Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZENG Boxin, DONG Taiwei, XIA Yanting, YANG Shun, HUANG Mengmeng, FEI Hui
- Characterization and comparison of two C-type lectins with novel key motifs from mud crab Scylla paramamosain
- Journal of Oceanology and Limnology, 40(4): 1628-1639
- http://dx.doi.org/10.1007/s00343-021-1153-3
Article History
- Received Jun. 9, 2021
- accepted in principle Jul. 21, 2021
- accepted for publication Sep. 6, 2021
2 Zhejiang Provincial Key Laboratory of Silkworm Bioreactor and Biomedicine, Zhejiang Sci-Tech University, Hangzhou 310018, China
Invertebrate defense against pathogens depends on innate immunity because of the lack of acquired immunity (Loker et al., 2004; Rowley and Powell, 2007). The innate immune system recognizes and responds to different types of pathogens, depending on the common structural and functional characteristics of microorganisms. As the first line of defense, the innate immune system keeps the host from infection and its defense function enables the host to effectively resist bacteria, fungi, and viral pathogens (Zhang et al., 2019; Tran et al., 2020). The conserved determinants of the origin of microorganisms and viruses are detected by the receptors of the innate immune system, initiate the activation of the signal cascade, which leads to the transcriptional regulation of inflammatory mediators that play a role in pathogen control and clearance, and ultimately leads to an effective immune response (Brubaker et al., 2015; Tran et al., 2020).
Based on the difference of molecular structure, animal lectins can be mainly classified into C-type lectins (CTL), P-type lectins, I-type lectins, S-type lectins, pentamer proteins, and so on (Drickamer and Taylor, 1993). In the lectin superfamily, C-type lectins play an important role in identifying and clearing foreign invasions and have attracted more and more attention in recent years (Christophides et al., 2002). C-type lectins are a kind of protein superfamily that needs Ca2+ to participate in selective carbohydrate binding (Weis et al., 1998). They are classified into 17 groups according to their structure and functions. Collectins mainly serve as pattern recognition receptor (PRR) participating in discrimination of pathogens (Cambi et al., 2005), while another group selectins mediate cellular adhesion to active opsonization (Patel et al., 2002). CTL has at least one characteristic carbohydrate recognition domain (CRD), which are composed of 115–130 amino acids (Drickamer, 1993; Zelensky and Gready, 2003). Glu-Pro-Asn (EPN)/Gln-Pro-Asp (QPD) and Trp-Asn-Asp (WND) are two highly conserved amino acid motifs in this domain, which are related to the specific recognition of carbohydrates (Weis et al., 1998). C-type lectins play an important role in invertebrate innate immune response, including activating prophenoloxidase (Christophides et al., 2002), microbial agglutination (Wang et al., 2009), inhibiting bacterial growth, and promoting blood cell phagocytosis. For example, Scylla paramamosain SpCTL-B can bind to a variety of Gram-positive bacteria, Gram-negative bacteria, and fungi (Wei et al., 2018).
Scylla paramamosain is an important mariculture crab in China, with high nutritional value, medicinal value, and economic value (Cao et al., 2013). However, in the process of cultivation, it is easy to be affected by water quality, bacteria, and virus infections, which leads to the low survival rate and causes huge losses to aquaculture of S. paramamosain. For example, bacterial infections such as Vibrio parahaemolyticus and Aeromonas hydrophila in 2010 led to a nearly 60% drop in the production of crabs in Shantou, China. Although there have been many coping strategies such as immune enhancers, how to resist increasingly serious diseases by improving the immunity of mud crabs has become an urgent problem in the crab farming industry.
In this study, we have identified and verified the functions of SpCTL-C and SpCTL-D derived from S. paramamosain. The pathogen-associated molecular patterns (PAMPs) binding activity, antibacterial activity, bacterial binding activity, blood coagulation activity, opsonization enhancement activity of blood cells, and the tissue distribution of the two lectins were studied systematically using molecular biology and immunological techniques, and preliminary exploration of its molecular recognition mechanism.
