Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZHAO Peipei, WU Qinghua, XIA Xuekui, GUO Shiyi, SHEN Sizhong, WANG Yujue, HUANG Aiyou
- Metabolomic and proteomic responses of Phaeodactylum tricornutum to hypoxia
- Journal of Oceanology and Limnology, 40(5): 1963-1973
- http://dx.doi.org/10.1007/s00343-021-1232-5
Article History
- Received Jul. 16, 2021
- accepted in principle Sep. 1, 2021
- accepted for publication Nov. 4, 2021
2 State Key Laboratory of Marine Resource Utilization in the South China Sea, Hainan University, Haikou 570228, China;
3 Laboratory of Development and Utilization of Marine Microbial Resource, Hainan University, Haikou 570228, China;
4 Key Laboratory of Tropical Hydrobiology and Biotechnology of Hainan Province, Haikou 570228, China;
5 College of Marine Sciences, Hainan University, Haikou 570228, China;
6 Department of Chemistry, University of Washington, Seattle WA 98195, USA
Plants and algae can respond to low oxygen by remodeling their metabolism pathways. The transition from mitochondrial respiration to fermentation is representative of anaerobic metabolism in most organisms (Geigenberger, 2003; Banti et al., 2013). Rapid activation of lactate dehydrogenase (LDH) catalyzing the reduction of pyruvate to lactate occurs in almost all plant species under hypoxic conditions (Rivoal et al., 1991; Sweetlove et al., 2000). Chlamydomonas produces hydrogen (H2) in the absence of oxygen. It is speculated that H2 may have evolved as a component of photoprotection, but the precise physiological functions of hydrogenase and H2 release in Chlamydomonas remains unclear (Grossman et al., 2011).
Additionally, hypoxia may lead to decreased rate of photorespiration, which can affect cellular carbon and nitrogen metabolism pathways. For example, in cyanobacteria, the enhanced photorespiration due to low carbon availability leads to an accumulation of 2-phosphoglycolate, a sign of carbon deficiency (Jiang et al., 2018). 2-phosphoglycolate also promotes the expression of carbon assimilation-related genes and thus regulates carbon and nitrogen balance (Zhang et al., 2018). Photorespiration is associated with intracellular (near ribulose-1, 5-bisphosphate carboxylase/oxygenase, RuBisCO) carbon-oxygen levels (Jiang et al., 2018). Thereby, if hypoxia can lead to a decrease in photorespiration and 2-phosphoglycolate levels, and in turn, reduce carbon assimilation, remains unknown. Hypoxia may also affect oxidative phosphorylation (Solaini et al., 2010), and thus alter the cellular energy state and induce changes in cellular carbon metabolism (glycolysis, TCA cycle, lactic acid metabolism, and glyoxylic acid cycle) to maintain an intracellular balance between matter and energy.
Diatoms are important unicellular photoautotrophic eukaryotes that play a crucial role in ecology; they contribute to one-fifth of the total primary productivity on the Earth (Falkowski et al., 1998; Field et al., 1998; Kroth, 2007). The success of diatoms in marine environments is attributed to their rapid response to nitrogen fluctuations. In addition, in some regions in modern oceans, high carbon and low oxygen trends are prevalent (Khangaonkar et al., 2021). Phaeodactylum tricornutum Bohlin is the first diatom whose complete genome is sequenced, which provides a universal molecular toolbox and a stable transgenic expression system (Bowler et al., 2008). Thus, it is a model organism for studying physiology, evolution, and biochemistry. In addition, P. tricornutum is widely distributed across coastal and inland waters (including but not limited to the Great Salt Lake), where the environment is unstable and temperature, light quality, salinity, and dissolved oxygen levels fluctuate rapidly (Rushforth et al., 1988; Yang and Zhang, 2020). Therefore, it is of great ecological and physiological significance to study the underlying response mechanism of diatoms in a low oxygen environment using P. tricornutum model. In this study, we compared the growth rates and pigment contents of P. tricornutum under different oxygen concentrations. Proteomic and metabolomic analyses were conducted. The results help to reveal the different metabolic changes to hypoxia of diatom from that of higher plants, which might be ascribed to their different habitats and needs further exploration in the future.
2 MATERIAL AND METHOD 2.1 Strain, culture conditions, and growth measurementsPhaeodactylum tricornutum were cultured axenically in f/2 medium supplemented with steam-sterilized NaHCO3-free artificial seawater, f/2 vitamins (0.5-μg/mL Biotin, 100-μg/mL Vitamin B1, 0.5-μg/mL Vitamin B12, filter sterilized), and inorganic nutrients (Guillard, 1975; Harrison et al., 1980). Cultures were incubated in cool-white-fluorescent light at 100 μmol/ (m2·s) in 12 h: 12 h light: dark cycle at 20 ℃ for approximately seven days. In low oxygen condition (LO), cultures were incubated in 2-L flasks containing 1.5-L medium, continuously aerated with a sterilized gas mixture of 2% O2, 0.04% CO2, and 97.96% N2, bubbled at a constant flow rate of 500 mL/min. In normal conditions (NC), sterilized air was used instead of the hypoxia gas mixture. Each treatment was conducted in triplicates. Cell growth was measured at 730 nm using a ultraviolet-visible (UV-VIS) spectrophotometer (UV-1800, Shimadzu, Japan) (Zhao et al., 2018). Cells were harvested after eight days of cultivation by centrifugation for 5 min at 4 000×g. Cell pellets were frozen instantly in liquid nitrogen and stored at -80 ℃ until protein extraction and further use.
