Institute of Oceanology, Chinese Academy of Sciences
Article Information
- XU Wei, NIU Jie, GAN Wenyu, GOU Siyu, ZHANG Shuai, QIU Han, JIANG Tianjiu
- Identification of paralytic shellfish toxin-producingmicroalgae using machine learning and deep learning methods
- Journal of Oceanology and Limnology, 40(6): 2202-2217
- http://dx.doi.org/10.1007/s00343-022-1312-1
Article History
- Received Oct. 1, 2021
- accepted in principle Nov. 28, 2021
- accepted for publication Jan. 11, 2022
2 Atmospheric Sciences and Global Change Division, Pacific Northwest National Laboratory, Richland 99354, WA, USA;
3 Key Laboratory of Eutrophication and Red Tide Prevention of Harmful Algae and Marine Biology, Jinan University, Guangzhou 510632, China
Harmful algal blooms (HABs) occurred frequently in the coastal areas of China in recent years. These blooms were harmful and resulted in massive fish mortality and greatly affected the sustainable development of marine ecosystems, aquatic wildlife resources, marine aquaculture, and even human health (Ignatiades and Gotsis-Skretas, 2010; Anderson et al., 2012a; O'Neil et al., 2012; Cusick and Sayler, 2013; Paerl et al., 2016). Studies have found that human poisoning syndromes are caused by ingestion of algal toxins through ingestion of contaminated seafood, direct exposure to algal toxins contaminated water, or inhalation of aerosol toxins (Van Dolah, 2001; Rasmussen et al., 2016; Wang et al., 2020). To date, the number of diseases and deaths caused by HABs pollution is increasing, especially in the Pacific Rim countries (Kumar and Sharma, 2021). Three types of red tide are toxic red tide, nontoxic red tide, and fish toxic red tide, respectively (Hallegraeff, 1993; Masó and Garcés, 2006). Paralytic shellfish poisoning (PSP) is one of the most dangerous biotoxins due to its wide distribution and strong toxicity (Andrinolo et al., 1999). Some algae that produce PSP have been identified, such as Gonyaulax, Alexandrium, and Gymnodinium (Sommer et al., 1937, 1948). Studies have shown that A. tamarense, A. catenella, and A. minutum are the main algae producing PSP, which belong to the Alexandrium genus (Kotaki, 1983; Yang and Li, 1998; Anderson et al., 2012b). The occurrence of toxic algal bloom would cause water quality deterioration, destroy the ecological balance of the water environment, restrict economic development, and even endanger human health. The identification of PSP microalgae is of great significance for the prevention and control of red tide hazards.
There are plenty of methods to classify PSP microalgae. At present, fluorescence spectroscopy (FSSS), high-performance liquid chromatography (HPLC), rRNA probes, and fluorescence in situ hybridization (FISH) have attracted wide attention as analytical techniques for distinguishing phytoplankton taxa groups due to its rapid, alternative, and sensitive characteristics (Jeffrey and Hallegraeff, 1980; Seppälä and Olli, 2008; Alexander et al., 2012; Tang et al., 2012). However, studies indicated that identification based on morphological characteristics alone is not sufficient for on-site monitoring of toxic microalgae (Tang et al., 2012). Moreover, some methods can identify only one species of algae at a time, but multiple algae rapidly, hence effective methods are required. Currently, three-dimensional (3D) fluorescence, also known as the excitation-emission matrix (EEM) or EEM diagram, as a common measure used in identifying HABs, can be completely obtained by varying the excitation wavelength (EX) and the emission wavelength (EM) concurrently (Lee et al., 1995; Millie et al., 1997; Beutler et al., 2002; Divya and Mishra, 2007). EEM FSSS combined with parallel factor (PARAFAC), principal component analysis (PCA), and wavelet decomposition (WD) using "db7" and "coif" wavelets are universal methods for quantitative and qualitative analysis of algae (Moberg et al., 2001; Lü et al., 2011; Duan et al., 2012; Cao et al., 2015). However, some steps in these analytical methods are complicated to operate, and the prediction accuracy of traditional forecasting methods such as empirical predicting, dynamic forecasting, and statistical forecasting is defective (Ma et al., 2007). Therefore, some new methods are required to simplify operations and enhance accuracy.
Image classification is a major application in the field of machine learning (ML), which is widely used in many fields, such as computer vision and network image retrieval (Aas and Eikvil, 1999; Sebastiani, 2002). With the advance of computer technology, ML has also been applied in environmental pollution control and biological hazard prevention (Alizadeh et al., 2018; Zhang, 2018a; Xu and Jackson, 2019). ML algorithms are based on statistical principles that dig hidden information in data sets and make calculable statistical predictions for unknown data without relying on predetermined model equations (Mosavi et al., 2019). k-nearest neighbor (k-NN) rule and support vector machines (SVM), as two types of ML models, have recently gained prominence in the field of precise classification. For example, Hingane et al. (2015) classified images of brain tumors using SVM and k-NN; Alimjan et al. (2018) also combined SVM and k-NN to distinguish remote sensing images; Zhuang et al. (2020) used deep k-NN for medical image classification, etc. Deep learning (DL) is also a type of machine learning that teaches computers to do what comes naturally to humans: learn from experience. Deep learning is especially suited for image recognition, which is significantly successful in solving problems such as facial recognition, motion detection, and many advanced driver assistance technologies (LeCun et al., 2015). In a taxonomic context, deep convolutional neural networks (CNN or ConvNet) are currently becoming the state-of-the-art technique in image classification. Convolutional neural network is a special branch of deep learning, which has been applied to a large number of research fields (Wang et al., 2018; Ozawa et al., 2020; Raphael et al., 2020; Zavala-Mondragon et al., 2020) and can recognize visual patterns hierarchically through learning (Sultana et al., 2019). LeCun et al. (1989a) presented the practical model of CNN and contrived LeNet-5 (LeCun et al., 1989b, 1998). The backpropagation algorithm was applied to train LeNet-5 to recognize visual patterns from the original image directly without using any separate feature engineering mechanism (LeCun, 1992). Presently, this algorithm has been developed quite maturely. These methods, however, have not been applied widely in the classification of algae.
Generally, ML and DL have shown higher classification accuracy, faster running speed, and simpler data preprocessing. At present, there are few studies on the horizontal classification of various algae genera, and the classification accuracy is not very well. In addition, the characteristic bands extracted from PSP microalgae and non-PSP microalgae by 3D fluorescence spectrum are not clear. Hence, the purposes of this study are to classify the algae at the genus level, and by combining SVM, k-NN, and CNN with the 3D FSSS, to develop a simple, rapid, and efficient 3D fluorescence method for classifying PSP and non-PSP microalgae, to contribute to the early warning of Marine toxic red tide outbreak. Another purpose of this study is to find the characteristic fluorescent bands of each microalga by using the spectral information extracted by CNN. The rest of this paper is organized as follows. Section 2 describes the experimental and data processing methods and models building process. Section 3 describes (i) the effects of two data processing methods, (ii) the identification accuracy comparisons of the models, and (iii) the differences in characteristic spectra between PSP and non-PSP microalgae. Then, in Section 4, the results and limitations of the current approach are discussed and outline the potential future studies. Finally, Section 5 summarizes the key findings of this research.
2 MATERIAL AND METHOD 2.1 Instrument and reagentThe fluorescence spectrophotometer used in this study is Hitachi F4600 made in Japan. The Rotary Evaporator is from Jiapeng Technology Co., Ltd., Shanghai, China. The high-speed centrifuge is Thermo electron corporation CR3i multifunction made in the USA. The microscope is XSZ-D2 OLYMPUS/CX31 from Precision Instrument Co., Ltd., Shanghai, China. Light Climate Box is from Carboniferous Instruments, Ningbo, China. Ultrasonic Cell Disruptor is Branson Sonifieal 540 made in the USA.
