Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZU Yao, MIAO Wenjie, LUO Yu, XU Chen, PAN Qianhui, CHEN Siyu, LI Jianhong
- Screening of nanobody against Microcystis from a human phage display nanobody library
- Journal of Oceanology and Limnology, 40(5): 1696-1705
- http://dx.doi.org/10.1007/s00343-022-1361-5
Article History
- Received Oct. 29, 2021
- accepted in principle Dec. 7, 2021
- accepted for publication Jan. 21, 2022
Harmful cyanobacterial blooms, in particular Microcystis blooms, were the most pervasive in temperate and tropical freshwater bodies throughout the world (Xiao et al., 2018). Extensive species of Microcystis can be responsible for harmful cyanobacterial blooms and lack effective analytical tools to identify them, making ecological monitoring research difficult. Until now, approximately 51 species of Microcystis genus had been classified and listed in AlgaeBase (Guiry and Guiry, 2019), however, their identification remains controversial.
Traditional identification of microalgae relies on visible traits by microscope, including cell size, gel sheath form, cell arrangement in a colony, etc. (Komárek and Komárková, 2002). It is difficult to accurately identify Microcystis species only based on their morphologic characteristics because they vary depending on the environment, especially colonial appearance (Otsuka et al., 2000; Li et al., 2013; Sun et al., 2016; Xiao et al., 2018). Specific DNA region sequencing is a very useful technique for microalgal identification. However, this procedure is time-consuming and cannot deliver results in real time. Additionally, when examining a water sample with mixed species under a microscope, the DNA analysis is ineffective.
Immunological techniques based on specific antibodies are a rapid, sensitive, and simplified method for pesticides (Zhu et al., 2020), viruses (Bhardwaj et al., 2020), bacteria (Zhang et al., 2021), biotoxins (Szkola et al., 2014), and harmful algae detection (Blanco et al., 2015). Antibodies, both polyclonal and monoclonal antibodies, have been generated for varied algae, including Chlamydomonas reinhardtii (Mitra et al., 2012), Alexandrium minutum (Mendoza et al., 1995; Gas et al., 2009; Carrera et al., 2010), and Prymnesium parvum (West et al., 2006). However, investigations on Microcystis identification using immunological approaches are still in the initial stage. Recently, Blanco et al.(2015, 2017) have developed a validated and cost-effective 17-polyclonal antibody microarray (containing three antibodies against M. aeruginosa, M. flos-aquae, and M. novacekii, respectively) for detecting and identifying cyanobacteria simultaneously. Only three polyclonal antibodies against Microcystis were found to be insufficient in terms of the present variety of Microcystis.
In recent years, phage display library has become a promising technology for rapidly and efficiently obtaining antibodies with high sensitivity and specificity for food safety and environmental contamination detection (Zhao et al., 2016; García-García et al., 2020; Lee et al., 2020). The method works by randomly cloning artificial antibody genes into phagemid vectors, then co-expressing them on the surface of phage capsid proteins, including single-domain antibody (sdAb), also known as nanobody (Nb) (Jiang et al., 2013; Qiu et al., 2018), single-chain antibody (scFv) (Xu et al., 2019a, b), and polypeptide (Lee et al., 2020). The candidate particles could be obtained by multiple rounds of biopanning from the library, and then their genes are sub-cloned into suitable vectors for over-expression (Xu et al., 2021). Nanobodies specific for surface antigens of A. minutum, C. reinhardtii, and M. aeruginosa were isolated from pre-immune or immune phage display libraries and effectively employed in diagnostics (Jiang et al., 2013, 2014; Mazzega et al., 2019; Folorunsho et al., 2021).
In this study, we employed a large diversity and capacity of human domain antibody display library for screening specific Microcystis nanobodies. The most two positive nanobodies were expressed in Escherichia coli BL21 and the DAb2 was specific for different Microcystis. The immuno-dot blot assay was used for Microcystis detection.
2 MATERIAL AND METHOD 2.1 Strain and reagentMicrocystis aeruginosa PCC7806 and Synechocystis sp. PCC6803 were obtained from Pasteur Culture Collection of Cyanobacteria, France. M. aeruginosa FACHB-905 was kindly provided by Dr. Han MENG (Nanjing Normal University). M. flos-aquae FACHB-1272, M. panniformis FACHB-1757, M. wesenbergii FACHB-929, Pediastrum duplex FACHB-1804 and Chroococcus sp. FACHB-193 were from Freshwater Algae Culture Collection at the Institute of Hydrobiology, Wuhan, China. Microcystis XW01, Anabaena sp. JK12, Chlorella sp., and Scenedesmus sp. JK37 were isolated from blooms in Xuanwu Lake, Nanjing, China and Taihu Lake, China. All strains were cultured in liquid BG-11 medium (Waterbury, 2006) at 28±2 ℃ under continuous illumination at 1 500 lx.