2.2 Metabolomics analysisSamples were freeze-dried and suspended in 1 mL of 50% v/v methanol-water buffer (vortexed for 1 min). Mixtures were centrifuged at 12 000×g for 5 min; the supernatant was lyophilized and re-dissolved in 0.45-mL water. Subsequently, a 50-μL 2, 2-dimethyl-2-silapentane-5-sulfonate (DSS) standard solution (Anachro, Canada) was added. Samples were mixed well before transferring into 5-mm nuclear magnetic resonance (NMR) tube (Norell, USA). Spectra were collected on the Bruker AV III 600-MHz spectrometer. The first increment of a 2D-1H, 1H-NOESY pulse sequence was utilized for the acquisition of 1H-NMR data and for suppressing the solvent signal. The mixing time was set to 100 ms along with a 990-ms pre-saturation time (~80-Hz gammaB1). A total of 128 scans for 15 min; spectra were collected at 25 ℃.
2.3 P. tricornutum pigment analysisPigment extraction and analysis from P. tricornutum were performed according to the previously described methods of Thayer and Björkman (1990), and Enriquez et al. (2010) with slight modifications. Briefly, the collected cells were freeze-dried, suspended in acetone/methanol (1/1, v/v) on ice for 1 h, and then centrifuged at 6 000×g for 5 min. The extraction was repeated thrice and the supernatant was pooled. After filtration, the supernatant was injected into the Agilent 1260 high performance liquid chromatography (HPLC) equipped with a reversed-phase column (YMC-Pack ODS-A, 10×250 mm). Analysis of the mobile phase, consisting of water, methanol, and acetonitrile, was programmed as follows: 0–15 min, linear gradient from I (15% water, 30% methanol, and 55% acetonitrile) to II (0% water, 15% methanol, and 85% acetonitrile); 15–30 min, linear gradient from II to III (100% methanol); and 30–40 min, III. The column temperature was set at 50 ℃ and the flow rate was 0.75 mL/min. Fucoxanthin was detected at 450 nm.
2.4 Preparation of P. tricornutum water-soluble proteinsCell pellets were ground in liquid nitrogen using a mortar and pestle. Pulverized cells were suspended in 10-mL phenol extraction buffer (Solarbio, T0250, containing 10-mmol/L dithiothreitol (DTT) and 1% protease inhibitor cocktail). Mixtures were subject to ultrasonic lysis. One volume of tris-phenol buffer was added to the mixtures. The cell lysate was centrifuged at 5 500×g for 10 min at 4 ℃ and the phenol supernatant was collected. Five volumes of 0.1 mol/L cold ammonium acetate-methanol buffer were added to the supernatant and precipitated overnight at -20 ℃. The precipitate was washed using cold methanol and acetone and dissolved in 8-mol/L urea. Protein concentration was determined by the Bradford assay (Bradford, 1976).
2.5 Label-free quantitative proteomicsTrichloroacetic acid at a final concentration of 20% was added to the samples, eddy mixed and precipitated at 4 ℃ for 2 h. The supernatant was discarded after centrifugation at 4 500×g for 5 min at 4 ℃. The supernatant was washed twice and precipitated with pre-chilled acetone. After the precipitate was dried, tetraethylammonium bromide (TEAB) at a final concentration of 200 mmol/L was added, the precipitate was dispersed by ultrasound, and trypsinized at a ratio of 1 50 (protease protein, m/m) overnight (enzymatic hydrolysis). DTT was added to a final concentration of 5 mmol/L and reduced at 56 ℃ for 30 min. Iodoacetamide (IAA) was added to make up to the final concentration to 11 mmol/L and incubated at room temperature for 15 min in dark (Wiśniewski et al., 2009).
The peptides were dissolved in mobile phase A (0.1% formic acid, 2% acetonitrile) and separated using a NanoElute ultra-high performance liquid phase system. Liquid phase gradient settings were as follows: 0–70 min, 6%–24% mobile phase B (0.1% formic acid in acetonitrile); 70–84 min, 24%–35% B; 84–87 min, 35%–80% B; 87–90 min, 80% B, and the flow rate was maintained at 450 nL/min. After separation on the ultra-high performance liquid phase platform, the peptide was injected into the capillary ion source for ionization and subsequently analyzed using timsTOF Pro instrument. The voltage of the ion source was set at 1.7 kV. High resolution time-of-flight (TOF) was used to detect and analyze the parent ion of the peptide and its secondary fragments. Maxquant software (v1.6.15.0) was used to search the secondary mass spectrum data. Search parameter settings and the process were as follows: The database is Phaeodactylum_tricornutum_strain_ CCAP_1055_556484_PR_20200709.fasta (10 465 sequences). Database searches were performed with trypsin digestion specificity. The Maxquant database search results show the label-free quantification (LFQ) intensity of each protein in different samples (the original intensity value of the protein is corrected between samples). After centralizing the LFQ intensity (I) of the protein in different samples, the relative quantitative value (R) of the protein in different samples was calculated by the formula of Rij=Iij/mean(Ij) (i represents the sample and j represents the protein). For multiple replicate sample experiments, calculate the average value of the relative quantitative value of each protein in multiple replicates, and then calculate the ratio of the average of the two samples, and this ratio was used as the final differential expression of the protein between the two samples ratio. Student t-test was conducted on the triplicates in each group. Significantly differentially expressed proteins were defined by the criteria of a fold change (FC) ≥1.5 or ≤0.67 (P < 0.05). Functional annotation of the proteins was performed using non-redundant protein (NR), Swiss Protein (SwissProt), Gene Ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases.