2.2 Microalgae culture experimentsThree strains of PSP microalgae, i.e., Alexandrium minutum (AMSY), Alexandrium pacificum (APQD), Gymnodinium catenatum (GCFC), and eighteen strains of non-PSP microalgae, i.e., Alexandrium tamarense (ATCZ), Amphidinium carterae (ACCZ), Prorocentrum donghaiense (PDCZ), Prorocentrum lima (PLCZ), Karenia mikimotoi (KMCZ), Dunaliella salina (DSCZ), Platymonas subcordiformis (PSCZ), Isochrysis galbana (IGCZ), Dictyocha speculum (DSCF), Chattonella marina (CMHK), Chattonella ovata (COHK), Heterosigma akashiwo (HACZ), Prymnesium parvum (PPCZ), Chaetoceros debilis (CDCZ), Skeletonema costatum (SCCZ), Thalassiosira weissflogii (TWCZ), Nitzschia closterium (NCCZ), Chlorella vulgaris (CVCZ) were selected in this study.
Natural seawater (pH is 7.5±0.1, and salinity is 28) was used to culture these microalgae. The microalgae were sterilized for about 25 min, with environmental conditions of 121 ℃ and 103.42 kPa, and then cooled to room temperature and filtered using 0.45-μm microporous membrane afterward. An algae culture medium with f/2 modified formula (Guillard and Ryther, 1962) was used in this work. Microalgae were cultivated in an artificial climate box that controls temperature (25 ℃), light intensity (150 μmol/(m2·s)), and light and dark cycle (L D=12 h 12 h) after inoculation (Guillard and Ryther, 1962). Each group consists of three parallel samples and would be applied to train and test the model, respectively.
2.3 Extraction of paralytic shellfish poisonFreezing and thawing the collected algal fluid, the microalgae cells were broken by an ultrasonic crusher (50% power, 2 s on, and 2 s off for 15 min). Then the seamollient with a crushing rate above 95% was examined. The supernatant was taken after centrifugation at 4 ℃ in an ultrafiltration centrifuge tube at 10 000×g working about 10 min and repeated the above steps (Jiang et al., 2011). Finally, centrifugal ultrafiltration fluid was preserved in a refrigerator at -20 ℃ to analyze toxicity with HPLC by centrifugation (4 ℃, 7 200×g, 10 min) using the ultrafiltration centrifuge tubes with 10 000 Dalton filter membrane aperture. The EX and the EM of 3D fluorescence spectrum were set to 400–600 nm and 650–750 nm, respectively. The appropriate concentration of microalgae cells was detected by 3D fluorescence spectroscopy and the corresponding spectral information was obtained by scanning. The data were saved in ASCII format for further analysis.
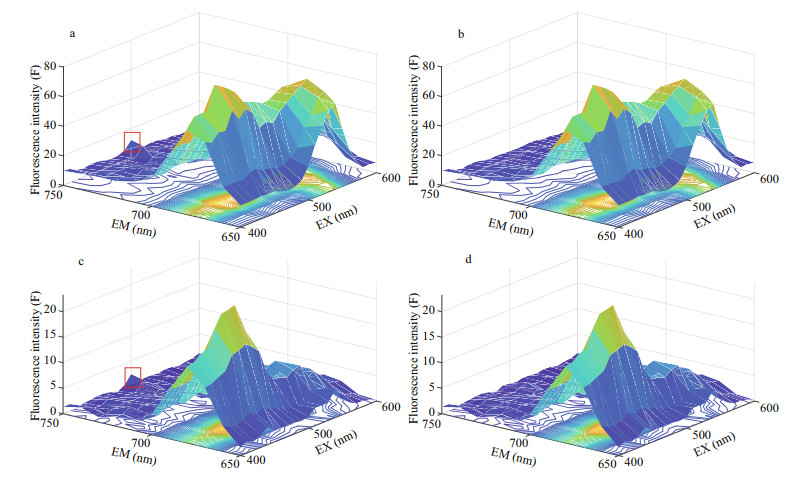
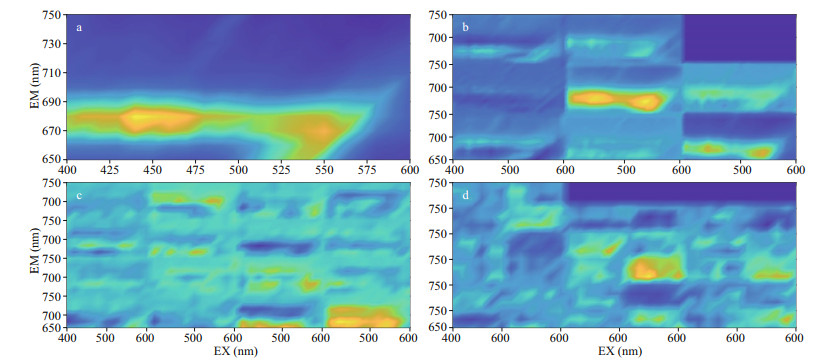
2.4 Preprocessing of 3D FSSS data 2.4.1 Removing scatteringTo avoid the influence of Rayleigh scattering and constituents of dissolved organic matter (CDOM), the excitation wavelength of 400–600 nm was selected in this study. In the original spectrogram of algae used in the experiment (Fig. 1a & c), the presence of scattering peaks caused by test tubes or instruments would affect the true fluorescence information of algae and should be removed. The original spectrogram of 18 species of non-PSP microalgae and 3 species of PSP microalgae are shown in Appendix. By comparing the scattered band data with the normal band data, the confidence interval of the normal data is obtained through the probability distribution, and the outliers are marked and interpolated by the Delaunay triangle interpolation (Zepp et al., 2004; Bahram et al., 2006).The method for removing the scattering peak (RS) has been described in Appendix. Data normalization transforms all fluorescence intensities to be between 0 and 1 as follows:
 (1)
(1)
|
| Fig.1 The original 3D fluorescence spectra of paralytic shellfish poison (PSP) (AMSY, (a) and (b)) and non-PSP microalgae (PPCZ, (c) and (d)) (a) and (c) represent microalgae spectra that have scattering peaks, and (b) and (d) represent those that scattering peaks have been removed. |
where X* denotes the normalized intensity values of the FSSS, Xn is the fluorescence intensity of each frequency point, Xmax is the maximum, and Xmin is the minimum fluorescence intensity.
2.4.2 Removing background noiseWavelet decomposition is the most commonly used method for signal denoising. The principle of wavelet analysis denoising is to decompose the signal into low and high frequency parts and extract the useful information according to the selected threshold (Merry, 2005). The Coiflets (Coif 2) wavelet was used to deal with the background noise in this study. The basic principle of feature extraction of 3D FSSS of algae for scale component and wavelet component is as follows:
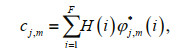 (2)
(2) (3)
(3)where H(i) (i=1, 2, ∙∙∙, n) is the EX or EM fluorescence spectrum of algae; i is the point of measurement; φj, m(i) and ψj, m(i) represent orthogonal scaling basis and wavelet basis, respectively; * stands for conjugate, and scale expansion (j) and scale function (m) shift with i. The scale space and wavelet space are orthogonal such that:
 (4)
(4)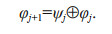 (5)
(5)Seventy percent of the whole data set (817 samples of non-PSP microalgae and 817 samples of PSP microalgae) were randomly selected for training the model, and the remaining data were used to verify the trained models and the verification results were reported as test accuracy. For k-NN and SVM, 5 k-fold cross-validation was used to verify the accuracy of the models during training. The PSP and non-PSP microalgae from the validation data set were also used to test the prediction accuracy of the model.