2 MATERIAL AND METHOD 2.1 MaterialEscherichia coli, Micrococcus luteus, and Staphylococcus aureus were purchased from the Microbial Culture Collection Center (Beijing, China). Vibrio fluvialis, V. parahaemolyticus, and Saccharomyces cerevisiae were obtained from the Global Bioresource Center (USA). Pichia pastoris was from Invitrogen. Aeromonas sobria, Edwardsiella tarda, and A. hydrophila were isolated by our laboratory. Vibrio alginolyticus were obtained from Minnan Normal University.
Scylla paramamosain were collected from a seafood market in Hangzhou, Zhejiang Province. Filtered seawater was used to acclimate the crabs for 7 days before experiments.
2.2 Method 2.2.1 Bioinformatics analysisThe cDNA and deduced amino acid sequences of SpCTL-C and SpCTL-D were analyzed using BLAST (http://www.ncbi.nlm.nih.gov/blast) and the Expert Protein Analysis System (http://www.expasy.org). ExPASy (http://web.expasy.org/computepi/) was used to predict the molecular weight and isoelectric point (pI) of SpCTL-C and SpCTL-D. The ClustalW Multiple Alignment program (http://www.ebi.ac.uk/clustalw/) was used to create a multiple sequence alignment. The signal peptide was predicted by the SignalP 3.0 server (http://www.cbs.dtu.dk/services/SignalP-3.0/). MEGA 7.0 was used to produce the phylogenetic tree, and the neighbor-joining (NJ) method was used for phylogenetic analysis.
2.2.2 Tissue distribution of SpCTL-C, SpCTL-D, and RNA extraction and cDNA synthesis of S.paramamosainTotal RNA was extracted from various tissues (muscle, gill, hemocytes, gonad, and hepatopancreas) of four randomly sampled individuals. Total RNA was isolated using TRIzol reagent (Sangon Biotech). The first strand synthesis was carried out based on Promega M-MLV RT usage information using the DNase I-treated total RNA as template and oligo (dT)-adaptor primer. The reactions were incubated at 42 ℃ for 1 h, terminated by heating at 95 ℃ for 5 min. The amplified cDNA was diluted and stored at -80 ℃.
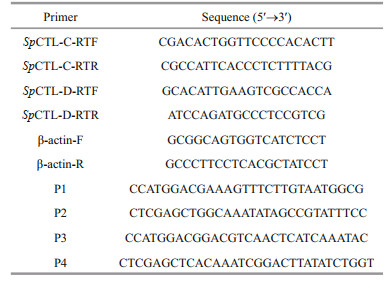
The tissue distribution and temporal expression of SpCTL-C and SpCTL-D transcript in different tissues were determined by quantitative real-time PCR (Q-PCR) as previously described (Huang et al., 2017). Specific primers of SpCTL-C, SpCTL-D, and internal reference gene β-actin were designed (Table 1). The reaction system was configured according to the instructions of the kit. The procedure was as follows: 95 ℃ for 1 min, followed by 40 cycles (95 ℃ for 5 s, 60 ℃ for 30 s). All samples were analyzed in four duplications and the data obtained from Q-PCR analysis were subjected to one-way analysis of variance (one-way ANOVA) followed by an unpaired, two-tailed t-test. A probability of P < 0.05 was considered statistically significant.
The cDNA fragments of SpCTL-C, SpCTL-D were amplified with primers P1–P4 (Table 1) with two restriction enzyme cleavage sites at 5′-ends (NcoI and XhoI). Target genes were digested with restriction enzymes and ligated to expression vector pET-32a. The recombinant plasmids (pET-32a-SpCTL-C and pET-32a-SpCTL-D) were transformed into antibiotic competent cells in E. coli DH5α. The SpCTL-C and SpCTL-D positive transformants were inoculated into LB medium containing a final concentration of 100-μg/mL ampicillin, and incubated at 37 ℃ with a shaker rotating 220 r/min until the absorbance at 600 nm (OD600) reached 0.4–0.6. Isopropyl-beta-D-thiogalactopyranoside (IPTG) was added to the LB medium to a final concentration of 0.01 mol/L, and cultured in a shaker at 18 and 30 ℃ converted to 220 r/min for 20 h and 4 h, respectively. After the cells were obtained by centrifugation, they were ultrasonically processed and centrifuged to obtain the soluble recombinant protein in the supernatant. Recombinant rSpCTL-C and rSpCTL-D were purified by Ni2+ chelating agarose column and eluted with 100 and 250-mol/L imidazole in tris buffered saline (TBS) buffer, respectively. After dialysis to TBS buffer, recombinant proteins were concentrated with PEG-20000, a certain concentration of recombinant protein was obtained. At the same time, recombinant thioredoxin (rTrx) was purified according to the same method. The obtained rSpCTL-C, rSpCTL-D, and rTrx were stored at -80 ℃ for the subsequent use in the experiments.