2.6 Protein validation by parallel reaction monitoring (PRM)To verify the protein expression levels in label-free quantitative proteomics, the expression levels of six selected proteins were quantified by PRM analyses. The method for protein extraction and tryptic digestion were the same as described above for the label-free experiment. The tryptic peptides were dissolved in 0.1% formic acid and directly loaded onto a homemade reversed-phase analytical column using the EASY-nLC 1000 ultra performance liquid chromatography (UPLC) system. Mobile phase A was 0.1% formic acid and 2% acetonitrile in water, and mobile phase C was 0.1% formic acid in 90% acetonitrile. Liquid phase gradient settings were as follows: 0–16 min, 7%–25% C; 16–22 min, 25%– 35% C; 22–26 min, 35%–80% C. The flow rate was set at 500 nL/min. The peptides were examined using the Q Exactive Plus tandem mass spectrometry (Thermo Fisher Scientific) platform where the nano-electrospray ionization (NSI) source was coupled online with the UPLC. The electrospray voltage was set to 2.1 kV. The primary m/z scan range was between 430 to 1 200, and the scanning resolution was 70 000. The scanning resolution of secondary mass spectrometry Orbitrap was 17 500. Data independent scanning (DIA) program was used and the fragmentation energy of higher-energy collisional dissociation was 28. The automatic gain control (AGC) of primary mass spectrometry was 3E6, and the maximum injection time was 50 ms. The AGC of secondary mass spectrometry was 1E5, the maximum injection time was 200 ms, and the isolation window was 1.4 m/z.
2.7 Statistical analysisThere were three replicates for each condition. The student t-test was used to determine significant differences. All data were represented as the mean of three independent experiments. The mean value was used to calculate the ratio (fold changes, FC). For metabolomics analysis, metabolites with P-value < 0.05 and P-value < 0.01 were marked with an asterisk and two asterisks, respectively. For proteomics analysis, the P-value of the differentially expressed proteins shown in the results were all less than 0.01 or 0.05, as indicated.
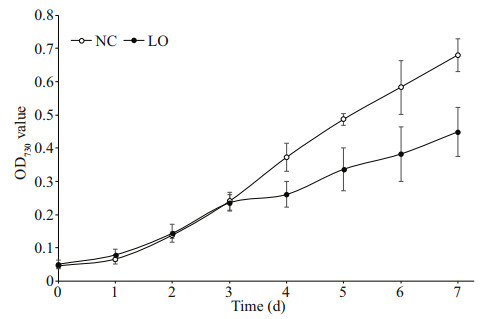
3 RESULT 3.1 Growth of P. tricornutumWe compared the growth rates of P. tricornutum under hypoxic and normal conditions. The growth rate of P. tricornutum under hypoxic condition was significantly lower as compared to that under normal conditions (Fig. 1), which indicated that hypoxia condition could inhibit the growth of P. tricornutum.
|
| Fig.1 OD730 value for the growth of P. tricornutum under different culture conditions LO: P. tricornutum cultured under hypoxic condition; NC: P. tricornutum cultured under normal condition. Bars represent standard deviation (SD). |
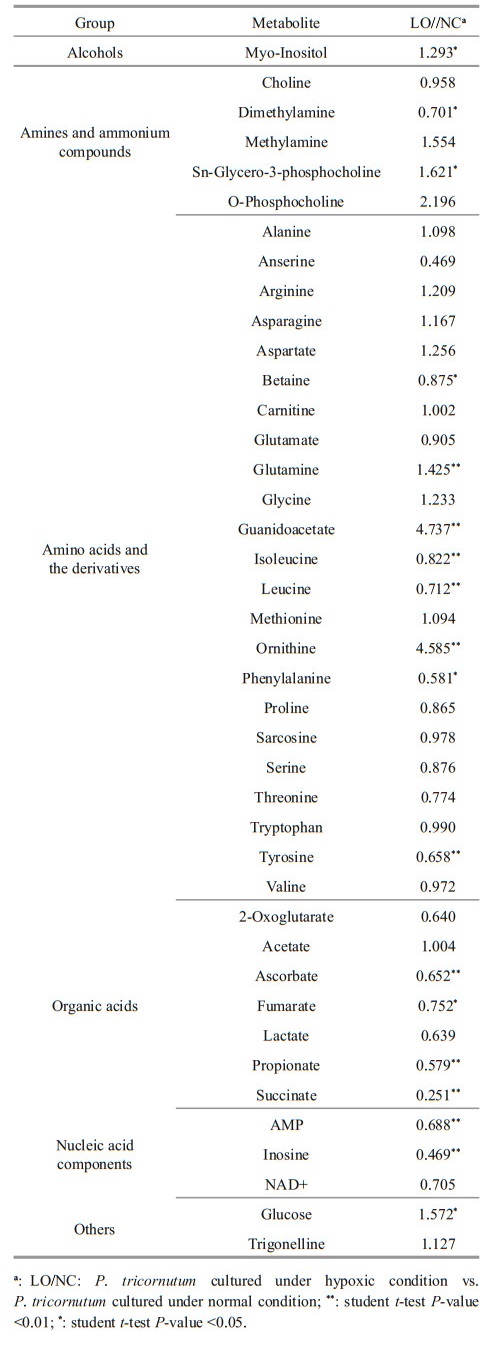
Metabolite profiling showed that the levels of amino acids such as serine, leucine, phenylalanine, and tyrosine, whose biosynthesis was the derivative of glycolysis were significantly reduced in LO culture (P. tricornutum cultured under hypoxic condition) (Table 1). Level of threonine, derivative from TCA cycle, was also reduced. However, levels of other amino acids derivative from the TCA cycle, such as glutamine, increased in LO culture. Ornithine level was also significantly increased in LO culture. Intermediates of the TCA cycle, including fumarate and succinate, were down-regulated in LO culture. For amines and ammonium compounds, the levels of dimethylamine and sn-glycero-3-phosphocholine were significantly altered between the two conditions (LO and NC).