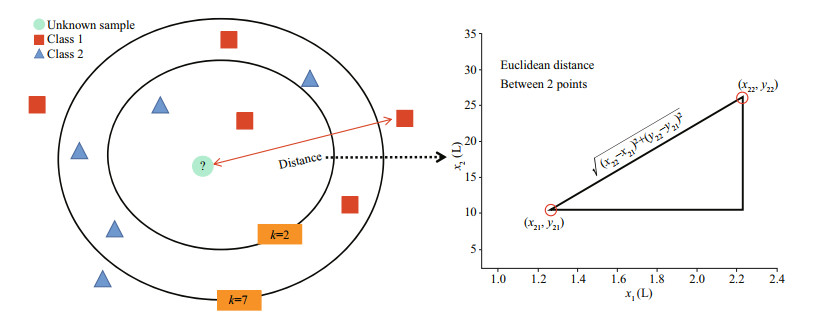
2.5.2 k-NN methodThe k-nearest neighbor (k-NN) model selects "k" known samples that are closest to the unknown sample by calculating the distance between the known and the unknown samples. Then, according to the principle of majority-voting, the unknown samples are classified into the same category as those in the most adjacent k samples, to determine the category of the unknown samples (Cover, 1968). Euclidean distance is generally used to judge the similarity of two samples in k-NN (Fig. 2).
|
| Fig.2 Principal diagram of k-nearest neighbor rule and Euclidean distance k represents the k samples closest to the unknown sample. |
The original data were saved in the matrix form, which needs to be transformed into a 1D vector. The PSP microalgae were labeled as "1" and the non-PSP microalgae as "0". This method is named KNN-1. The wavelet characteristic spectrum is the projection on the wavelet space of the original FSSS signal of the phytoplankton pigment extracted. The projection on each space is the characteristic segment of the wavelet spectrum, which represents the spectral information of the original signal. Therefore, the fourth scale component of the coif wavelet function was chosen to combine with k-NN, which is named KNN-2 and has the same steps as KNN-1 to distinguish the characteristic spectrum of algal toxicity.
2.5.3 SVM methodClassification using SVM is achieved by implementing linear or nonlinear separation surfaces in the input space (Fig. 3). Attractive for many classification applications, SVM attempts to find an optimal hyperplane that maximizes the boundary between classes by using a small number of training cases (Cherkassky, 1997; Vishwanathan and Narasimha Murty, 2002). It is based on statistical learning theory (Cherkassky, 1997) and seeks the optimal hyperplane in the high-dimensional space as a decision function (Boser et al., 1992). This may be illustrated with the training data set comprising k cases and be represented by {xi, yi}, i=1, 2, 3, ∙∙∙, k, where x RN is an N-dimensional space and y {-1, +1} is the class label. These training patterns are linearly separable if there exists a vector w (determining the orientation of a discriminating plane) and a scalar b (determining the offset of the discriminating plane from the origin) such that
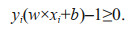 (6)
(6)
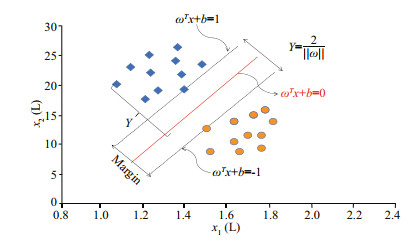
|
| Fig.3 Principal diagram of support vector machines x represents an N-dimensional space, y represents the class label, ω represents vector, b represents scalar, T stands for transpose. |
The hypothesis space can be defined by
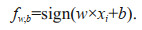 (7)
(7)The SVM finds the separating hyperplanes for which the distance among the classes measured along a line perpendicular to the hyperplane, is maximized. This can be achieved by solving the following constrained optimization problem:
 (8)
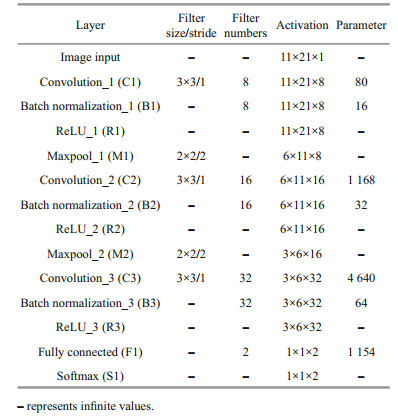
(8)The whole process of establishing the algal toxicity discriminating model by CNN is described in detail as follows. In this study, a database containing 1 144 images for training and 490 images for testing has been created. The CNN model applied in this study contains 14 weighted layers, including 3 convolutional layers, 3 batch normalization layers, 3 relu layers, 2 max pooling layers, 1 fully connected layer, 1 softmax layer, and 1 output layer (Table 1 and Fig. 4). The "sigmoid" function was used as the activation function to perform the non-linear transform before the pooling operation. The output layer used Euclidean Radial Basis Function units (RBF) (Buhmann, 2000) to classify the microalgae.
|
|
| Fig.4 Visualization of the activations of a convolutional neural network a. the input layer; b–d. the first (b), second (c), and third (d) convolution layers, respectively. |
The classification accuracy (%) (ACC) is the ratio between the number of correctly predicted samples (NP) and the total number of predicted samples (NT).
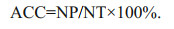 (9)
(9)The statistical analyses of the correlation among different variables were performed using IBM SPSS Statistics 21. The influences of scattering peaks and background noise removal on model results were determined by ANOVA. In all tests, the differences and correlations were statistically significant as P < 0.05.
3 RESULTAll the predictions in this work are computed using a personal computer. The CPU is i5-4200H, the graphics card is NVIDIA GTX 950M and the RAM is 12 G. The original size of input data is 11×21. The scale of the dataset is so small that the computation costs of training and running the models are almost negligible.
3.1 Results of removal and non-removal scattering peaksScattering peak affects the extraction of characteristic spectrum in the traditional algal toxicity discrimination. However, ML and DL can learn the characteristics of samples automatically, which means that the removal of the scattering peak may not be necessary for ML and DL. Thus, the results using the inputs with and without scattering peak removed (RS and NRS) were compared.
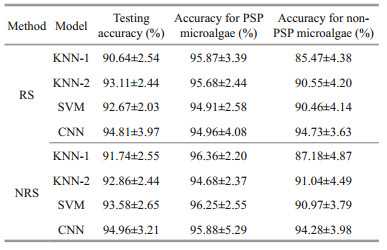
The results in Table 2 show the testing accuracy of the four models for each of the microalgae. The results with NRS using KNN-1 (91.74%), SVM (93.58%), and CNN (94.96%) are slightly better than those with RS (90.64%, 92.67%, and 94.81% using KNN-1, SVM, and CNN, respectively) while the accuracy of RS using KNN-2 (93.11%) is slightly higher than that of NRS (92.86%). In general, there was no significant difference in classification accuracy between RS and NRS cases. Therefore, this work chose the data without RS peak removed to train the model in the following experiments.
|
In this work, WD is chosen to remove background noise (RBN). For the k-NN model, there are two kinds of methods, which are KNN-1 and KNN-2. For the KNN-1, the raw data was converted into 1D vectors, while for the KNN-2, the 1D vectors were further processed by the fourth dimension of "coif 2" WD and then were applied to predict by k-NN. The samples were randomly selected from the whole dataset. 70% of algal data was used to train the models and the rest 30% was used to evaluate the model performance. Since the sample for each model training was randomly selected, the accuracy was the averaged value of the prediction after multiple training sessions.
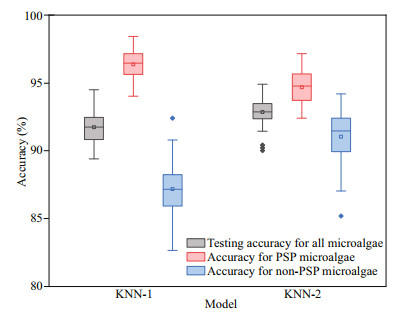
Figure 5 shows the classification results and accuracies of the k-NN models. The overall test accuracy of KNN-2 (92.86%) is higher than that of KNN-1 (91.74%). The accuracy for PSP microalgae with KNN-1 is 96.36%, which is higher than that with KNN-2 (94.68%). While the accuracy for non-PSP microalgae with KNN-1 (87.18%) is lower than that with KNN-2 (91.04%). Generally, the test accuracy of KNN-1 and KNN-2 are not significantly different. Hence, the preprocessing with RBN has little influence on the accuracy of PSP microalgae classification using the k-NN method.
