, Siano R, Labry C, et al. 2021. Variable inter and intraspecies alkaline phosphatase activity within single cells of revived dinoflagellates. The ISME Journal, 15(7): 2057-2069.
DOI:10.1038/s41396-021-00904-2 |
Gómez F, López-García P, Takayama H, et al. 2015. Balechina and the new genus Cucumeridinium gen. nov.(Dinophyceae), unarmored dinoflagellates with thick cell coverings. Journal of Phycology, 51(6): 1088-1105.
DOI:10.1111/jpy.12346 |
Gómez F. 2012. A checklist and classification of living dinoflagellates (Dinoflagellata, Alveolata). CICIMAR Oceánides, 27(1): 65-140.
DOI:10.37543/oceanides.v27i1.111 |
Gu H F, Sun J, Kooistra W H C F, et al. 2008. Phylogenetic position and morphology of thecae and cysts of Scrippsiella (Dinophyceae) species in the East China Sea. Journal of Phycology, 44(2): 478-494.
DOI:10.1111/j.1529-8817.2008.00478.x |
Gu H, Mertens K N, Derrien A, et al. 2022. Unravelling the Gonyaulax baltica species complex: cysttheca relationship of Impagidinium variaseptum, Spiniferites pseudodelicatus sp. nov. and S. ristingensis (Gonyaulacaceae, Dinophyceae), with descriptions of Gonyaulax bohaiensis sp. nov, G. amoyensis sp. nov. and G. portimonensis sp. nov. Journal of Phycology, 58(3): 465-486.
DOI:10.1111/jpy.13245 |
Guillard R R L. 1975. Culture of phytoplankton for feeding marine invertebrates. In: Proceedings of the 1st Conference on Culture of Marine Invertebrate Animals Greenport. Springer, New York. p. 29-60.
|
Hall T A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41: 95-98.
|
Head M J. 1996. Modern dinoflagellate cysts and their biological affnities. In: Jansonius J, McGregor D C eds. Palynology: Principles and Applications. American Association of Stratigraphic Palynologists Foundation, Dallas. p. 1197-1248.
|
Hu Z X, Deng Y Y, Luo Z H, et al. 2020a. Characterization of the unarmored dinoflagellate Pseliodinium pirum (Ceratoperidiniaceae) from Jiaozhou Bay, China. Phycological Research, 68(1): 3-13.
DOI:10.1111/pre.12385 |
Hu Z X, Li Z, Deng Y Y, et al. 2020b. Morphology, ultrastructure, and molecular phylogeny of the unarmoured dinoflagellate Kirithra sigma sp. nov. (Ceratoperidiniaceae, Dinophyceae). Phycologia, 59(5): 385-396.
DOI:10.1080/00318884.2020.1771660 |
Hu Z X, Liu Y Y, Deng Y Y, et al. 2022. The notorious harmful algal blooms-forming dinoflagellate Prorocentrum donghaiense produces sexual resting cysts, which widely distribute along the coastal marine sediment of China. Frontiers in Marine Science, 9: 826736.
DOI:10.3389/fmars.2022.826736 |
Hu Z X, Xu N, Gu H F, et al. 2021. Morpho-molecular description of a new HAB species, Pseudocochlodinium profundisulcus gen. et sp. nov., and its LSU rRNA genebased genetic diversity and geographical distribution. Harmful Algae, 108: 102098.
DOI:10.1016/j.hal.2021.102098 |
Hughes A R, Inouye B D, Johnson M T J, et al. 2008. Ecological consequences of genetic diversity. Ecology Letters, 11(6): 609-623.
DOI:10.1111/j.1461-0248.2008.01179.x |
Kang W, Wang Z H, Fu Y H, et al. 2009. Investigation on germination of phytoplankton resting cells in sediment traps collected from Daya Bay, South China Sea. China Environmental Science, 29(12): 1285-1290.
|
Kang W, Wang Z H. 2018. Identification of a marine woloszynskioid dinoflagellate Biecheleriopsis adriatica and germination of its cysts from southern Chinese coasts. Journal of Environmental Sciences, 66: 246-254.