Domain antibody (DAb) phage display library (based on a single human VH framework (V3-23/ D47) with diversity introduced in the antigen-binding site and constructed in an ampicillin resistance pR2 phagemid vector, the size of the library is 3×109) was purchased from the Medical Research Council (MRC) Laboratory of Molecular Biology (Cambridge, UK) and used to pan specific Microcystis nanobodies. Helper phage M13K07 and Escherichia coli TG1 (E. coli TG1) were obtained from Nanobody (NB) Biolab (Chengdu, China). E. coli BL21 was purchased from Vazyme (Nanjing, China). Anti-HIS monoclonal antibody and His-Tag mAb (horseradish peroxidase (HRP) conjugated) were purchased from GenScript (Nanjing, China). Skim milk and trypsin were obtained from Sigma-Aldrich (Beijing, China). Six-well and 96-well plates were purchased from Corning (Beijing, China).
2.2 Preparation of protein extractMicrocystis aeruginosa PCC7806 lysates were used as binding templates to select specific antibodies from the DAb phage display library. 50-mL of cells with an optical density of 0.6 at 720 nm (OD720) were harvested by centrifugation at 8 000×g for 5 min at 4 ℃ and washed with phosphate-buffered saline (PBS, pH 7.4) three times. The pellets were re-suspended in 1-mL PBS solution (pH 7.4) and frozen at -80 ℃, then thawed the frozen repertoire on ice. Following triple freezing, the mixtures were centrifuged for 25 min at 8 000×g and 4 ℃. The clear supernatant was collected and stored in small aliquots at -20 ℃ until used. Protein concentration was estimated by the Bradford method (Bradford. 1976), using bovine serum albumin (BSA) as a standard.
2.3 Amplification of the DAb phage display libraryAmplification of the antibody library was described with some modifications in Xu et al. (2018). The frozen stock of phagemid in E. coli TG1 was thawed on ice and added into 250-mL 2×TY-AG medium (16-g tryptone, 10-g yeast extract, and 5-g NaCl in 1-L double distilled water with 100-μg/mL ampicillin and 1% glucose). The mixtures were cultured for 2 h at 37 ℃ and 250 r/min until cells density reached 0.6 at OD600, then KM13K07 helper phages (approximately 1012 pfu/mL) were added and cultured for 1 h at 30 ℃ in a water bath. The cultures were centrifuged at 3 200×g and 30 ℃ for 30 min, then precipitates were re-suspended with 500-mL 2×TY-AK (containing 100-μg/mL ampicillin and 50-μg/mL kanamycin) for growing overnight at 25 ℃ and 250 r/min. The next day, after being centrifuged for 30 min at 3 300×g and 4 ℃, a 500-mL supernatant was obtained, 125-mL PEG/NaCl (containing 20% polyethylene-glycol (PEG) and 2.5-mol/L NaCl) were added into the supernatant, then kept the mixtures on ice for 1 h. The mixtures were centrifuged for 30 min at 3 300×g and 4 ℃ and the precipitates were re-suspended with 5-mL sterile PBS solution and centrifuged at 11 600×g and 4 ℃ for 10 min. The supernatant was the amplified antibody library and the titer of phagemids was estimated by series dilutions of 10-fold and adjusted to 109 CFU/mL for next study.
2.4 Enrichment and biopanning of phage particles binding to M. aeruginosa PCC7806 extractEnrichment and biopanning of particles were performed as described with some modifications by Xu et al. (2018). Six-well plate was coated with 2-mL M. aeruginosa PCC7806 lysates (the first round of screening was 200 μg/mL and the remaining two rounds were 100 μg/mL and 50 μg/mL) for standing overnight at 4 ℃. The next day, the plate was washed with 3-mL PBST solution (containing 0.1% Tween 20 in PBS solution) three times and blocked with 3-mL MPBS solution (containing 5% skim milk powder in PBS solution) for 2 h at 37 ℃. After washing three times with 3-mL PBST solution, 1-mL particles (2.13×109 pfu/mL) mixed with 2-mL MPBS solution was added. The plate was shaken for 1 h at 120 r/min and room temperature, then standing for 1 h at 37 ℃. After being washed ten times with 3-mL PBST solution, unbound phages were removed and bounding ones were eluted by 1-mL trypsin solution (0.2-mg/mL trypsin in PBS solution). The eluent was the first round enriched library. Particles collected from every round were quantified at 109 CFU/mL and evaluated by calculating the output/input and polyclonal phage enzyme-linked immunosorbent assay (ELISA) (Zhang et al., 2012).