|
| Fig.5 Model performances of the k-nearest neighbor method with and without wavelet decomposition The middle lines in each box represent the mean values, the top and bottom ones are for the maximum and minimum values, the little squares are the median values, the diamonds are the outliers, and the top and bottom of the rectangles represent 75% and 25% quantile, respectively. |
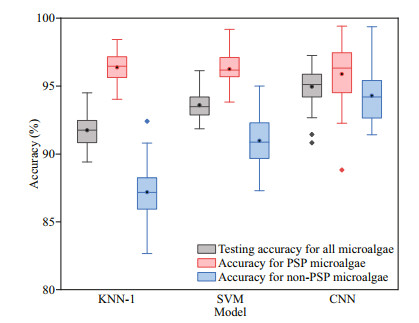
It has been shown in Section 3.2 that WD has no significant influence on the results of verification accuracy. Therefore, in this section, only results of KNN-1, SVM, and CNN are compared as shown in Fig. 6. All models were trained more than 40 times to verify the stability. The mean testing accuracies are above 91.74%, and the identification accuracies of PSP microalgae are higher than those of non-PSP. Comparing the results of the three models, the average test accuracy from high to low is CNN (94.96%), SVM (93.58%), and KNN-1 (91.74%), respectively. The range of verification accuracy of SVM is the smallest, which indicates that SVM is relatively stable in PSP and non-PSP microalgae classification.
|
| Fig.6 Comparisons of model performances of k-nearest neighbor (KNN-1), support vector machine (SVM), and convolutional neural network (CNN) The middle lines in each box represent the mean values, the top and bottom ones are for the maximum and minimum values, the little squares are the median values, the diamonds are the outliers, and the top and bottom of the rectangles represent 75% and 25% quantile, respectively. |
The accuracy of the three models can sometimes reach above 99.0% and the mean values are 96.36% (KNN-1), 96.25% (SVM), and 95.88% (CNN), respectively for identifying PSP microalgae. When identifying non-PSP microalgae, CNN does the best with the mean value of 94.28%. KNN-1 shows poor predictive performance in identifying non-PSP microalgae, which is just the opposite of the case of the PSP microalgae. In general, CNN performs better than the other two models regarding both accuracy and stability.
3.3.2 Prediction result of all modelsThe 690 samples of PSP microalgae that were not used for training were predicted. The accuracies are above 91.43% in the three models. Comparing the results of all models, the accuracy of KNN-1 (96.09%) is better than CNN (93.95%) and SVM (91.43%). And these results are consistent with the PSP microalgae testing results in Section 3.3.1.
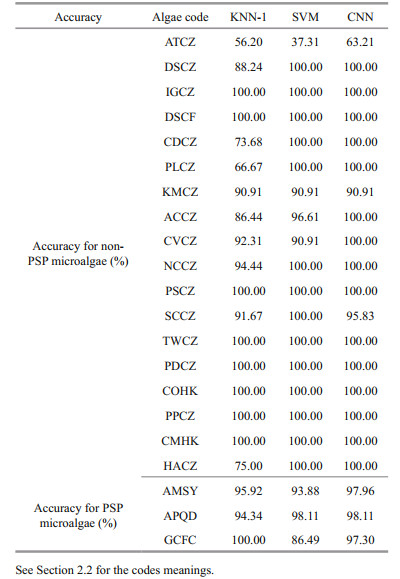
3.4 Result of single species of microalgae 3.4.1 Result of single species of microalgae in the optimum conditionsPSP microalgae with the highest concentration (104 cells/mL) and stronger fluorescence activity are selected as samples. The concentration of non-PSP microalgae is the same. Each microalgae species is selected to be cultured under the optimum conditions described in Section 2.2. The discrimination results of the three models for each species of microalgae are shown in Table 3.
|
For non-PSP microalgae, the accuracies of IGCZ, DSCF, PSCZ, TWCZ, PDCZ, COHK, PPCZ, and CMHK of the three models are all 100%. For DSCZ, CDCZ, PLCZ, NCCZ, and HACZ, the accuracies of CNN and SVM are also 100%, while the accuracies of KNN-1 are 88.24%, 73.68%, 66.67%, 94.44%, and 75.00%, respectively. Moreover, only CNN is 100% correct for the test of ACCZ and CVCZ, while KNN-1 is 86.44% and 92.31% and SVM is 96.61% and 90.91% accurate for the same cases. The discrimination accuracy of ATCZ is the lowest of the three models, i.e., KNN-1 is 56.20%, SVM is 37.31%, and CNN is 63.21%. In general, the identification accuracies of CNN for non-PSP microalgae are higher than those of SVM and KNN-1.
For PSP microalgae, the accuracies of CNN in the identification of AMSY, APQD, and GCFC are above 97%. The accuracy of KNN-1 for GCFC is 100%, while the accuracy of SVM is 86.49%. In addition, the average discriminant accuracy for APQD of the three models is higher than those of the other two microalgae species. Overall, the three models are effective in the identification of PSP microalgae, which is also consistent with the results in Section 3.3.
3.4.2 Result of PSP microalgae at different concentrationsThe experimental microalgae samples of AMSY, APQD, and GCFC are concentrated as 102, 103, and 104 cells/mL, respectively. The training set of the models includes the mixed data of all algae species with different concentrations with labeled categories (the training targets). Training with mixed data could also improve the robustness, adaptability, and generalization capability of the model. The results are shown in Fig. 7. The accuracies of the three models for 102-cells/mL microalgae samples are low, especially for GCFC. The accuracy of CNN is 59.57%, SVM is 70.21%, and KNN-1 is 87.23%, for classifying GCFC at 102 cells/mL. The accuracies of the three models are the highest for microalgae at 103 cells/mL. In addition, the accuracies of CNN for the three microalgae species are above 99% when the concentration is 103 cells/mL. The accuracies for microalgae samples at 104 cells/mL of the three models are slightly lower than those at 103 cells/mL, but all the accuracies are higher than 95% except for GCFC by SVM (86.49%). Furthermore, the classification accuracies of GCFC for all concentrations of SVM are generally lower than that of the other two models.

|
| Fig.7 Results of k-nearest neighbor, support vector machine, and convolutional neural network models for predicting 3 species of the paralytic shellfish poisoning alga at 102-cells/mL, 103-cells/mL, and 104-cells/mL concentration, respectively The light blue columns represent the prediction accuracy for the A. catenella microalgae, the dark blue ones for the A. pacificum microalgae, and the light green ones for the G. catenatum microalgae. |
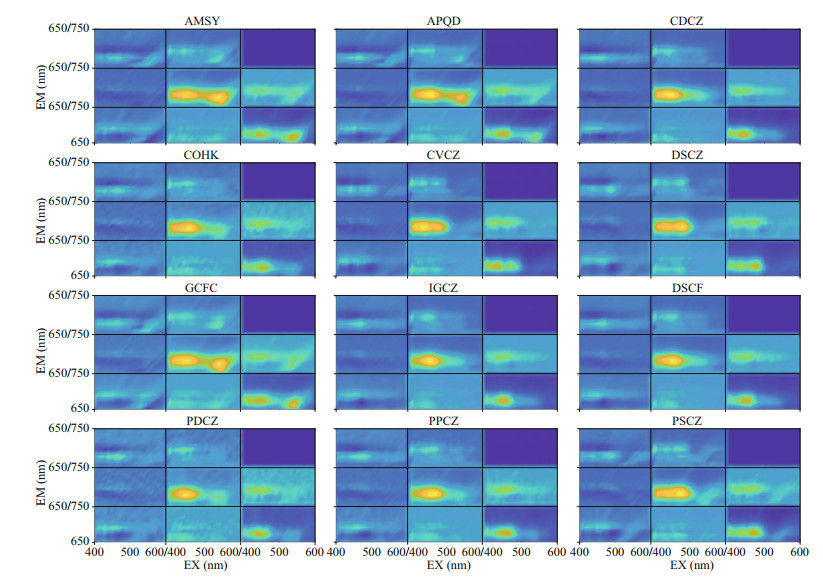
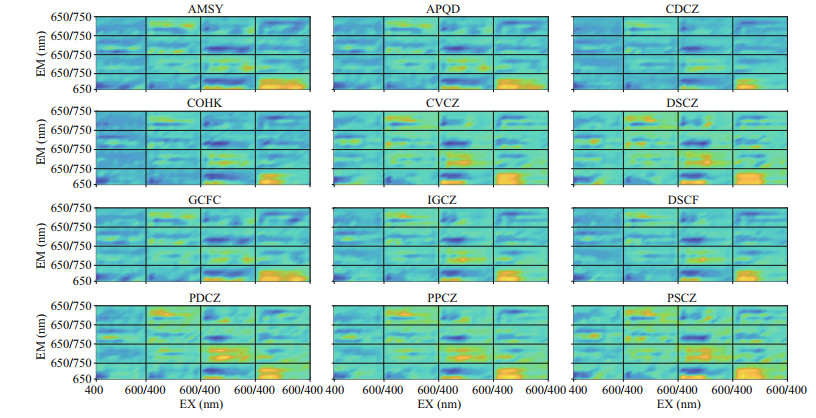
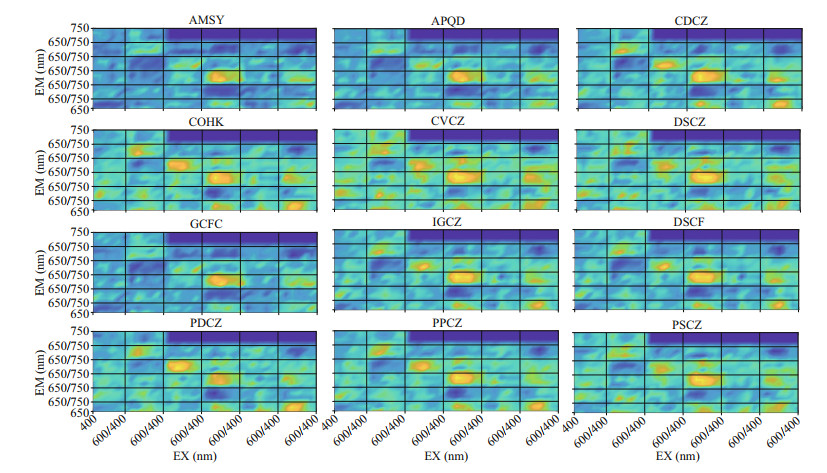