DOI:10.1016/j.jes.2017.04.031 |
Katoh K, Misawa K, Kuma K I, et al. 2002. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research, 30(14): 3059-3066.
DOI:10.1093/nar/gkf436 |
Keafer B A, Buesseler K O, Anderson D M. 1992. Burial of living dinoflagellate cysts in estuarine and nearshore sediments. Marine Micropaleontology, 20(2): 147-161.
DOI:10.1016/0377-8398(92)90004-4 |
Kim S Y, Roh Y H, Shin H H, et al. 2018. Decadal-scale variations of sedimentary dinoflagellate cyst records from the Yellow Sea over the last 400 years. Estuarine, Coastal and Shelf Science, 200: 91-98.
DOI:10.1016/j.ecss.2017.10.006 |
Klouch K Z, Schmidt S, Andrieux-Loyer F, et al. 2016. Historical records from dated sediment cores reveal the multidecadal dynamic of the toxic dinoflagellate Alexandrium minutum in the Bay of Brest (France). FEMS Microbiology Ecology, 92(7): fiw101.
DOI:10.1093/femsec/fiw101 |
Kong F Z, Yu R C, Zhang Q C, et al. 2012. Pigment characterization for the 2011 bloom in Qinhuangdao implicated "brown tide" events in China. Chinese Journal of Oceanology and Limnology, 30(3): 361-370.
DOI:10.1007/s00343-012-1239-z |
Kremp A, Elbrächter M, Schweikert M, et al. 2005. Woloszynskia halophila (Biecheler) comb. nov.: abloom-forming cold-water dinoflagellate co-occurring with Scrippsiella hangoei (Dinophyceae) in the Baltic Sea. Journal of Phycology, 41(3): 629-642.
DOI:10.1111/j.1529-8817.2005.00070.x |
Kremp A, Hinners J, Klais R, et al. 2018. Patterns of vertical cyst distribution and survival in 100-year-old sediment archives of three spring dinoflagellate species from the northern Baltic Sea. European Journal of Phycology, 53(2): 135-145.
DOI:10.1080/09670262.2017.1386330 |
Kumar S, Stecher G, Li M, et al. 2018. MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6): 1547-1549.
DOI:10.1093/molbev/msy096 |
Lee B, Park M G. 2020. Distribution and genetic diversity of the toxic benthic dinoflagellate genus Ostreopsis in Korea. Harmful Algae, 96: 101820.
DOI:10.1016/j.hal.2020.101820 |
Li L, Wang Y J, Liu D Y. 2021. Phytoplankton shifts in the central Bohai Sea over the last 250 years reflect eutrophication and input from the Yellow River. Ecological Indicators, 126: 107676.
DOI:10.1016/j.ecolind.2021.107676 |
Limoges A, Van Nieuwenhove N, Head M J, et al. 2020. A review of rare and less well known extant marine organic-walled dinoflagellate cyst taxa of the orders Gonyaulacales and Suessiales from the northern Hemisphere. Marine Micropaleontology, 159: 101801.
DOI:10.1016/j.marmicro.2019.101801 |
Lin S H, Hu Z X, Deng Y Y, et al. 2020. An assessment on the intrapopulational and intraindividual genetic diversity in LSU rDNA in the harmful algal blooms-forming dinoflagellate Margalefidinium (=Cochlodinium) fulvescens based on clonal cultures and bloom samplesfrom Jiaozhou Bay. China. Harmful Algae, 96: 101821.
DOI:10.1016/j.hal.2020.101821 |
Liu Y Y, Deng Y Y, Shang L X, et al. 2021. Geographic distribution and historical presence of the resting cysts of Karenia mikimotoi in the seas of China. Harmful Algae, 109: 102121.
DOI:10.1016/j.hal.2021.102121 |
Liu Y Y, Hu Z X, Deng Y Y, et al. 2020a. Evidence for production of sexual resting cysts by the toxic dinoflagellate Karenia mikimotoi in clonal cultures and marine sediments. Journal of Phycology, 56(1): 121-134.