According to the results of the output/input and polyclonal phages ELISA, 1.5-mL E. coli TG1 were infected by 0.5 mL of particles eluted in the third round, then spread on TYE-AG solid medium (containing 100-μg/mL ampicillin and 1% glucose) for culturing overnight at 37 ℃. The next day, individual colonies were randomly picked and inoculated into 96-well plates with 100 μL/well of 2×TY-AG medium for culturing overnight at 37 ℃. Two microliters of cultures were transferred to another plate for culturing 2 h at 250 r/min and 37 ℃, then 20 μL/well of KM13K07 helper phages (approximately 1012 pfu/mL) were added into the plates for rescuing 2 h at 250 r/min and 37 ℃. The plates were centrifuged at 3 300×g for 30 min at room temperature and the precipitates were re-suspended with 200-μL 2×TY-AK medium followed by incubating overnight at 250 r/min and 30 ℃. The next day, the plates were centrifuged at 3 300×g for 30 min and the supernatants were used for monoclonal phage ELISA (Xu et al., 2018). A positive monoclonal phage antibody was considered according to the OD450 ratio of P/N > 3 (Wang et al., 2012). P/N=positive (coated with lysates) / negative (coated with carbonate-buffered saline).
2.5 Colony PCR and DNA sequencingThe positive phage nanobody genes were identified by colony PCR and DNA sequencing. The PCR reaction conditions were as follows (primers were LMB3: CAGGAAACAGCTATGAC and pHEN: CTATGCGGCCCCATTCA): 98 ℃ for 3 min, 30 cycles of 98 ℃ for 10 s, 52 ℃ for 10 s and 72 ℃ for 15 s, and final extension at 72 ℃ for 2 min. The amplified products were examined by 1% agarose gel electrophoresis and sequenced by Sangon Bio. Co. Ltd. (Shanghai, China). The alignment of the protein sequence was performed using DNAMAN 9.0 software.
2.6 Soluble expression and purification of nanobodiesThe most two positive nanobody genes (DAb2 and DAb3) were cloned between Nco I and Not I sites of pET-29a(+) and pET-26b(+) vectors for soluble expression, respectively. In brief, the genes were amplified by PCR using primers DAb-Nco I-F CATGCCATGGGCCATGCCATGGGCCAGGT and DAb-Not I-R ATAAGAATGCGGCCGCGCTCGA-GACGGTG, then the products were digested with Nco I and Not I and ligated into pET-29a(+) or pET-26b(+) vectors which had been digested with the same restriction endonucleases. The recombine plasmids were transferred into E. coli BL21 and the positive clones were identified by colony PCR and DNA sequencing using universal primers (T7 promoter and T7 terminator). A well-sequenced single colony was inoculated into 5 mL of Luria-Bertani (LB) medium containing 50-μg/mL kanamycin and incubated at 37 ℃ overnight. Five milliliters of cultures were inoculated into 500 mL of LB medium containing 50-μg/mL kanamycin for culturing until cells density reached 0.6 at OD600, then adding isopropyl-β-D-thiogalactoside (IPTG) with a final concentration of 0.2 mmol/L to induce the antibodies expression for 16 h at 28 ℃ and 150 r/min. The cells were harvested by centrifugation at 8 000×g for 10 min and 4 ℃ and re-suspended with 10-mL ice-cold lysis buffer (50-mmol/L sodium phosphate, 300-mmol/L sodium chloride, and 10-mmol/L imidazole, pH 8.0), then lysed by sonication on ice, working 15 s and intermittent 45 s for 15 min. After being centrifuged at 8 000×g for 30 min and 4 ℃, the supernatant of whole cell lysate was used for preparing pure antibodies by a His60 Ni superflow resin & gravity column. The purified antibodies were dialyzed against PBS solution and concentrated to 3.5 mL using Amicon Ultra-15 centrifugal filter devices (Millipore). Then, the proteins were checked by 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE).
2.7 Gel electrophoresis and immunoblotting analysisPrior to electrophoretic analysis, protein samples were boiled in SDS-PAGE buffer (300-mmol/L Tris-HCl, 10% w/v SDS, 30% v/v glycerol, 9.3% w/v dithiothreitol, and 0.012% bromophenol blue). Then, the proteins were separated with 10% SDS-PAGE gels and electrophoretically transferred to immobilon-P transfer membrane (ABM, catalog number B500). Prior to Western blot analysis, the membrane was blocked with 5% skim milk dissolved TBS solution with gentle rocking for 1.5 h at room temperature. The membrane was washed three times (15 min each) in TBST solution (containing 0.1% Tween 20 in TBS solution) and incubated with purified antibodies (DAb2 and DAb3) for 2 h at room temperature. After being washed with TBST solution, the membrane was incubated with Anti-HIS monoclonal antibody (1 : 10 000 v/v dilutions in TBST solution) followed by a 1-h incubation at room temperature. After washing, the membrane was incubated with goat anti-mouse IgG (HRP) (1 : 10 000 v/v dilutions in TBST solution) for 1 h at room temperature with gentle rocking. After intensive washing as noted above, immunoreactive proteins were visualized by incubation with enhanced chemiluminescent substrate (Tanon, catalog number 180–500) and images were generated by a cooled charge-coupled device (CCD) camera (Tanon-4100).