Figures 8–10 show the visualization of the kernels of three convolution layers of PSP microalgae and non-PSP microalgae, respectively. Some differences have been found by comparing images of the convolution kernel for PSP and non-PSP microalgae. In the first layer (Fig. 8), PSP microalgae have specific spectrums when the EX is 500–560 nm and the EM is 650–690 nm. In the second layer (Fig. 9), PSP microalgae have specific spectrums when the EX is 500–580 nm and the EM is 650–690 nm. In addition, non-PSP microalgae have specific spectrums when the EX is 440–520 nm and the EM is 690–700 and 720–730 nm. In the third layer (Fig. 10), non-PSP microalgae have specific spectrums when the EX is 410–480 nm and the EM is 650–720 nm.
|
| Fig.8 Visualization of the first convolutional layer of the convolutional neural network for the paralytic and non-paralytic shellfish toxigenic microalgae AMSY, APQD, and GCFC stand for A. catenella, A. pacificum, and G. catenatum, respectively, which are the paralytic shellfish poisoning microalgae. |
|
| Fig.9 Visualization of the second convolutional layer of the convolutional neural network for the paralytic and non-paralytic shellfish toxigenic microalgae AMSY, APQD, and GCFC stand for A. catenella, A. pacificum, and G. catenatum, respectively, which are the paralytic shellfish poisoning microalgae. |
|
| Fig.10 Visualization of the third convolutional layer of the convolutional neural network for the paralytic and non-paralytic shellfish toxigenic microalgae AMSY, APQD, and GCFC stand for A. catenella, A. pacificum, and G. catenatum, respectively, which are for paralytic shellfish poisoning microalgae. |
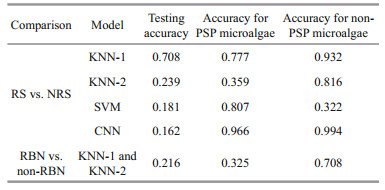
The effect of the RS peak on the accuracy of discrimination has been studied in this paper. Pearson correlation results show that there is no significant correlation between RS peak and NRS peak in each model. The RS peak of each sample has the same size and position. The EX of RS of each sample is 510– 560 nm, and the EM is 650–680 nm. Because ML and DL can automatically learn the features of samples so that the same scattering features may not be extracted. Comparing the Pearson correlation results of RS and NRS in Table 4, the P values of testing accuracy, PSP accuracy, and non-PSP accuracy are all greater than 0.05, which proves that removing the scattering peaks have no significant effect on the classification results of the models. Thus, the accuracies of the test of the four models are not significantly different whether removing the RS peak or not. Therefore, there is no need to consider the effect of the RS peak when DL and ML are used to classify PSP microalgae.
|
Comparing the Pearson correlation results of KNN-1 and KNN-2 in Table 4, the P values of testing accuracy, PSP accuracy, and non-PSP accuracy are all greater than 0.05, which proves that background noise has no significant effect on the classification using k-NN. While the dataset is processed by WD, some main features may be erased which can make the training accuracy deteriorate. Thus, there is no need to consider the effect of the background noise when DL and ML are used to classify PSP microalgae.
4.2 The application performance of all modelsThe mean accuracies of the three models are shown in Fig. 6. In general, the results of CNN are better than those of the other two, and SVM has higher accuracy than KNN-1. The principal rule of k-NN is searching the nearest neighbor or the k-nearest neighbor for classification. The classification principle of SVM is to construct a supercritical plane to classify samples in infinite-dimensional space. However, CNN can classify images by extracting image features for multiple learning and training directly (Sultana et al., 2019). This illustrates that CNN can learn more discriminative feature representation to enhance classification accuracy. KNN-1 has slightly higher accuracy than CNN in PSP recognition but has lower accuracy in non-PSP recognition. Overall, DL shows good performance in algal toxicity classification, especially with large data samples. But, KNN-1 and CNN algorithms can be used to distinguish red tide algae in practical application to improve the accuracy of early warning.
4.3 The application performance of single species of microalgaeThe reason that the discrimination accuracy of the three models for ATCZ is low (Table 3) is ATCZ and APQD belong to the same genus, which can cause ATCZ and APQD to have similar spectra features. Because ML and DL still have some deficiencies in similar image recognition, further optimization of parameters is needed to solve such problems in future research. In addition, the number of samples of APQD used in model training is also much larger than that of ATCZ, which may lead to the failure to extract some features of ATCZ and easy to produce differences for classification. Thus, the sample size of ATCZ needs to be increased for training in future studies to determine whether this guess is correct. Moreover, the accuracies of KNN-1 for CDCZ, PLCZ, and HACZ are poor, which also indicates that KNN-1 needs to be further optimized in the discrimination of non-PSP microalgae. It must be pointed out that the models are only capable of classifying these 21 microalgae species since they were only trained using the data for them. In order to classify the different species of microalgae, the spectral data of the specific microalgae should be provided to train the model. With the increase of training data, the model performance could be further improved. The variety of data will be increased for training to classify more microalgae species in future work.
The experimental results show that when the concentration of microalgae is 102 cells/mL, the toxic spectral information of GCFC detected by the 3D fluorescence spectrum is not sufficient. This may result in less extraction of toxic features for training the models. Therefore, further optimization of the model is needed to improve the discrimination effect of microalgae at low concentrations to enhance the practicability of models.