2.2.4 PAMPS binding experiments of rSpCTL-C and rSpCTL-D based on enzyme-linked immuno sorbent assay (ELISA)PAMPs binding assays were performed as previously described with some modifications (Yu et al., 2007). An amount of 10-μg peptidoglycan (PGN), glucan (Glucan), and lipopolysaccharide (LPS) in 100-μL carbonate-bicarbonate buffer (pH=9.6) were used to coat a 96-well microliter plate (Costar) and incubated at 4 ℃ overnight, respectively. The incubated PAMPs were poured out and washed with phosphate buffered saline-Tween (PBS-T) (pH 7.4) for three times. Nonspecific binding to the wells was prevented by addition 3% bovine serine albumin (BSA) at 37 ℃ for 1 h. Then the plate was washed by PBS-T for three times, and 100-μL rSpCTL-C or rSpCTL-D were added to each well using two-fold dilution method (Ca2+ concentration 10 mol/L). The same concentration of rTrx was set as a negative control. Both of them were incubated at 18 ℃ for 3 h. After three times washing with PBS-T, 100-μL horseradish peroxidase (HRP)-labelled rabbit anti-6 × His-tag polyclonal antibody (Abcam, diluted 1꞉4 000 in PBS-T) was added to each well and incubated at 37 ℃ for 1 h. After three times washing, detection was performed following the instructions of the El-TMB Chromogenic Reagent kit (Sangon). The absorbance under OD450 for detection was measured. The filled wells with 100-μL TBS (pH=8.0) were used as blanks. Three replicates were set for each group.
2.2.5 Microbial binding experiments of rSpCTL-C and rSpCTL-DThe experimental bacteria (M. luteus, S. aureus, V. fluvialis, V. parahaemolyticus, S. cerevisiae, A. sobria, V. alginolyticus, E. tarda, and A. hydrophila) were cultured and resuspended to a final concentration at 1×108 CFU. Then they were used to coat a 96-well microliter plate (Costar) and incubated at 4 ℃ overnight. After rinsing the plates, 200-μL BSA solution was added to each well and sealed at 37 ℃ for 2 h. An aliquot of 100-μL recombinant protein (Ca2+ concentration 10 mol/L) was added to each well and incubated at 18 ℃ for 3 h after three times rinse. After washing the plates, 100-μL HRP labeled rabbit anti-His tag antibody was added to each well (diluted with 1꞉3 000), and incubated at 37 ℃ for 1 h. After rinsing the board for three times, color development was carried out according to the instructions of EL-TMB kit, and absorbance was measured under 450 nm (OD450).
2.2.6 rSpCTL-C and rSpCTL-D microbial agglutination experimentsThe experimental bacteria (S. aureus, M. luteus, V. fluvialis, A. sobria, P. pastoris, and S. cerevisiae) were cultured to the logarithmic growth stage, and then washed three times with 0.1-mol/L NaHCO3 buffer (pH=9.0) after centrifugation. The bacteria were mixed with 0.1-mg/mL FITC dye and incubated for 2 h at room temperature for staining. After cleaning excess dye, FITC-labeled microbes were resuspended to TBS buffer (50-mol/L TBS, 10-mol/L CaCl2) with concentration of 1×1010 cell/mL. An aliquot of 25-μL rSpCTL-C and rSpCTL-D were incubated with 10-μL bacterial suspension respectively in dark for 1 h at room temperature. The agglutination activity of rSpCTL-C and rSpCTL-D were observed under the fluorescence microscope. rTrx of the same concentration was used as negative control.