|
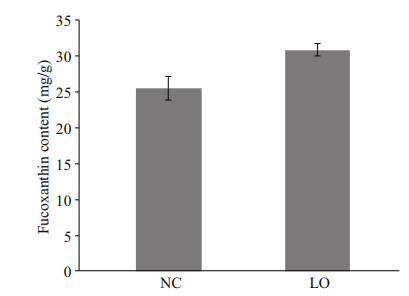
The fucoxanthin content of P. tricornutum increased significantly (P < 0.01) under hypoxic condition, which was 30.85±0.84 mg/g dry weight (DW) and 25.46±1.61 mg/g DW under hypoxic condition and normal conditions, respectively (Fig. 2).
|
| Fig.2 Fucoxanthin content of P. tricornutum under different culture conditions LO: P. tricornutum cultured under hypoxic condition; NC: P. tricornutum cultured under normal condition. Bars represent standard deviation (SD). |
The proteomic analysis yielded 271 differential expression proteins with 129 was up-regulated and 142 was down-regulated (LO/NC: P. tricornutum cultured under hypoxia condition vs. P. tricornutum cultured under normal condition) (data are available via ProteomeXchange with project ID: PXD028642). The potential cellular functions of the differentially expressed proteins identified were searched for in UniProt (http://www.uniprot.org/) and classified into several categories (Figs. 3-4). More than half of the proteins had no notation in the database. Annotated proteins were classed into categories and described in Fig. 3. Here annotated proteins were proteins with annotation in UniProt (http://www.uniprot.org/). Predicted proteins were proteins which had no notation and only were named as 'Predicted protein' or 'Hypothetical protein' in the database.

|
| Fig.3 Functional categories of differentially expressed proteins |
|
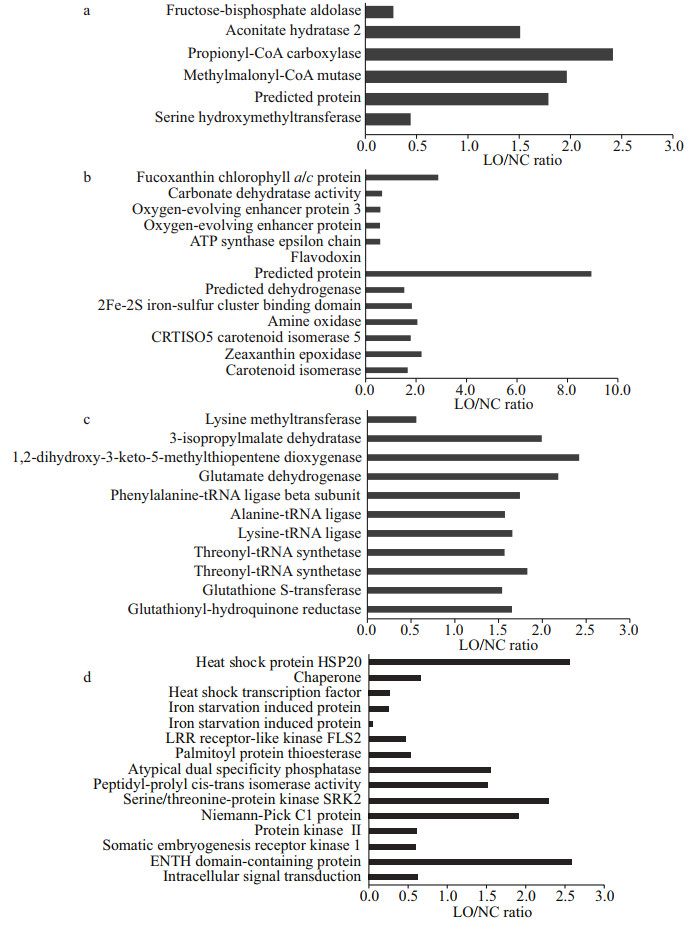
| Fig.4 Differentially expressed proteins in LO/NC group a. proteins involved in central carbon mechanism; b. proteins involved in photosynthesis, electron transport, and biosynthesis of photosynthetic pigments; c. proteins involved in amino acid metabolism; d. proteins involved in stress tolerance and signal transduction. LO/NC: P. tricornutum cultured under hypoxic condition vs. P. tricornutum cultured under normal condition. |
A fructose-bisphosphate aldolase (UniProt accession number, B7GE67) related to the glycolysis pathway was down-regulated in LO culture (Fig. 4a). Aconitate hydratase 2/2-methylisocitrate dehydratase (B7FUR4), which catalyzes the interconversion between citric acid and isocitrate in the TCA cycle, was significantly increased in LO culture. PropionylCoA carboxylase (B7GCL6) and methylmalonylCoA mutase (B7FX73), responsible for the first and third step reactions of propionyl-CoA to succinylCoA conversions, were up-regulated by 2.42 fold and 1.97 fold respectively in LO culture. Propionyl-CoA is an end-product of β-oxidized odd chain fatty acids and it enters the TCA cycle after its conversion to succinyl-CoA. Serine hydroxymethyltransferase (B5Y594), an enzyme that participates in photorespiration, was decreased in LO culture.