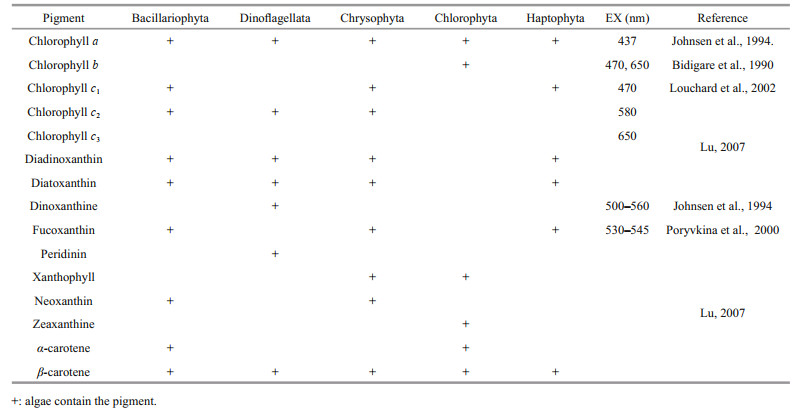
4.4 Difference in CNN between PSP and non-PSP microalgaeAlgal pigments include photosynthetic pigments and auxiliary photosynthetic pigments. The compound pigment is chlorophyll, which is directly involved in photosynthesis (Cai et al., 2006), and the auxiliary pigments are composed of carotenoids and phycobilins. The absorbed light can only be used for photosynthesis if it is first transmitted to chlorophyll a. The main pigment species and fluorescence wavelengths in each microalgae cell are shown in Table 5. Studies have shown that the synthesis of PSP may be closely related to photosynthesis (Taroncher-Oldenburg et al., 1997, 1999; Kellmann et al., 2008). Glutamy1-tRNA (gltX) regulatory gene which participates in chlorophyll synthesis cooperates with the glutamate N-acetyltransferase/amino acid N-acetyltransferase (argJ) gene which produces the paralytic toxin precursor arginine to synthesize paralytic toxin and chlorophyll (Yang et al., 2010; Jaeckisch et al., 2011; Zhang et al., 2018b). At present, nine proteins whose functions are known are associated with toxin production, and 6 of them have functions related to photosynthesis (Han et al., 2021). Therefore, it is feasible to identify toxin-producing algae by their fluorescence spectra.
|
The dinoxanthine, which is the characteristic pigment of Dinoflagellata, has a characteristic spectrum of 500–560 nm. The different bands found in Fig. 9 are also between 500–560 nm, which may be caused by dinoxanthine. In addition, chlorophyll concentration will affect the fluorescence intensity, resulting in spectral differences. Toxin synthesis in Dinoflagellata is accompanied by an increase in chlorophyll concentration (Zhang et al., 2018b), increasing the fluorescence intensity of the spectrum of PSP microalgae. With the deepening of the convolution layer, the fluorescence spectral characteristics are further extracted, making the fluorescence intensity of non-PSP microalgae at 410–480 nm higher than that of PSP microalgae. Differences in chlorophyll c1 in each algae species also lead to this result. As shown in Table 5, there is no chlorophyll c1 in Dinoflagellata, while Chrysophyta, Haptophyta, and Bacillariophyta all contain chlorophyll c1, and the absorption peak of chlorophyll c1 is at 470 nm. Therefore, this may lead to differences in the fluorescence spectra of microalgae. In addition, the different types of carotene and carotenoid in each algae cell also affect the different bands of algae. The absorption band of carotene is located between 425–500 nm and carotenoids have strong absorption at about 500–550 nm, which may also cause the difference in fluorescence spectrum at 410–520 nm.
Comparing our results with previous studies (Zhang, 2008; Huan et al., 2013; Qi et al., 2016), the accuracy of classifying the toxic microalgae has been enhanced by using ML and DL. In addition, there are few studies on the classification of microalgae at the species level using ML and DL. Therefore, this work shed a light on developing a new method to help classify PSP and non-PSP microalgae rapidly and accurately.
5 CONCLUSIONThe proposed ML and DL methods can classify the PSP and non-PSP microalgae and have achieved better results than previous studies. From the experiments, some of the conclusions are summarized. First, the scattering peaks in the FSSS are not necessary to be removed in advance since it has little effect on the determination of ML and DL. Second, the background noise in the FSSS is also not necessary to be removed in advance, the classification accuracy of ML and DL are not significantly affected. Third, the verification accuracies of the four models were above 91.74%, and the identification accuracy of PSP microalgae is higher than that of non-PSP microalgae; the accuracy of CNN (94.96%) is a bit better than SVM (93.58%), KNN-2 (92.86%), and KNN-1 (91.74%). Forth, the accuracies of the three models for non-PSP microalgae such as IGCZ, DSCF, PSCZ, TWCZ, PDCZ, COHK, PPCZ, and CMHK are all 100%, and the accuracies of CNN in the identification of PSP microalgae, such as AMSY, APQD, and GCFC, are above 97%. The accuracy of KNN-1 for GCFC is 100%, while the accuracy of SVM is 86.49%. Fifth, the accuracies of ML and DL are the lowest when PSP microalgae concentration was 102 cells/mL and the highest when it was 103 cells/mL. Finally, when the EM is 650– 690 nm, the fluorescence characteristics bands (EX) occur differently at 410–480 nm and 500–560 nm for PSP and non-PSP microalgae, respectively. Overall, DL shows good performance in algal toxicity classification.
Establishing a model that can accurately identify algae under complex conditions is helpful to prevent the occurrence of the marine red tide disaster. Classifying algae using ML and DL can reduce the complexity of data processing and improve the accuracy and speed of classification. It is also worthwhile to study the classification of mixed microalgae by adjusting the model parameters and structure. The accurate classification of microalgae in different growth environments will be explored in the future study, to make the model more universally adaptable. Through the methods of this study, the microalgae database is established to improve the understanding of Marine red tide microalgae.
6 DATA AVAILABILITY STATEMENTThe raw data and code supporting the conclusions of this article will be made available by the authors once it was accepted for publication, without undue reservation.
Electronic supplementary material
Supplementary material (Appendix) is available in the online version of this article at https://doi.org/10.1007/s00343-022-1312-1.
Aas K, Eikvil L. 1999. Text Categorization: A Survey. Norwegian Computing Center.
|
Alexander R, Gikuma-Njuru P, Imberger J. 2012. Identifying spatial structure in phytoplankton communities using multi-wavelength fluorescence spectral data and principal component analysis. Limnology and Oceanography: Methods, 10(6): 402-415.
DOI:10.4319/lom.2012.10.402 |
Alimjan G, Sun T L, Liang Y, et al. 2018. A new technique for remote sensing image classification based on combinatorial algorithm of SVM and KNN. International Journal of Pattern Recognition and Artificial Intelligence, 32(7): 1859012.
DOI:10.1142/S0218001418590127 |
Alizadeh J M, Kavianpour M R, Danesh M, et al. 2018. Effect of river flow on the quality of estuarine and coastal waters using machine learning models. Engineering Applications of Computational Fluid Mechanics, 12(1): 810-823.
DOI:10.1080/19942060.2018.1528480 |
Anderson D M, Cembella A D, Hallegraeff G M. 2012a. Progress in understanding harmful algal blooms: paradigm shifts and new technologies for research, monitoring, and management. Annual Review of Marine Science, 4: 143-176.
DOI:10.1146/annurev-marine-120308-081121 |
Anderson D M, Alpermann T J, Cembella A D, et al. 2012b. The globally distributed genus Alexandrium: multifaceted roles in marine ecosystems and impacts on human health. Harmful Algae, 14: 10-35.
DOI:10.1016/j.hal.2011.10.012 |
Andrinolo D, Michea L F, Lagos N. 1999. Toxic effects, pharmacokinetics and clearance of saxitoxin, a component of paralytic shellfish poison (PSP), in cats. Toxicon, 37(3): 447-464.
DOI:10.1016/S0041-0101(98)00173-1 |
Bahram M, Bro R, Stedmon C, et al. 2006. Handling of Rayleigh and Raman scatter for PARAFAC modeling of fluorescence data using interpolation. Journal of Chemometrics, 20(3-4): 99-105.
DOI:10.1002/cem.978 |
Beutler M, Wiltshire K H, Meyer B, et al. 2002. A fluorometric method for the differentiation of algal populations in vivo and in situ. Photosynthesis Research, 72(1): 39-53.
DOI:10.1023/A:1016026607048 |
Bidigare R R, Ondrusek M E, Morrow J H et al. 1990. In-vivo absorption properties of algal pigments. In: Proceedings of SPIE 1302, Ocean Optics X. SPIE, Orlando, USA. p. 290-302, https://doi.org/10.1117/12.21451.
|
Boser B E, Guyon I M, Vapnik V N. 1992. A training algorithm for optimal margin classifiers. In: Proceedings of the 5th Annual Workshop on Computational Learning Theory. ACM, Pittsburgh, USA. p. 144-152, https://doi.org/10.1145/130385.130401.
|
Buhmann M D. 2000. Radial basis functions. Acta Numerica, 9: 1-38.