2.8 Protein extraction and immuno-dot blot analysisAbout 107 cells were broken with 50-μL PBS solution and 0.06–0.08-g glass beads (0.1-mm diameter) using a high-throughput tissue grinder (SCIENTZ) before being centrifuged at 10 000×g for 10 min. Immuno-dot blot assay was performed to evaluate the nanobodies specificity using the supernatant of different Microcystis lysates (M. aeruginosa PCC7806, M. aeruginosa FACHB-905, Microcystis XW01, M. flos-aquae FACHB-1272, M. panniformis FACHB-1757, and M. wesenbergii FACHB-929) and some companion species with Microcystis (Synechocystis sp. PCC6803, Chroococcus sp. FACHB-193, Anabaena sp. JK12, Chlorella sp., Pediastrum duplex FACHB-1804, and Scenedesmus sp. JK37). In addition, a water sample collected from Taihu Lake with more than 90% of Microcystis biomass was also used for detection. 5 μL of the supernatant were spotted onto immobilon-P transfer membrane (ABM, catalog number B500) and allowed to dry up at room temperature. After being blocked with 5% (w/v) skim milk in TBST solution for 1.5 h at room temperature, the membrane was washed three times (15 min each) and incubated with the nanobodies (5-μg/mL diluted in TBS solution) for 2 h at room temperature. After washing, the membrane was incubated with His-Tag mAb (HRP conjugated) (1 5 000 dilutions in TBS solution) for 1 h at room temperature. After intensive washing as noted above, immunoreactive proteins were visualized by incubation with enhanced chemiluminescent substrate (Tanon, catalog number 180–500) and images were generated by a cooled CCD camera (Tanon-4100).
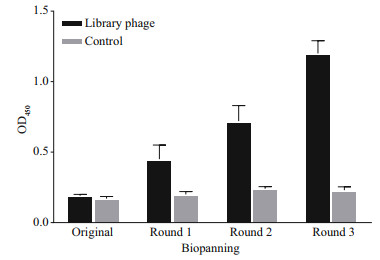
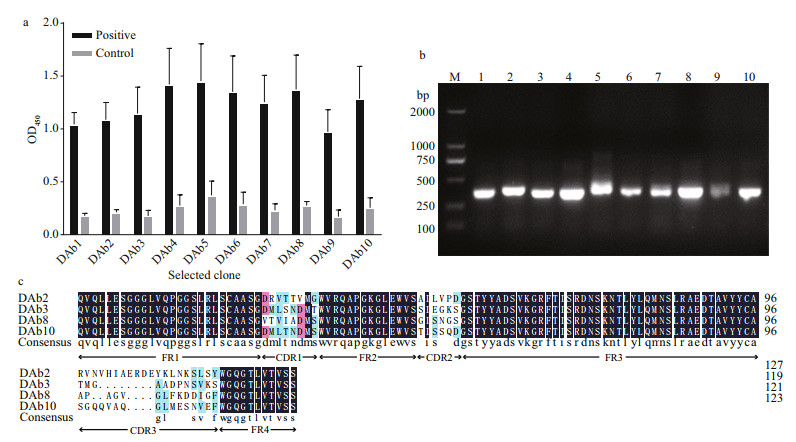
3 RESULT 3.1 Screening and identification of Microcystis particlesComparing the third round enriched library to the original, the OD450 value was increased approximately 6.3-fold by polyclonal phage ELISA (Fig. 1). The result indicated that the target particles were enriched obviously. After three rounds of biopanning, about 960 clones (10×96-well plates) were analyzed by the phage ELISA and 10 positive particles whose OD450 ratios of P/N > 3 were obtained. The ratio of the most positive particle named DAb3 was 6.43 and the lowest named DAb5 was 3.90 (Fig. 2a). All positive particles displayed a real and similar size (approximately 400 bp) gene fragments by colony PCR on a gel (Fig. 2b). Then the particles were checked by DNA sequencing, four (DAb2, DAb3, DAb8, and DAb10) of which possessed different amino acid sequences of the nanobodies displayed the same immunoglobulin frameworks from FR1 to FR4, but more or less differences in the complementarity determining regions from CDR1 to CDR3 (Fig. 2c).