3.4.2 Photosynthesis, electron transport, and biosynthesis of photosynthetic pigmentsFucoxanthin chlorophyll a/c protein (B7G4U8), the antenna protein in diatoms, was up-regulated by 2.88-fold in LO culture (Fig. 4b). Two oxygen-evolving enhancer proteins (B7FZ94, B7FZ96) were down-regulated in LO culture. An amine oxidase (B7GBV4) and a zeaxanthin epoxidase (B7FUR7) were up-regulated by 2.05-fold and 2.22-fold, respectively, in LO culture. These two up-regulated proteins indicated that the production of photosynthetic pigments could be enhanced in LO culture.
3.4.3 Amino acid metabolismA lysine methyltransferase (B5Y4M4) was down-regulated, while a 3-isopropylmalate dehydratase (B7FPZ8), a 1, 2-dihydroxy-3-keto-5-methylthiopentene dioxygenase (B7FQT4), and a glutamate dehydrogenase (B7G3X3) were up-regulated in LO culture (Fig. 4c). A phenylalanine-tRNA ligase beta subunit (A0T0H4), an alanine-tRNA ligase (B7G7J0), and two threonyl-tRNA synthetases (B5Y5F7, B7S423) were also up-regulated in LO culture.
3.4.4 Stress tolerance and signal transductionA heat shock protein HSP20 (B7G195) was up-regulated in LO culture (Fig. 4d), while a heat shock transcription factor (B7FWU3) was down-regulated in LO culture. Two iron starvation induced proteins (B7FYL2, B7G4H8) were down-regulated in LO culture. The abundance of other proteins involved in stress tolerance and signal transduction was also altered, which implied that stress response mechanisms of P. tricornutum were turned on under hypoxic conditions. The target protein levels detected by PRM were consistent with those in label-free quantitative proteomics, which indicated that the label-free quantitative proteomic results were highly reliable and reproducible (Supplementary Fig.S1).
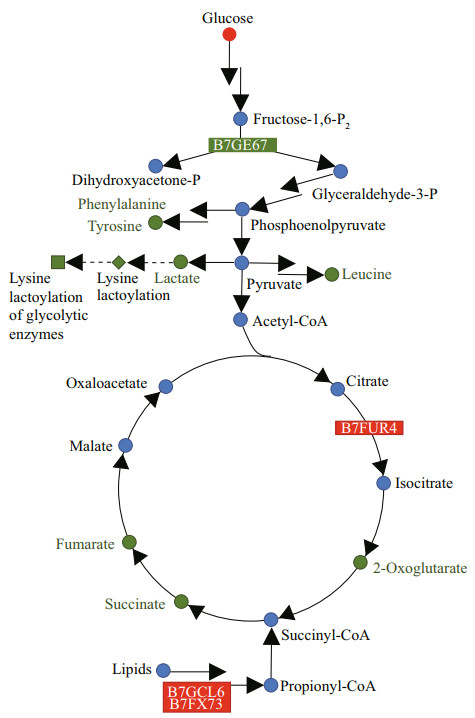
4 DISCUSSION 4.1 P. tricornutum resets carbon metabolism under hypoxic conditionThe response of P. tricornutum under LO condition was different from that of higher plants. In almost all plant species under hypoxic conditions, the rapid activation of LDH, which catalyzes the reduction of pyruvate to lactate, was observed (Banti et al., 2013), whereas a down-regulation of the glycolytic enzyme fructose-bisphosphate aldolase (B7GE67) and the glycolytic by-products lactate was observed in P. tricornutum under LO condition (Fig. 5). We proposed that this difference might be due to their different structures. The structures of higher plants are more complex, which divide into photosynthetic and non-photosynthetic tissues. The response of non-photosynthetic tissues to hypoxia is different to that of photosynthetic tissues, with non-photosynthetic tissues are more susceptible to hypoxia. For gray poplar, glycolytic flux and ethanolic fermentation were stimulated in roots but not in leaves (Kreuzwieser et al., 2009). It is probable that in photosynthetic tissues, O2 generation from photosynthesis is enough for the demand of cells. As P. tricornutum is single cell photosynthetic organism, its response to LO condition might be similar to photosynthetic tissues of higher plants. That is probably why lactate was not increase but decrease in P. tricornutum. Besides, lactate play an important role in regulation of glycolytic enzymes. A preliminary experiment on lactoylation proteomics shows that the lysine lactoylation of the whole proteome decreases in P. tricornutum cultured under hypoxic conditions (unpublished data), which was coincides with the decreased lactate levels. The lysine lactoylation is enriched in glycolytic enzymes and results in decreased glycolytic output (Gaffney et al., 2020). This is a feedback mechanism similar to that through 3-phosphoglyceryl-Lys, leading to a decrease in the levels of glycolytic metabolites (Moellering and Cravatt, 2013). The down-regulation of glycolytic enzyme and by-products might weaken the feedback mechanism in P. tricornutum.