DOI:10.1017/S0962492900000015 |
Cai Q S, Li R X, Zhen Y, et al. 2006. Detection of two Prorocentrum species using sandwich hybridization integrated with nuclease protection assay. Harmful Algae, 5(3): 300-309.
DOI:10.1016/j.hal.2005.08.002 |
Cao J R, Huan Q L, Wu N, et al. 2015. Effects of temperature, light intensity and nutrient condition on the growth and hemolytic activity of six species of typical ichthyotoxic algae. Marine Environmental Science, 34(3): 321-329.
(in Chinese with English abstract) |
Cherkassky V. 1997. The nature of statistical learning theory. IEEE Transactions on Neural Networks, 8(6): 1564.
DOI:10.1109/TNN.1997.641482 |
Cortes C, Vapnik V N. 1995. Support-vector networks. Machine Learning, 20(3): 273-297.
DOI:10.1007/BF00994018 |
Cover T M. 1968. Estimation by the nearest neighbor rule. IEEE Transactions on Information Theory, 14(1): 50-55.
DOI:10.1109/TIT.1968.1054098 |
Cusick K D, Sayler G S. 2013. An overview on the marine neurotoxin, saxitoxin: genetics, molecular targets, methods of detection and ecological functions. Marine Drugs, 11(4): 991-1018.
DOI:10.3390/md11040991 |
Divya O, Mishra A K. 2007. Multivariate methods on the excitation emission matrix fluorescence spectroscopic data of diesel-kerosene mixtures: a comparative study. Analytica Chimica Acta, 592(1): 82-90.
DOI:10.1016/j.aca.2007.03.079 |
Duan Y L, Su R G, Shi X Y, et al. 2012. Differentiation of phytoplankton populations by in vivo fluorescence based on high-frequency component of wavelet. Chinese Journal of Lasers, 39(7): 0715003.
(in Chinese with English abstract) DOI:10.3788/CJL201239.0715003 |
Guillard R R L, Ryther J H. 1962. Studies of marine planktonic diatoms: I. Cyclotella nana hustedt, and Detonula confervacea (cleve) gran. Canadian Journal of Microbiology, 8(2): 229-239.
DOI:10.1139/m62-029 |
Hallegraeff G M. 1993. A review of harmful algal blooms and their apparent global increase. Phycologia, 32(2): 79-99.
DOI:10.2216/i0031-8884-32-2-79.1 |
Han J, Park J S, Park Y, et al. 2021. Effects of paralytic shellfish poisoning toxin-producing dinoflagellate Gymnodinium catenatum on the marine copepod Tigriopus japonicus. Marine Pollution Bulletin, 163: 111937.
DOI:10.1016/j.marpolbul.2020.111937 |
Hingane M C, Matkar S B, Mane A B, et al. 2015. Classification of MRI brain image using SVM classifier. IJSTE -International Journal of Science Technology & Engineering, 1(9): 24-28.
|
Huan Q, Huang X, Wu N, et al. 2013. Identification of Ichthyotoxic Microalgae Species and Its Hemolytic Activity by Three-Dimensional Fluorescence Spectra. Spectroscopy and Spectral Analysis, 33(2): 399-403.
(in Chinese with English abstract) DOI:10.3964/j.issn.1000-0593(2013)02-0399-05 |
Ignatiades L, Gotsis-Skretas O. 2010. A review on toxic and harmful algae in Greek Coastal Waters (E. Mediterranean Sea). Toxins, 2(5): 1019-1037.
DOI:10.3390/toxins2051019 |
Jaeckisch N, Yang I, Wohlrab S, et al. 2011. Comparative genomic and transcriptomic characterization of the toxigenic marine dinoflagellate Alexandrium ostenfeldii. PLoS One, 6(12): e28012.
DOI:10.1371/journal.pone.0028012 |
Jeffrey S W, Hallegraeff G M. 1980. Studies of phytoplankton species and photosynthetic pigments in a warm core eddy of the East Australian Current. I. Summer populations. Marine Ecology Progress Series, 3: 285-294.
DOI:10.3354/meps003285 |
Jiang T, Wang R, Wu N, et al. 2011. Study on hemolytic activity of Chattonella marina Hong Kong strain. Environmental Science, 32(10): 2920-2925.
(in Chinese with English abstract) |
Johnsen G, Samset O, Granskog L, et al. 1994. In vivo absorption characteristics in 10 classes of bloom-forming phytoplankton: taxonomic characteristics and responses to photoadaptation by means of discriminant and HPLC analysis. Marine Ecology Progress Series, 105: 149-157.
DOI:10.3354/meps105149 |
Kellmann R, Mihali T K, Jeon Y J, et al. 2008. Biosynthetic intermediate analysis and functional homology reveal a saxitoxin gene cluster in cyanobacteria. Applied and Environmental Microbiology, 74(13): 4044-4053.
DOI:10.1128/AEM.00353-08 |
Kotaki Y, Tajiri M, Oshima Y, et al. 1983. Identification of a calcareous red alga as the primary source of paralytic shellfish toxins in coral reef crabs and gastropods. Bulletin of the Japanese Society of Scientific Fisheries, 49(2): 283-286.
DOI:10.2331/suisan.49.283 |
Kumar M S, Sharma S A. 2021. Toxicological effects of marine seaweeds: a cautious insight for human consumption. Critical Reviews in Food Science and Nutrition, 61(3): 500-521.
DOI:10.1080/10408398.2020.1738334 |
LeCun Y. 1992. A theoretical framework for back-propagation. In: Mehra P, Wah B eds. Artificial Neural Networks: Concepts and Theory. IEEE, Los Alamitos.
|
LeCun Y, Bengio Y, Hinton G. 2015. Deep learning. Nature, 521(7553): 436-444.
DOI:10.1038/nature14539 |
LeCun Y, Boser B, Denker J S, et al. 1989a. Backpropagation applied to handwritten zip code recognition. Neural Computation, 1(4): 541-551.
DOI:10.1162/neco.1989.1.4.541 |
LeCun Y, Boser B, Denker J S et al. 1989b. Handwritten digit recognition with a back-propagation network. In: Proceedings of the 2nd International Conference on Neural Information Processing Systems. Morgan Kaufmann, Denver, USA. p. 396-404.
|
LeCun Y, Bottou L, Bengio Y, et al. 1998. Gradient-based learning applied to document recognition. Proceedings of the IEEE, 86(11): 2278-2324.
DOI:10.1109/5.726791 |
Lee T Y, Tsuzuki M, Takeuchi T, et al. 1995. Quantitative determination of cyanobacteria in mixed phytoplankton assemblages by an in vivo fluorimetric method. Analytica Chimica Acta, 302(1): 81-87.
DOI:10.1016/0003-2670(94)00425-L |
Louchard E M., Reid R P, Stephens C F, et al. 2002. Derivative analysis of absorption features in hyperspectral remote sensing data of carbonate sediments. Optics Express, 10(26): 1573-1584.
DOI:10.1364/OE.10.001573 |
Lu L. 2007. Study on Fluorescence Spectra for Identifying Phytoplankton Community. Ocean University of China, Qingdao, China. (in Chinese with English abstract)
|
Lü G C, Zhao W H, Wang J T. 2011. Applications of three-dimensional fluorescence spectrum of dissolved organic matter to identification of red tide algae. Spectroscopy and Spectral Analysis, 31(1): 141-144.
(in Chinese with English abstract) DOI:10.3964/j.issn.1000-0593(2011)01-0141-04 |
Ma Y M, Gao J Y, Wang Q H. 2007. Forecast model for red tide on artificial neural network. Marine Forecasts, 24(1): 38-44.
(in Chinese with English abstract) DOI:10.3969/j.issn.1003-0239.2007.01.006 |
Masó M, Garcés E. 2006. Harmful microalgae blooms (HAB); problematic and conditions that induce them. Marine Pollution Bulletin, 53(10-12): 620-630.