2.2.7 Encapsulation experiment of rSpCTL-C and rSpCTL-DNi-NTA agarose particles were equilibrated in 5-mol/L TBS-Ca2+ buffer, incubated with rSpCTL-C, rSpCTL-D, and rTrx, respectively, shaken overnight at 4 ℃. After melting with 1% agarose, the 48-well cell culture plate was coated, and the extracted hemocytes of carb were added to each well separately, and the cells were settled for 10 min. Then three protein-coated agarose particles (120–150) were added into each well, and three groups of each protein were set in parallel, incubated at 18 ℃ for 6 h, and the encapsulation was observed under the optical microscope. The encapsulation rate = (the number of encapsulated beads/all beads tested)×100%. The data are expressed in means±SE (n=3).
2.2.8 Phagocytosis-promoting experiment of rSpCTL-C and rSpCTL-DThe experimental bacteria (M. luteus, E. coli) were cultured to logarithmic growth phase and after centrifugation were washed for three times with 0.1-mol/L NaHCO3 (pH=9.0). The bacteria were stained with 0.1-mg/mL FITC and incubated at room temperature for 2 h. After washing the excess stain, bacteria were resuspended with TBS-Ca2+ to a bacterial concentration of 1×1010 cells/mL. The blood cells were re-suspended in TBS group, negative control rTrx group, and rSpCTL-C, rSpCTL-D group according to the final concentration of 5×107 /mL. M. luteus and E. coli with the final concentration of 1×1010 /mL was added and incubated at room temperature for 1 h. Flow cytometry (AccuriTM C6 Plus, BD biosciences, USA) was used to detect the phagocytosis rate of blood cells and the phagocytosis percentage was defined as (the hemocytes ingesting bacteria/all hemocytes tested) ×100%. All Assay was performed in triplicate. The data are expressed as means ± SE (n=3).
2.2.9 Antibacterial activity assay of rSpCTL-C and rSpCTL-DVarious microbes (V. parahaemolyticus, V. fluvialis, P. pastoris, S. cerevisiae, M. luteus, and V. alginolyticus) were cultured to the logarithmic growth phase and were washed 3 times with TBS-Ca2+ and resuscitated to 104 CFU. 50 μL the same concentration of rSpCTL-C and rSpCTL-D were mixed with bacterial suspension at 1꞉1 and incubated at room temperature for 1 h. 50-μL TBS plus bacterial solution was used as blank control, and 50-μL rTrx was used as negative control. Take 20-μL mixture to 96-well cell culture plate, add 200-μL corresponding culture medium, incubate in a constant temperature shaker for 12–16 h until reaching the platform stage and read OD600 every half an hour. The microbial growth curve was analyzed and drawn by Origin 8.0 software and Excel.
2.2.10 Statistical analysisAll experimental data were analyzed using one-way analysis of variance (one-way analysis of variance, ANOVA). The expression level was analyzed using the software GraphPad Prims 8.0. The results are expressed as mean±SE. P < 0.05 means significant difference, and P < 0.01 means extremely significant difference.