DOI:10.1111/jpy.12925 |
Liu Y Y, Hu Z X, Deng Y Y, et al. 2020b. Evidence for resting cyst production in the cosmopolitan toxic dinoflagellate Karlodinium veneficum and the cyst distribution in the China seas. Harmful Algae, 93: 101788.
DOI:10.1016/j.hal.2020.101788 |
Lundholm N, Ribeiro S, Andersen T J, et al. 2011. Buried alive-germination of up to a century-old marine protist resting stages. Phycologia, 50(6): 629-640.
DOI:10.2216/11-16.1 |
Lundholm N, Ribeiro S, Godhe A, et al. 2017. Exploring the impact of multidecadal environmental changes on the population genetic structure of a marine primary producer. Ecology and Evolution, 7(9): 3132-3142.
DOI:10.1002/ece3.2906 |
Luo Z H, Hu Z X, Tang Y Z, et al. 2018. Morphology, ultrastructure, and molecular phylogeny of Wangodinium sinense gen. et sp. nov. (Gymnodiniales, Dinophyceae)and revisiting of Gymnodinium dorsalisulcum and Gymnodinium impudicum. Journal of Phycology, 54(5): 744-761.
DOI:10.1111/jpy.12780 |
Luo Z H, Yang W D, Xu B, et al. 2015. Morphology, ultrastructure, and phylogeny of Protodinium simplex and Biecheleriopsis cf. adriatica (Dinophyceae) from the China Sea. Nova Hedwigia, 101(1-2): 251-268.
DOI:10.1127/nova_hedwigia/2015/0268 |
Mertens K N, Gu H F, Gurdebeke P R, et al. 2020. A review of rare, poorly known, and morphologically problematic extant marine organic-walled dinoflagellate cyst taxa of the orders Gymnodiniales and Peridiniales from the northern Hemisphere. Marine Micropaleontology, 159: 101773.
DOI:10.1016/j.marmicro.2019.101773 |
Moestrup Ø, Lindberg K, Daugbjerg N. 2009. Studies on woloszynskioid dinoflagellates V. ultrastructure of Biecheleriopsis gen. nov., with description of Biecheleriopsis adriatica sp. nov. Phycological Research, 57(3): 221-237.
DOI:10.1111/j.1440-1835.2009.00541.x |
Murray S A, Garby T, Hoppenrath M, et al. 2012. Genetic diversity, morphological uniformity and polyketide production in dinoflagellates (Amphidinium, Dinoflagellata). PLoS One, 7(6): e38253.
DOI:10.1371/journal.pone.0038253 |
Nishimura T, Sato S, Tawong W, et al. 2013. Genetic diversity and distribution of the ciguatera-causing dinoflagellate Gambierdiscus spp. (Dinophyceae) in coastal areas of Japan. PLoS One, 8(4): e60882.
DOI:10.1371/journal.pone.0060882 |
Ok J H, Jeong H J, Lee S Y, et al. 2021. Shimiella gen. nov. and Shimiella gracilenta sp. nov. (Dinophyceae, Kareniaceae), a kleptoplastidic dinoflagellate from Korean waters and its survival under starvation. Journal of Phycology, 57(1): 70-91.
DOI:10.1111/jpy.13067 |
Posada D. 2008. jModelTest: phylogenetic model averaging. Molecular Biology and Evolution, 25(7): 1253-1256.
DOI:10.1093/molbev/msn083 |
Price A M, Baustian M M, Turner R E, et al. 2018. Dinoflagellate cysts track eutrophication in the northern Gulf of Mexico. Estuaries and Coasts, 41(5): 1322-1336.
DOI:10.1007/s12237-017-0351-x |
Ribeiro S, Berge T, Lundholm N, et al. 2011. Phytoplankton growth after a century of dormancy illuminates past resilience to catastrophic darkness. Nature Communications, 2: 311.
DOI:10.1038/ncomms1314 |
Ronquist F, Huelsenbeck J P. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19(12): 1572-1574.