|
| Fig.1 Polyclonal phage ELISA based on each round of the enriched library (quantified to 109-CFU/mL phage particles) for protein lysates of M. aeruginosa PCC7806 The positive coated antigen was protein lysates, while the negative control was CBS. The OD450 values were the means±SD from triplicate measurements. |
|
| Fig.2 Monoclonal phage ELISA that the selected phage particles for protein lysates of M. aeruginosa PCC7806 (a); the colony PCR amplified products of the ten positive monoclonal phage antibody genes, which were named DAb1–10 (b); amino acid sequences of the obtained positive monoclonal phage antibodies (c) In (a), during the test, the positive coated antigen was protein lysates and the negative control was CBS. The OD450 values were the means±SDs of triplicate measurements. In (b), lane M: DNA marker; lanes 1-10: correspond to the DAb1-10, respectively. In (c), the immunoglobulin frameworks (FR1-4) and the complementarity determining regions (CDR1-3) were indicated by the NCBI database. |
In this study, the most two positive antibodies (DAb2 and DAb3) were chosen and expressed in E. coli BL21, the expressed antibodies were abundant in the supernatant of whole cell lysate by ultrasonic breaking (Fig. 3a). Consequently, the purified DAb2 and DAb3 antibodies were obtained from the supernatant of whole cell lysates by using a His60 Ni superflow resin & gravity column, which existed in a clear protein band (approximately 14.5 or 12 kDa) on SDS-PAGE gal as shown in Fig. 3a. The concentration of the purified DAb2 and DAb3 in PBS solution were 506 and 972 μg/mL, i.e., 3.54 and 6.80 mg/L in the 500-mL original culture.

|
| Fig.3 SDS-PAGE analysis of the two most positive nanobody proteins (DAb2 and DAb3) expressed in E. coli BL21 (a); the Western blot (WB) using purified nanobodies DAb2 (b) and DAb3 (c) as the detection antibodies In (a), lane M: protein marker; lane 1: total proteins of bacteria without IPTG; lane 2: total proteins of bacteria with 0.2-mmol/L IPTG; lane 3: the supernatant of whole cell lysate by ultrasonic breaking; lane 4: purified nanobody proteins from the supernatant of whole cell lysate. SDS-PAGE and the Western blot analysis of protein extracts from M. aeruginosa PCC7806. In (b & c), immunoreactivity assay was performed using decreasing concentrations of protein extracts from M. aeruginosa PCC7806. Lane M: protein marker; lanes 1-3: 20-, 10-, and 5-μg protein extracts of M. aeruginosa PCC7806, respectively. |
The immunoreactivity of purified antibodies was examined by Western blot. As shown in Fig. 3b & c, both DAb2 and DAb3, exhibited positive immunoreactivity with M. aeruginosa PCC7806 polypeptides and they probably recognize the same antigen, only the difference in sensitivity modifies the apparent distribution pattern.
3.3 Antibodies specificity and sensitivity of the immuno-dot blot assayThe DAb3 did not react with any algae examined in this study, therefore the DAb2 was used for all subsequent immunoassays. As shown in Fig. 4a, the DAb2 bound efficiently to all cultured Microcystis strains (M. aeruginosa, M. flos-aquae, M. panniformis, and M. wesenbergii) and the field water sample. However, it exhibited an extremely weak signal for Synechocystis sp. PCC6803 and Chroococcus sp., no cross-reaction with the other cyanobacterial and green algal strains (Anabaena sp. JK12, Chlorella sp., Pediastrum duplex FACHB-1804, and Scenedesmus sp. JK37). The detection limit of the immuno-dot blot assay was less than 240 cells of M. aerginosa PCC7806 (Fig. 4b).

|
| Fig.4 Various algae lysates were tested in the immuno-dot blot test to check the specificity of antibody DAb2 (a); detection limit of the immuno-dot blot assay (b) In (a), Dot 1: elution buffer (PBS); Dot 2: M. aeruginosa PCC7806; Dot 3: M. aeruginosa FACHB-905; Dot 4: Microcystis XW01; Dot 5: field water sample; Dot 6: M. flos-aquae FACHB-1272; Dot 7: M. panniformis FACHB-1757; Dot 8: M. wesenbergii FACHB-929; Dot 9: Synechocystis sp. PCC6803; Dot 10: Chroococcus sp. FACHB-193; Dot 11: Anabaena sp. JK12; Dot 12: Chlorella sp.; Dot 13: Pediastrum duplex FACHB-1804; Dot 14: Scenedesmus sp. JK37. The detection limit of the assay was performed with the proteins of serial dilutions of M. aeruginosa PCC7806 cells (3×104-2.4×102). Proteins were extracted with 50-μL PBS using a high-throughput tissue grinder and 5-μL lysates were spotted onto immobilon-P transfer membrane. The membrane was incubated with the DAb2 and without the DAb2 as a control. |
Microcystis identification is mainly based on morphological characteristics determined by the microscopic method. It is a time-consuming and arduous process that requires specialist taxonomic knowledge. The immunological method could quickly and reliably identify Microcystis species and require only small sample volumes and a simple sample preparation based on a short lysis step or working with intact cells (Blanco et al., 2015, 2017; Folorunsho et al., 2021).