DOI:10.1016/j.marpolbul.2006.08.006 |
Merry R J E. 2005. Wavelet Theory and Applications: A Literature Study. Eindhoven University of Technology Department of Mechanical Engineering Control Systems Technology Group.
|
Millie D F, Schofield O M, Kirkpatrick G J, et al. 1997. Detection of harmful algal blooms using photopigments and absorption signatures: a case study of the Florida red tide dinoflagellate, Gymnodinium breve. Limnology and Oceanography, 42(5): 1240-1251.
DOI:10.4319/lo.1997.42.5_part_2.1240 |
Moberg L, Karlberg B, Sørensen K, et al. 2002. Assessment of phytoplankton class abundance using absorption spectra and chemometrics. Talanta, 56(1): 153-160.
DOI:10.1016/S0039-9140(01)00555-0 |
Mosavi A, Salimi M, Faizollahzadeh Ardabili S, et al. 2019. State of the art of machine learning models in energy systems, a systematic review. Energies, 12(7): 1301.
DOI:10.3390/en12071301 |
O'Neil J M, Davis T W, Burford M A, et al. 2012. The rise of harmful cyanobacteria blooms: the potential roles of eutrophication and climate change. Harmful Algae, 14: 313-334.
DOI:10.1016/j.hal.2011.10.027 |
Ozawa T, Ishihara S, Fujishiro M, et al. 2020. Automated endoscopic detection and classification of colorectal polyps using convolutional neural networks. Therapeutic Advances in Gastroenterology, 13: 175628482091065.
DOI:10.1177/1756284820910659 |
Paerl H W, Gardner W S, Havens K E, et al. 2016. Mitigating cyanobacterial harmful algal blooms in aquatic ecosystems impacted by climate change and anthropogenic nutrients. Harmful Algae, 54: 213-222.
DOI:10.1016/j.hal.2015.09.009 |
Poryvkina L, Babichenko S, Leeben A. 2000. Analysis of phytoplankton pigments by excitation spectra of fluorescence. In: Proceedings of EARSeL-SIG-Workshop LIDAR. FRG, Dresden, Germany. p. 224-232.
|
Qi X L, Wu Z Z, Zhang C S, et al. 2016. A fluorescence technology for discriminating toxic algae by support sector machine regression. Periodical of Ocean University of China, 46(12): 73-80.
(in Chinese with English abstract) |
Raphael A, Dubinsky Z, Iluz D, et al. 2020. Deep neural network recognition of shallow water corals in the Gulf of Eilat (Aqaba). Scientific Reports, 10(1): 12959.
DOI:10.1038/s41598-020-69201-w |
Rasmussen S A, Andersen A J C, Andersen N G, et al. 2016. Chemical diversity, origin, and analysis of phycotoxins. Journal of Natural Products, 79(3): 662-673.
DOI:10.1021/acs.jnatprod.5b01066 |
Sebastiani F. 2002. Machine learning in automated text categorization. ACM Computing Surveys, 34(1): 1-47.
DOI:10.1145/505282.505283 |
Seppälä J, Olli K. 2008. Multivariate analysis of phytoplankton spectral in vivo fluorescence: estimation of phytoplankton biomass during a mesocosm study in the Baltic Sea. Marine Ecology Progress Series, 370: 69-85.
DOI:10.3354/meps07647 |
Sommer H, Monnier R P, Riegel B, et al. 1948. Paralytic shellfish poison. I. Occurrence and concentration by ion exchange. Journal of the American Chemical Society, 70(3): 1015-1018.
DOI:10.1021/ja01183a038 |
Sommer H, Whedonc W F, Kofoid A, et al. 1937. Relation of paralytic shellfish poison to certain plankton organisms of the genus Gonyaulax. Archives of Pathology, 24(5): 537-559.
|
Sultana F, Sufian A, Dutta P. 2019. Advancements in image classification using convolutional neural network. IEEE.
DOI:10.1109/ICRCICN.2018.8718718 |
Tang X H, Yu R C., Zhou M J, et al. 2012. Application of rRNA probes and fluorescence in situ hybridization for rapid detection of the toxic dinoflagellate Alexandrium minutum. Chinese Journal of Oceanology and Limnology, 30(2): 256-263.
DOI:10.1007/s00343-012-1142-7 |
Taroncher-Oldenburg G, Kulis D M, Anderson D M. 1997. Toxin variability during the cell cycle of the dinoflagellate Alexandrium fundyense. Limnology and Oceanography, 42(5): 1178-1188.
DOI:10.4319/lo.1997.42.5_part_2.1178 |
Taroncher-Oldenburg G, Kulis D M, Anderson D M. 1999. Coupling of saxitoxin biosynthesis to the G1 phase of the cell cycle in the dinoflagellate Alexandrin fundyense: temperature and nutrient effects. Natural Toxins, 7(5): 207-219.
DOI:10.1002/1522-7189(200009/10)7:5<207::AID-NT61>3.0.CO;2-Q |
Van Dolah F M, Roelke D, Greene R M. 2001. Health and ecological impacts of harmful algal blooms: risk assessment needs. Human and Ecological Risk Assessment: An International Journal, 7(5): 1329-1345.
DOI:10.1080/20018091095032 |
Vishwanathan S V M, Narasimha Murty M. 2002. SSVM: a simple SVM algorithm. In: Proceedings of 2002 International Joint Conference on Neural Networks. IEEE, Honolulu, USA. p. 2393-2398.
|
Wang L, Xu X, Dong H, et al. 2018. Multi-pixel simultaneous classification of PolSAR image using convolutional neural networks. Sensors (Basel), 18(3): 769.
DOI:10.3390/s18030769 |
Wang Q, Pang W J, Mao Y D, et al. 2020. Changes of extracellular polymeric substance (EPS) during Microcystis aeruginosa blooms at different levels of nutrients in a eutrophic microcosmic simulation device. Polish Journal of Environmental Studies, 29(1): 349-360.
DOI:10.15244/pjoes/102367 |
Xu C M, Jackson S A. 2019. Machine learning and complex biological data. Genome Biology, 20(1): 76.
DOI:10.1186/s13059-019-1689-0 |
Yang I, John U, Beszteri S. 2010. Comparative gene expression in toxic versus non-toxic strains of the marine dinoflagellate Alexandrium minutum. BMC Genomics, 11: 248.
DOI:10.1186/1471-2164-11-248 |
Yang P, Li X L. 1998. Study on marine algal toxic food poisoning (review). Chinese Journal of Food Hygiene, 10(1): 40-43, 45.
(in Chinese) |
Zavala-Mondragon L A, Lamichhane B, Zhang L, et al. 2020. CNN-SkelPose: a CNN-based skeleton estimation algorithm for clinical applications. Journal of Ambient Intelligence and Humanized Computing, 11(6): 2369-2380.
DOI:10.1007/s12652-019-01259-5 |
Zepp R G, Sheldon W M, Moran M A. 2004. Dissolved organic fluorophores in southeastern US coastal waters: correction method for eliminating Rayleigh and Raman scattering peaks in excitation-emission matrices. Marine Chemistry, 89(1-4): 15-36.
DOI:10.1016/j.marchem.2004.02.006 |
Zhang F, Su R, Wang X Z, et al. 2008. Fluorescence Characteristics Extraction and Differentiation of Phytoplankton. Chinese Journal of Lasers, 35(12).
(in Chinese with English abstract) |
Zhang J, Qiu H, Li X Y, et al. 2018a. Real-time nowcasting of microbiological water quality at recreational beaches: a wavelet and artificial neural network-based hybrid modeling approach. Environmental Science & Technology, 52(15): 8446-8455.
DOI:10.1021/acs.est.8b01022 |
Zhang S F, Zhang Y, Lin L, et al. 2018b. iTRAQ-Based quantitative proteomic analysis of a toxigenic dinoflagellate Alexandrium catenella at different stages of toxin biosynthesis during the cell cycle. Marine Drugs, 16(12): 491.
DOI:10.3390/md16120491 |
Zhuang J X, Cai J B, Wang R X et al. 2020. Deep kNN for medical image classification. In: Proceedings of the 23rd International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, Lima, Peru. p. 127-136, https://doi.org/10.1007/978-3-030-59710-8_13.
|
 2022, Vol. 40
2022, Vol. 40