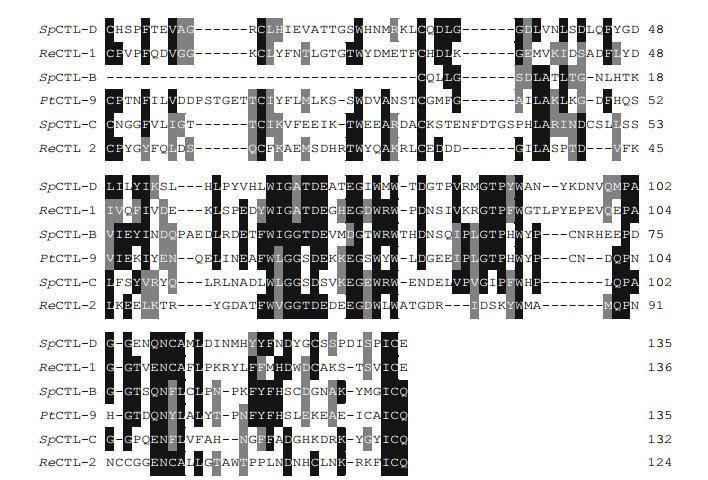
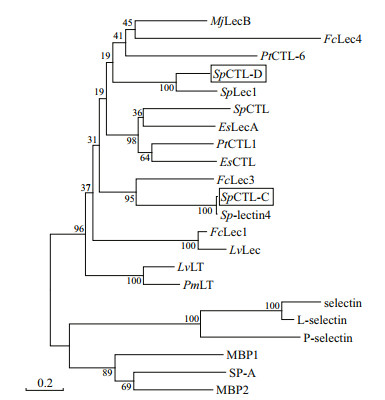
3 RESULT 3.1 cDNA cloning and sequence analysis of SpCTL-C and SpCTL-DThe cDNA sequences of SpCTL-C and SpCTL-D were screened from GenBank with accession numbers KC757381.1 and HQ325748.1, respectively. The open reading frame (ORF) of SpCTL-C encodes 158 amino acids. The ORF of SpCTL-D encodes 176 amino acids. Both of them contains a single CRD. The predicted theoretical isoelectric point and molecular weight of SpCTL-C are 5.29 and 17.775 kDa, while SpCTL-D is 4.89 and 19.698 kDa, respectively. According to multiple sequence alignments, SpCTL-C and SpCTL-D both contain CRDs which have two pairs of disulfide bonds formed by four highly conserved cysteines maintaining the stability of the structure. The key motif determining the carbohydrate binding specificity was QPA and MPA in SpCTL-C and SpCTL-D respectively. Multiple sequence alignment analysis of SpCTL-C and SpCTL-D against the CTLs from various species show that they differ in amino acid composition (Fig. 1). Figure 1 shows that some amino acid residues, such as cysteine and tryptophan residues, are highly conserved in the CTLs of different species. A phylogenetic tree was constructed by using NJ algorithm to analyze the sequence alignment based on the alignment of C-type lectin amino acid sequences from S. paramamosain, Eriocheir sinensis, Penaeus japonicus, Penaeus vannamei, Portunus trituberculatus, Penaeus monodon, Homo sapiens, Gallus gallus, Salmo salar, Mus musculus, and two distinct groups were separated in the phylogenetic tree (Fig. 2).

|
| Fig.1 Multiple sequence alignment of CRD in SpCTL-C, SpCTL-D, and other CTLs The sequence of other CTLs is as follows: ReCTL-1 (MN230985.1, Rimicaris exoculata); SpCTL-B (LN994606, S. paramamosain); PtCTL-9 (MT882671, P. trituberculatus); ReCTL-2 (MN230986, R. exoculata). The relationships among the residues are indicated as follows: identical residues (white letters on a black background), similar residues (white letters on a grey background); and non-similar residues (black letters on a white background). |
|
| Fig.2 Phylogenetic analysis of SpCTL-C and SpCTL-D A thouthand bootstraps were performed on the NJ trees to check the repeatability of the results. The C-type lectin sequences used for phylogenetic analysis are as follows: MjLecB (ADG85668.1, P. japonicus); FcLec4 (AHA85979.1, Penaeus chinensis); PtCTL-6 (QGV11036.1, P. trituberculatus); SpLec1 (AEO92001.1, S. paramamosain); EsLecA (AFF59979.1, E. sinensis); SpCTL (AFU52948.1, S. paramamosain); PtCTL-1 (ARD09216.1, P. trituberculatus); EsCTL (AGC31447.1, E. sinensis); FcLec3 (ACJ06431.1, P. chinensis); Sp-lectin4 (AIC80996.1, S. paramamosain); FcLec1 (ABA54612.1, P. chinensis); LvLec (ABU62825.1, P. vannamei); LvLT (ABI97374.1, P. vannamei); PmLT (ABI97373.1, P. monodon); selectin (AAI09160.1, M. musculus); L-selectin (NP_035476.1, M. musculus); P-selectin (NP_002996.2, H. sapiens); MBP1 (NP_001134969.1, S. salar); SP-A (NP_075623.2, M. musculus); MBP2 (AAB94071.1, G. gallus); SpCTL-C and SpCTL-D. |
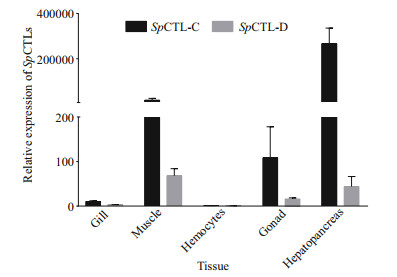
The mRNA transcripts of SpCTL-C and SpCTL-D were detected in gill, muscle, gonad, and hepatopancreas (Fig. 3). They were most abundant in hepatopancreas, moderately expressed in gonad and muscle, and relatively lower in gill and hemocytes. The expression level of SpCTL-C in hepatopancreas, gonad, muscle, and gill were 266 119-, 108.29-, 15 419-, and 10.13-fold relative to that in hemocytes, respectively. The expression level of SpCTL-D in hepatopancreas, gonad, muscle and gill were 43.82, 16.14-, 68.08-, and 2.89-fold relative to that in hemocytes, respectively.