|
| Fig.5 Central carbon metabolism regulation network of P. tricornutum under hypoxic condition The green symbols and boxes represent down-regulated metabolites and proteins, respectively. The blue symbols represent metabolites with no change in content. The red symbols and boxes represent up-regulated metabolites and proteins, respectively. The dashed lines indicate that some reaction steps were omitted. |
An up-regulation of aconitate hydratase 2/2-methylisocitrate dehydratase (B7FUR4), which catalyzes citric acid to isocitrate in the TCA cycle, was observed under LO condition. Yet this did not lead to the increase of downstream products, isocitrate and 2-oxoglutarate. Meanwhile, the intermediate products of the TCA cycle, including succinate, 2-oxoglutarate, and fumarate, showed a downward trend (Table 1; Fig. 5). Amino acids derived from glycolysis and TCA cycle also showed a downward trend (Table 1; Fig. 5), including leucine derived from pyruvate, phenylalanine and tyrosine from phosphoenolpyruvate and erythrose-4-phosphate, and so on. We speculated that this might be a result of the down-regulation of glycolysis products as well as TCA cycle products in LO condition. Besides, a rapid consumption of TCA metabolic intermediates might also result in their depletion. For example, the glyoxylate cycle, which requires plenty of isocitrate, is particularly important in plant seeds (Eastmond et al., 2000). The stored lipids in the seeds generate sugars through acetyl-CoA, which timely supply the energy and carbon skeleton and promote germination and growth. The glyoxylate cycle improves the ability of the organism to use acetyl-CoA. With only a small amount of oxaloacetic acid as a primer, the glyoxylate cycle can continually to produce succinate to replenish four carbon units for the TCA cycle. In this study, we observed the up-regulated propionyl-CoA carboxylase (B7GCL6) and methylmalonyl-CoA mutase (B7FX73). This might promote the conversion of propionyl-CoA to succinyl-CoA and enhanced the use of lipids and replenishment of the TCA cycle. Thus, P. tricornutum could reset its carbon mechanism to cope with hypoxic stress.
4.2 P. tricornutum enhances biosynthesis of photosynthetic pigment under hypoxic conditionAlgae accumulate carotenoids under stress conditions. In this study, we observed an increase in fucoxanthin content in P. tricornutum under LO condition. Accordingly, the key enzyme of photosynthesis, fucoxanthin chlorophyll a/c protein (B7G4U8), was upregulated. Meanwhile, an amine oxidase (B7GBV4) and a zeaxanthin epoxidase (B7FUR7), which participate in carotenoid biosynthesis, were up-regulated. Amine oxidase catalyzes the conversion of phytoene to phytofluene, and phytofluene to ζ-carotene. ζ-carotene can be converted to zeaxanthin in a six-step reaction. Zeaxanthin epoxidase catalyzes the conversion of zeaxanthin to antheraxanthin and antheraxanthin to violaxanthin. Violaxanthin can be converted to neoxanthin and subsequently fucoxanthin (Dambek et al., 2012). Fucoxanthin in fucoxanthin-chlorophyll a complexes, resembles the light-harvesting complexes, and functions as an antenna and transfers excitation energy to chlorophyll (Owens and Wold, 1986; Papagiannakis et al., 2005). Pigment analysis showed that fucoxanthin content in P. tricornutum increased in LO culture, consistent with proteomic results. Thus, this response may provide continued energy supply for P. tricornutum under hypoxic condition. This might be another response of P. tricornutum to LO condition.
4.3 P. tricornutum initiates other stress resistance mechanisms under hypoxic conditionUp-regulated heat shock proteins might be also one of LO responses of P. tricornutum. Whether oxidative stress exists under hypoxic condition remains controversial. It seems counterintuitive since oxygen is needed for the production of reactive oxygen species (ROS). However, activation of ROS-regulated genes or ROS-associated is reported in different plants under hypoxic conditions (Banti et al., 2013). Heat Shock Proteins (HSPs) are the most common among the ROS-related proteins, which are also induced by anoxia (Mustroph et al., 2010; Pucciariello et al., 2012). Some ROS-related transcription factors (TFs) such as HsfA2 and ZAT12, associated with ROS production are also regulated by hypoxia (Pucciariello et al., 2012). Our results show an up-regulation in levels of heat shock protein HSP20 (B7G195), and a down-regulation of a heat shock transcription factor (B7FWU3), which corresponded to the findings in plants. The synthesis of iron starvation-induced proteins (B7FYL2, B7G4H8) was also up-regulated under LO condition. These proteins usually related with silicon and iron starvation (Zhao et al., 2014, 2018).
The high expression of ornithine and arginine under hypoxic condition may also be one of the stress resistance mechanisms of P. tricornutum. In plants, the existence of the urea cycle has been demonstrated in some species by the isotope tracer method (Slocum, 2005), but the evidence is limited. It is speculated that ornithine in chloroplasts is the initial substrate for the urea cycle in plants and is the precursor of polyamines; ornithine in mitochondria also participates in the synthesis of proline and other amino acids (Winter et al., 2015). Arginine is a crucial amino acid having several functions in plants. It is not only a component of the proteins, a medium for nitrogen storage and transport but also an osmotic regulator. Arginine not only participates in the synthesis of proline and nitric oxide (NO) through the urea cycle and the citrulline-NO cycle but also in the biosynthesis of polyamines (Winter et al., 2015). Thus, metabolic intermediates of the urea cycle play an important role in stress signaling transduction, regulation of ionic balance, generation of antioxidants and osmotic protective substances, the precursors of proline, polyamines, and NO biosynthesis. Additionally, it is directly involved in plant resistance through the synthesis of osmotic protective substances and antioxidants (Liu et al., 2015).
5 CONCLUSIONUnder hypoxic condition, the growth rate of P. tricornutum was reduced. Carbon metabolism in P. tricornutum was reset under an adverse environmental condition, such as the enhancement of lipid utilization. Other stress resistance mechanisms were activated to cope with hypoxia-induced adversity. Due to the difference in habitat as compared to the higher plants, P. tricornutum showed metabolic alterations, which needs further investigation in the future.
6 DATA AVAILABILITY STATEMENTThe data that support the findings of this study are available from the corresponding author upon reasonable request.
Electronic supplementary material
Supplementary material (Supplementary Fig.S1) is available in the online version of this article at https://doi.org/10.1007/s00343-021-1232-5.