Polyclonal antibodies have been widely used. Blanco et al.(2015, 2017) have successfully produced three polyclonal antibodies against M. aeruginosa, M. flos-aquae, and M. novacekii. However, the drawbacks of polyclonal antibody including burdensome immune programs and difficulty in reproducing should be unavoidable. Additionally, when animals injected with toxic cyanobacterial cells or lysates, the target antibodies could not be got due to the toxicity of antigen (Gas et al., 2009). Phage display antibody library has become a promising technology to rapidly obtain specific antibodies for any antigen, independently of its toxicity or immunogenicity by a fast in vitro high-throughput screening without immunization (Xu et al., 2018). In this study, we selected specific nanobodies against Microcystis by panning in vitro a human domain antibody display library and then conveniently converted them into applicable reagents for diagnostics. Our goal is to mass-produce a genetic engineering antibody, which could provide a guarantee for the development of immune methods for Microcystis detection.
The species selectivity of the anti-Microcystis nanobody (the DAb2) produced in this work enabled to use it for the detection of Microcystis cultivated in the laboratory and grown in the field. Although the DAb2 could recognize all Microcystis tested in this study, it should be emphasized the affinity of the DAb2 varied among the different Microcystis species. It exhibited higher immunoreactivity with M. aeruginosa strains (PCC7806 and FACHB-905), M. flos-aquae FACHB-1272, M. panniformis FACHB-1757, and Microcystis XW01 than M. wesenbergii FACHB-929 (Fig. 4a). Previous studies showed that M. wesenbergii can be distinguished from other Microcystis because of the significant gel sheath of colonies and divergence in the cpcBA-IGS sequence (Komárek and Komárková, 2002; Tan et al., 2010). The results indicated that M. wesenbergii could have a distant evolutionary relationship with other Microcystis species. The DAb2 weakly recognized Synechocystis sp. PCC6803 and Chroococcus sp., both of which belong to the same Chroococcales as Microcystis. The faint signal could be exploited as a negative control in the detection of Microcystis by the immuno-dot blot assay. Recently, Folorunsho et al. (2021) have selected three specific nanobodies against M. aeruginosa by whole-cell biopanning from a naïve phage display library. However, whether the antibodies reacted with the other Microcystis species were not clear. The DAb2 could be more effectively applied for simultaneous detection of Microcystis species.
The immuno-dot blot assay based on the DAb2 showed that 240 cells of M. aeruginosa PCC7806 displayed an obvious signal, it is inferred from the signal that the detection limit could be less than the 200 cells, similar to Blanco's polyclonal antibody (the detection limit was about 102–103 cells) (Blanco et al., 2015). However, it existed a big gap compared with the records by Folorunsho et al. (2021) that reached to 1.1 and 2.2 cells/mL for the strains M. aeruginosa PCC7806 and PCC7005. To improve the detection sensitivity, some signal amplification strategies would be introduced, such as using antibody-conjugated AuNPs (Thiruppathiraja et al., 2011) and fluorescent labeled-antibody (Yu et al., 2011; Niu et al., 2012). In addition, more sensitive detection methods, such as flow cytometry based on fluobodies (Mazzega et al., 2019), thermal lens spectrometry (TLS) diagnostic method (Folorunsho et al., 2021), and electrochemical immunodetection (Adkins et al., 2017) could be used in the immune detection of Microcystis species.
5 CONCLUSIONIn this study, we have screened a specific nanobody DAb2 against Microcystis from the human domain antibody display library. The DAb2 intensively recognized M. aeruginosa, M. flos-aquae and M.panniformis, and weakly recognized M.wesenbergii by the immuno-dot blot assay. The antibody could be used for Microcystis immunodetection.
6 DATA AVAILABILITY STATEMENTThe data generated in this study are available from the corresponding author or the first author upon reasonable request.
Adkins J A, Boehle K, Friend C, et al. 2017. Colorimetric and electrochemical bacteria detection using printed paper-and transparency-based analytic devices. Analytical Chemistry, 89(6): 3613-3621.
DOI:10.1021/acs.analchem.6b05009 |
Bhardwaj J, Kim M W, Jang J. 2020. Rapid airborne influenza virus quantification using an antibody-based electrochemical paper sensor and electrostatic particle concentrator. Environmental Science & Technology, 54(17): 10700-10712.
DOI:10.1021/acs.est.0c00441 |
Blanco Y, Moreno-Paz M, Parro V. 2017. Experimental protocol for detecting Cyanobacteria in liquid and solid samples with an antibody microarray chip. Journal of Visualized Experiments, 120: e54994.