DOI:10.1093/bioinformatics/btg180 |
Scholin C A, Herzog M, Sogin M, et al. 1994. Identification of group- and strain-specific genetic markers for globally distributed Alexandrium (Dinophyceae) II. Sequence analysis of a fragment of the LSU rRNA gene. Journal of Phycology, 30(6): 999-1011.
DOI:10.1111/j.0022-3646.1994.00999.x |
Shang L X, Hu Z X, Deng Y Y, et al. 2019. Metagenomic sequencing identifies highly diverse assemblages of dinoflagellate cysts in sediments from ships' ballast tanks. Microorganisms, 7(8): 250.
DOI:10.3390/microorganisms7080250 |
Siano R, Lassudrie M, Cuzin P, et al. 2021. Sediment archives reveal irreversible shifts in plankton communities after World War II and agricultural pollution. Current Biology, 31(12): 2682-2689.e7.
DOI:10.1016/j.cub.2021.03.079 |
Silvestro D, Michalak I. 2012. raxmlGUI: a graphical frontend for RAxML. Organisms Diversity & Evolution, 12(4): 335-337.
DOI:10.1007/s13127-011-0056-0 |
Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30(9): 1312-1313.
DOI:10.1093/bioinformatics/btu033 |
Steidinger K A, Meave del Castillo M E. 2018. Guide to the Identification of Harmful Microalgae in the Gulf of Mexico, Volume I: Taxonomy. Florida Fish and Wildlife Conservation Commission, Fish and Wildlife Research Institute, St. Petersburg, p384.
|
Takahashi K, Benico G, Lum W M, et al. 2019. Gertia stigmatica gen. et sp. nov. (Kareniaceae, Dinophyceae), a new marine unarmored dinoflagellate possessing the peridinin-type chloroplast with an eyespot. Protist, 170(5): 125680.
DOI:10.1016/j.protis.2019.125680 |
Takahashi K, Moestrup Ø, Jordan R W, et al. 2015. Two new freshwater woloszynskioids Asulcocephalium miricentonis gen. et sp. nov. and Leiocephalium pseudosanguineum gen. et sp. nov. (Suessiaceae, Dinophyceae) lacking an apical furrow apparatus. Protist, 166(6): 638-658.
DOI:10.1016/j.protis.2015.10.003 |
Takahashi K, Moestrup Ø, Wada M, et al. 2017. Dactylodinium pterobelotum gen. et sp. nov., a new marine woloszynskioid dinoflagellate positioned between the two families Borghiellaceae and Suessiaceae. Journal of Phycology, 53(6): 1223-1240.
DOI:10.1111/jpy.12575 |
Tamura K, Nei M, Kumar S. 2004. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences of the United States of America, 101(30): 11030-11035.
DOI:10.1073/pnas.0404206101 |
Tang Y Z, Gu H F, Wang Z F, et al. 2021. Exploration of resting cysts (stages) and their relevance for possibly HABscausing species in China. Harmful Algae, 107: 102050.
DOI:10.1016/j.hal.2021.102050 |
Tang Y Z, Hu Z X, Deng Y Y. 2016. Characteristical life history (resting cyst) provides a mechanism for recurrence and geographic expansion of harmful algal blooms of dinoflagellates: a review. Studia Marina Sinica, 51: 132-154.
(in Chinese with English abstract) DOI:10.12036/hykxjk20160730001 |
Van Nieuwenhove N, Head M J, Limoges A, et al. 2020. An overview and brief description of common marine organic-walled dinoflagellate cyst taxa occurring in surface sediments of the northern Hemisphere. Marine Micropaleontology, 159: 101814.
DOI:10.1016/j.marmicro.2019.101814 |
Wang Z H, Fu Y H, Kang W, et al. 2013. Germination of phytoplankton resting cells from surface sediments in two areas of the Southern Chinese coastal waters. Marine Ecology, 34(2): 218-232.
DOI:10.1111/maec.12009 |
White T J, Bruns T, Lee S et al. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M A, Gelfand D H, Sninsky J J et al eds. PCR Protocols. Elsevier, Amsterdam. p. 315-322, https://doi.org/10.1016/B978-0-12-372180-8.50042-1.
|
 2022, Vol. 40
2022, Vol. 40