Banti V, Giuntoli B, Gonzali S, Loreti E, Magneschi L, Novi G, Paparelli E, Parlanti S, Pucciariello C, Santaniello A, Perata P. 2013. Low oxygen response mechanisms in green organisms. International Journal of Molecular Sciences, 14(3): 4734-4761.
DOI:10.3390/ijms14034734 |
Bowler C, Allen A E, Badger J H, Grimwood J, Jabbari K, Kuo A, Maheswari U, Martens C, Maumus F, Otillar R P, Rayko E, Salamov A, Vandepoele K, Beszteri B, Gruber A, Heijde M, Katinka M, Mock T, Valentin K, Verret F, Berges J A, Brownlee C, Cadoret J P, Chiovitti A, Choi C J, Coesel S, De Martino A, Detter J C, Durkin C, Falciatore A, Fournet J, Haruta M, Huysman M J J, Jenkins B D, Jiroutova K, Jorgensen R E, Joubert Y, Kaplan A, Kröger N, Kroth P G, La Roche J, Lindquist E, Lommer M, Martin-jézéquel V, Lopez P J, Lucas S, Mangogna M, Mcginnis K, Medlin L K, Montsant A, Secq M P O, Napoli C, Obornik M, Parker M S, Petit J L, Porcel B M, Poulsen N, Robison M, Rychlewski L, Rynearson T A, Schmutz J, Shapiro H, Siaut M, Stanley M, Sussman M R, Taylor A R, Vardi A, Von Dassow P, Vyverman W, Willis A, Wyrwicz L S, Rokhsar D S, Weissenbach J, Armbrust E V, Green B R, Van De peer Y, Grigoriev I V. 2008. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature, 456(7219): 239-244.
DOI:10.1038/nature07410 |
Bradford M M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72(1-2): 248-254.
DOI:10.1016/0003-2697(76)90527-3 |
Dambek M, Eilers U, Breitenbach J, Steiger S, Büchel C, Sandmann G. 2012. Biosynthesis of fucoxanthin and diadinoxanthin and function of initial pathway genes in Phaeodactylum tricornutum. Journal of Experimental Botany, 63(15): 5607-5612.
DOI:10.1093/jxb/ers211 |
Eastmond P J, Germain V, Lange P R, Bryce J H, Smith S M, Graham I A. 2000. Postgerminative growth and lipid catabolism in oilseeds lacking the glyoxylate cycle. Proceedings of the National Academy of Sciences of the United States of America, 97(10): 5669-5674.
DOI:10.1073/pnas.97.10.5669 |
Enriquez M M, LaFountain A M, Budarz J, Fuciman M, Gibson G N, Frank H A. 2010. Direct determination of the excited state energies of the xanthophylls diadinoxanthin and diatoxanthin from Phaeodactylum tricornutum. Chemical Physics Letters, 493(4-6): 353-357.
DOI:10.1016/j.cplett.2010.05.051 |
Falkowski P G, Barber R T, Smetacek V. 1998. Biogeochemical controls and feedbacks on ocean primary production. Science, 281(5374): 200-206.
DOI:10.1126/science.281.5374.200 |
Field C B, Behrenfeld M J, Randerson J T, Falkowski P. 1998. Primary production of the biosphere: integrating terrestrial and oceanic components. Science, 281(5374): 237-240.
DOI:10.1126/science.281.5374.237 |
Gaffney D O, Jennings E Q, Anderson C C, Marentette J O, Shi T D, Oxvig A M S, Streeter M D, Johannsen M, Spiegel D A, Chapman E, Roede J R, Galligan J J. 2020. Non-enzymatic lysine lactoylation of glycolytic enzymes. Cell Chemical Biology, 27(2): 206-213.
DOI:10.1016/j.chembiol.2019.11.005 |
Geigenberger P. 2003. Response of plant metabolism to too little oxygen. Current Opinion in Plant Biology, 6(3): 247-256.
DOI:10.1016/s1369-5266(03)00038-4 |
Grossman A R, Catalanotti C, Yang W Q, Dubini A, Magneschi L, Subramanian V, Posewitz M C, Seibert M. 2011. Multiple facets of anoxic metabolism and hydrogen production in the unicellular green alga Chlamydomonas reinhardtii. New Phytologist, 190(2): 279-288.
DOI:10.1111/j.1469-8137.2010.03534.x |
Guillard R R L. 1975. Culture of phytoplankton for feeding marine invertebrates. In: Smith W L, Chanley M H eds. Culture of Marine Invertebrate Animals. Springer, Boston. p. 29-60, https://doi.org/10.1007/978-1-4615-8714-9_3.
|
Harrison P J, Waters R E, Taylor F J R. 1980. A broad spectrum artificial sea water medium for coastal and open ocean phytoplankton. Journal of Phycology, 16(1): 28-35.
DOI:10.1111/j.0022-3646.1980.00028.x |
Jiang Y L, Wang X P, Sun H, Han S J, Li W F, Cui N, Lin G M, Zhang J Y, Cheng W, Cao D D, Zhang Z Y, Zhang C C, Chen Y X, Zhou C Z. 2018. Coordinating carbon and nitrogen metabolic signaling through the cyanobacterial global repressor NdhR. Proceedings of the National Academy of Sciences of the United States of America, 115(2): 403-408.
DOI:10.1073/pnas.1716062115 |
Khangaonkar T, Nugraha A, Premathilake L, Keister J, Borde A. 2021. Projections of algae, eelgrass, and zooplankton ecological interactions in the inner Salish Sea-for future climate, and altered oceanic states. Ecological Modelling, 441: 109420.