DOI:10.3791/54994 |
Blanco Y, Quesada A, Gallardo-Carreño I, et al. 2015. CYANOCHIP: an antibody microarray for high-taxonomical-resolution cyanobacterial monitoring. Environmental Science & Technology, 49(3): 1611-1620.
DOI:10.1021/es5051106 |
Bradford M M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72(1-2): 248-254.
DOI:10.1016/0003-2697(76)90527-3 |
Carrera M, Garet E, Barreiro A, et al. 2010. Generation of monoclonal antibodies for the specific immunodetection of the toxic dinoflagellate Alexandrium minutum Halim from Spanish waters. Harmful Algae, 9(3): 272-280.
DOI:10.1016/j.hal.2009.11.004 |
Folorunsho O G, Oloketuyi S F, Mazzega E, et al. 2021. Nanobody-dependent detection of Microcystis aeruginosa by ELISA and thermal lens spectrometry. Applied Biochemistry and Biotechnology, 193(9): 2729-2741.
DOI:10.1007/s12010-021-03552-6 |
García-García A, Madrid R, González I, et al. 2020. A novel approach to produce phage single domain antibody fragments for the detection of gluten in foods. Food Chemistry, 321: 126685.
DOI:10.1016/j.foodchem.2020.126685 |
Gas F, Pinto L, Baus B, et al. 2009. Monoclonal antibody against the surface of Alexandrium minutum used in a whole-cell ELISA. Harmful Algae, 8(3): 538-545.
DOI:10.1016/j.hal.2008.08.027 |
Guiry M D, Guiry G M. 2019. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org. Accessed on 2019-12-1.
|
Jiang W Z, Cossey S, Rosenberg J N, et al. 2014. A rapid live-cell ELISA for characterizing antibodies against cell surface antigens of Chlamydomonas reinhardtii and its use in isolating algae from natural environments with related cell wall components. BMC Plant Biology, 14(1): 244.
DOI:10.1186/s12870-014-0244-0 |
Jiang W Z, Rosenberg J N, Wauchope A D, et al. 2013. Generation of a phage-display library of single-domain camelid VHH antibodies directed against Chlamydomonas reinhardtii antigens, and characterization of VHHs binding cell-surface antigens. The Plant Journal, 76(4): 709-717.
DOI:10.1111/tpj.12316 |
Komárek J, Komárková J. 2002. Review of the European Microcystis-morphospecies (Cyanoprokaryotes) from nature. Czech Phycology, 2: 1-24.
|
Lee J, Kim J H, Kim B N, et al. 2020. Identification of novel paraben-binding peptides using phage display. Environmental Pollution, 267: 115479.
DOI:10.1016/j.envpol.2020.115479 |
Li M, Zhu W, Gao L, et al. 2013. Changes in extracellular polysaccharide content and morphology of Microcystis aeruginosa at different specific growth rates. Journal of Applied Phycology, 25(4): 1023-1030.
DOI:10.1007/s10811-012-9937-7 |
Mazzega E, Beran A, Cabrini M, et al. 2019. In vitro isolation of nanobodies for selective Alexandrium minutum recognition: a model for convenient development of dedicated immuno-reagents to study and diagnostic toxic unicellular algae. Harmful Algae, 82: 44-51.
DOI:10.1016/j.hal.2019.01.002 |
Mendoza H, López-Rodas V, González-Gil S, et al. 1995. The use of polyclonal antisera and blocking of antibodies in the identification of marine dinoflagellates: species-specific and clone-specific antisera against Gymnodinium and Alexandrium. Journal of Experimental Marine Biology and Ecology, 186(1): 103-115.
DOI:10.1016/0022-0981(94)00160-F |
Mitra M, Dewez D, García-Cerdán J G, et al. 2012. Polyclonal antibodies against the TLA1 protein also recognize with high specificity the D2 reaction center protein of PSII in the green alga Chlamydomonas reinhardtii. Photosynthesis Research, 112(1): 39-47.
DOI:10.1007/s11120-012-9733-x |
Niu W M, He E Q, Wu Q G, et al. 2012. Use of fluorescent europium chelates as labels for detection of microcystin-LR in Taihu Lake, China. Journal of Rare Earths, 30(9): 941-946.
DOI:10.1016/S1002-0721(12)60158-6 |
Otsuka S, Suda S, Li R H, et al. 2000. Morphological variability of colonies of Microcystis morphospecies in culture. The Journal of General and Applied Microbiology, 46(1): 39-50.
DOI:10.2323/jgam.46.39 |
Qiu Y L, Li P, Dong S, et al. 2018. Phage-mediated competitive chemiluminescent immunoassay for detecting Cry1Ab toxin by using an anti-idiotypic camel nanobody. Journal of Agricultural and Food Chemistry, 66(4): 950-956.
DOI:10.1021/acs.jafc.7b04923 |
Sun Q Q, Zhu W, Li M, et al. 2016. Morphological changes of Microcystis aeruginosa colonies in culture. Journal of Limnology, 75(1): 14-23.
DOI:10.4081/jlimnol.2015.1225 |
Szkola A, Linares E M, Worbs S, et al. 2014. Rapid and simultaneous detection of ricin, staphylococcal enterotoxin B and saxitoxin by chemiluminescence-based microarray immunoassay. Analyst, 139(22): 5885-5892.
DOI:10.1039/C4AN00345D |
Tan W H, Liu Y, Wu Z X, et al. 2010. CpcBA-IGS as an effective marker to characterize Microcystis wesenbergii (Komárek) Komárek in Kondrateva (cyanobacteria). Harmful Algae, 9(6): 607-612.
DOI:10.1016/j.hal.2010.04.011 |
Thiruppathiraja C, Kamatchiammal S, Adaikkappan P, et al. 2011. An advanced dual labeled gold nanoparticles probe to detect Cryptosporidium parvum using rapid immuno-dot blot assay. Biosensors and Bioelectronics, 26(11): 4624-4627.
DOI:10.1016/j.bios.2011.05.006 |
Wang Y, Zhang X, Zhang C Z, et al. 2012. Isolation of single chain variable fragment (scFv) specific for Cry1C toxin from human single fold scFv libraries. Toxicon, 60(7): 1290-1297.
DOI:10.1016/j.toxicon.2012.08.014 |
Waterbury J B. 2006. The cyanobacteria-isolation, purification and identification. In: Dworkin M, Falkow S, Rosenberg E et al eds. The Prokaryotes. Springer, New York. p. 1053-1073, https://doi.org/10.1007/0-387-30744-3_38.
|
West N J, Bacchieri R, Hansen G, et al. 2006. Rapid quantification of the toxic alga Prymnesium parvum in natural samples by use of a specific monoclonal antibody and solid-phase cytometry. Applied and Environmental Microbiology, 72(1): 860-868.
DOI:10.1128/AEM.72.1.860-868.2006 |
Xiao M, Li M, Reynolds C S. 2018. Colony formation in the cyanobacterium Microcystis. Biological Reviews, 93(3): 1399-1420.
DOI:10.1111/brv.12401 |
Xu C X, He D, Zu Y, et al. 2021. Microcystin-LR heterologous genetically engineered antibody recombinant and its binding activity improvement and application in immunoassay. Journal of Hazardous Materials, 406: 124596.
DOI:10.1016/j.jhazmat.2020.124596 |
Xu C X, Liu X Q, Liu Y, et al. 2019a. High sensitive single chain variable fragment screening from a microcystin-LR immunized mouse phage antibody library and its application in immunoassay. Talanta, 197: 397-405.
DOI:10.1016/j.talanta.2019.01.064 |
Xu C X, Miao W J, He Y, et al. 2019b. Construction of an immunized rabbit phage display antibody library for screening microcystin-LR high sensitive single-chain antibody. International Journal of Biological Macromolecules, 123: 369-378.
DOI:10.1016/j.ijbiomac.2018.11.122 |
Xu C X, Yang Y, Liu L W, et al. 2018. Microcystin-LR nanobody screening from an alpaca phage display nanobody library and its expression and application. Ecotoxicology and Environmental Safety, 151: 220-227.
DOI:10.1016/j.ecoenv.2018.01.003 |
Yu H W, Jang A, Kim L H, et al. 2011. Bead-based competitive fluorescence immunoassay for sensitive and rapid diagnosis of cyanotoxin risk in drinking water. Environmental Science & Technology, 45(18): 7804-7811.
DOI:10.1021/es201333f |
Zhang W Q, Lin M Q, Yan Q, et al. 2021. An intracellular nanobody targeting T4SS effector inhibits Ehrlichia infection. Proceedings of the National Academy of Sciences, 118(18): e2024102118.
DOI:10.1073/pnas.2024102118 |
Zhang X, Liu Y, Zhang C Z, et al. 2012. Rapid isolation of single-chain antibodies from a human synthetic phage display library for detection of Bacillus thuringiensis (Bt) Cry1B toxin. Ecotoxicology and Environmental Safety, 81: 84-90.
DOI:10.1016/j.ecoenv.2012.04.021 |
Zhao A Z, Tohidkia M R, Siegel D L, et al. 2016. Phage antibody display libraries: a powerful antibody discovery platform for immunotherapy. Critical Reviews in Biotechnology, 36(2): 276-289.
DOI:10.3109/07388551.2014.958978 |
Zhu Y Y, Wu J J X, Han L J, et al. 2020. Nanozyme sensor arrays based on heteroatom-doped graphene for detecting pesticides. Analytical Chemistry, 92(11): 7444-7452.
DOI:10.1021/acs.analchem.9b05110 |
 2022, Vol. 40
2022, Vol. 40