DOI:10.1016/j.ecolmodel.2020.109420 |
Kreuzwieser J, Hauberg J, Howell K A, Carroll A, Rennenberg H, Millar A H, Whelan J. 2009. Differential response of gray poplar leaves and roots underpins stress adaptation during hypoxia. Plant Physiology, 149(1): 461-473.
DOI:10.1104/pp.108.125989 |
Kroth P. 2007. Molecular biology and the biotechnological potential of diatoms. In: León R, Galván A, Fernández E eds. Transgenic Microalgae as Green Cell Factories. Springer, New York. p. 23-33, https://doi.org/10.1007/978-0-387-75532-8_3.
|
Liu J H, Wang W, Wu H, Gong X Q, Moriguchi T. 2015. Polyamines function in stress tolerance: from synthesis to regulation. Frontiers in Plant Science, 6: 827.
DOI:10.3389/fpls.2015.00827 |
Moellering R E, Cravatt B F. 2013. Functional lysine modification by an intrinsically reactive primary glycolytic metabolite. Science, 341(6145): 549-553.
DOI:10.1126/science.1238327 |
Mustroph A, Lee S C, Oosumi T, Zanetti M E, Yang H J, Ma K, Yaghoubi-Masihi A, Fukao T, Bailey-Serres J. 2010. Cross-kingdom comparison of transcriptomic adjustments to low-oxygen stress highlights conserved and plant-specific responses. Plant Physiology, 152(3): 1484-1500.
DOI:10.1104/pp.109.151845 |
Owens T G, Wold E R. 1986. Light-harvesting function in the diatom Phaeodactylum tricornutum. I. Isolation and characterization of pigment-protein complexes. Plant Physiology, 80(3): 732-738.
DOI:10.1104/pp.80.3.732 |
Papagiannakis E, Van Stokkum I H M, Fey H, Büchel C, Van Grondelle R. 2005. Spectroscopic characterization of the excitation energy transfer in the fucoxanthin-chlorophyll protein of diatoms. Photosynthesis Research, 86(1): 241-250.
DOI:10.1007/s11120-005-1003-8 |
Pucciariello C, Parlanti S, Banti V, Novi G, Perata P. 2012. Reactive oxygen species-driven transcription in Arabidopsis under oxygen deprivation. Plant Physiology, 159(1): 184-196.
DOI:10.1104/pp.111.191122 |
Rivoal J, Ricard B, Pradet A. 1991. Lactate dehydrogenase in Oryza sativa l. Seedlings and roots: identification and partial characterization. Plant Physiology, 95(3): 682-686.
DOI:10.1104/pp.95.3.682 |
Rushforth S R, Johansen J R, Sorensen D L. 1988. Occurrence of Phaedactylum tricornutum in the Great Salt Lake, Utah, USA. Great Basin Naturalist, 48(3): 324-326.
|
Slocum R D. 2005. Genes, enzymes and regulation of arginine biosynthesis in plants. Plant Physiology and Biochemistry, 43(8): 729-745.
DOI:10.1016/j.plaphy.2005.06.007 |
Solaini G, Baracca A, Lenaz G, Sgarbi G. 2010. Hypoxia and mitochondrial oxidative metabolism. Biochimica et Biophysica Acta (BBA)-Bioenergetics, 1797(6-7): 1171-1177.
DOI:10.1016/j.bbabio.2010.02.011 |
Sweetlove L J, Dunford R, Ratcliffe R G, Kruger N J. 2000. Lactate metabolism in potato tubers deficient in lactate dehydrogenase activity. Plant, Cell & Environment, 23(8): 873-881.
DOI:10.1046/j.1365-3040.2000.00605.x |
Thayer S S, Björkman O. 1990. Leaf xanthophyll content and composition in sun and shade determined by HPLC. Photosynthesis Research, 23(3): 331-343.
DOI:10.1007/bf00034864 |
Winter G, Todd C D, Trovato M, Forlani G, Funck D. 2015. Physiological implications of arginine metabolism in plants. Frontiers in Plant Science, 6: 534.
DOI:10.3389/fpls.2015.00534 |
Wiśniewski J R, Zougman A, Nagaraj N, Mann M. 2009. Universal sample preparation method for proteome analysis. Nature methods, 6(5): 359-362.
DOI:10.1038/nmeth.1322 |
Yang L Q, Zhang Y Y. 2020. Effects of atrazine and its two major derivatives on the photosynthetic physiology and carbon sequestration potential of a marine diatom. Ecotoxicology and Environmental Safety, 205: 111359.
DOI:10.1016/j.ecoenv.2020.111359 |
Zhang C C, Zhou C Z, Burnap R L, Peng L. 2018. Carbon/ nitrogen metabolic balance: lessons from Cyanobacteria. Trends in Plant Science, 23(12): 1116-1130.
DOI:10.1016/j.tplants.2018.09.008 |
Zhao P P, Gu W H, Huang A Y, Wu S C, Liu C H, Huan L, Gao S, Xie X J, Wang G C. 2018. Effect of iron on the growth of Phaeodactylum tricornutum via photosynthesis. Journal of Phycology, 54(1): 34-43.
DOI:10.1111/jpy.12607 |
Zhao P P, Gu W H, Wu S C, Huang A Y, He L W, Xie X J, Gao S, Zhang B Y, Niu J F, Lin A P, Wang G C. 2014. Silicon enhances the growth of Phaeodactylum tricornutum Bohlin under green light and low temperature. Scientific Reports, 4: 3958.
DOI:10.1038/srep03958 |
 2022, Vol. 40
2022, Vol. 40