|
| Fig.3 Real time expression of SpCTL-C and SpCTL-D The mRNA transcripts of SpCTL-C and SpCTL-D could be detected in gill, muscle, gonad, and hepatopancreas. Data are shown as mean ± SE (n=3). |
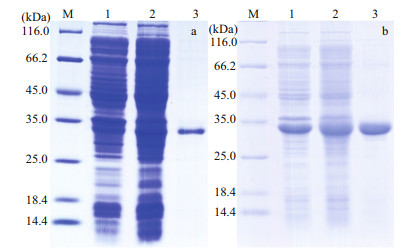
Prokaryotic expression recombinant plasmids pET-32a (+)-SpCTL-C and pET-32a (+)-SpCTL-D were transformed into E. coli BL21 respectively. After induction of expression by IPTG, their expression was detected using SDS-PAGE (Fig. 4). The results showed that rSpCTL-C was soluble expressed at 16 ℃ and the final induction concentration of IPTG was 0.01 mol/L, while rSpCTL-D was soluble expressed at 30 ℃ and the final induction concentration of IPTG was 0.01 mol/L.
|
| Fig.4 Expression and purification of recombinant SpCTL-C (a) and SpCTL-D (b) a. lane M: Protein molecular standard; lane 1: negative control of rSpCTL-C without IPTG induction; lane 2: expression of rSpCTL-C after IPTG induction; lane 3: purified rSpCTL-C; b. lane M: Protein molecular standard; lane 1: negative control of rSpCTL-D without IPTG induction; lane 2: expression of rSpCTL-D after IPTG induction; lane 3: purified rSpCTL-D. |
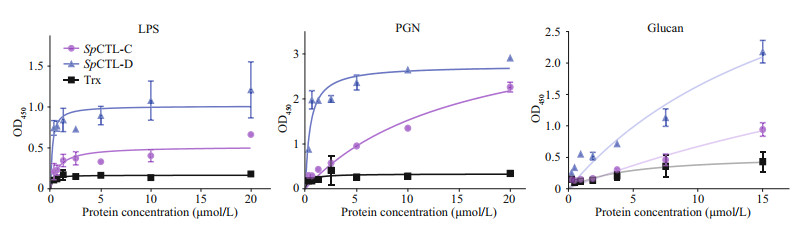
Both rSpCTL-C and rSpCTL-D had binding affinity to LPS, PGN, and Glucan. Meanwhile, rSpCTL-D had a higher binding affinity than that of rSpCTL-C, and no binding activity was detected in the negative control (Fig. 5).
|
| Fig.5 ELISA analysis of the interaction between rSpCTL-C, rSpCTL-D, and the PAMPs Plates were coated with three PAMPs, and then incubated with rSpCTL-C, rSpCTL-D, and rTrx with different concentration. After adding rabbit anti-6 × His-tag polyclonal antibodies (diluted to 1꞉4 000), follow the instructions of El-TMB Chromogenic Reagent to detect the interaction between rSpCTL-C, rSpCTL-D, rTrx, and PAMPs (Sangon) at 450 nm. |
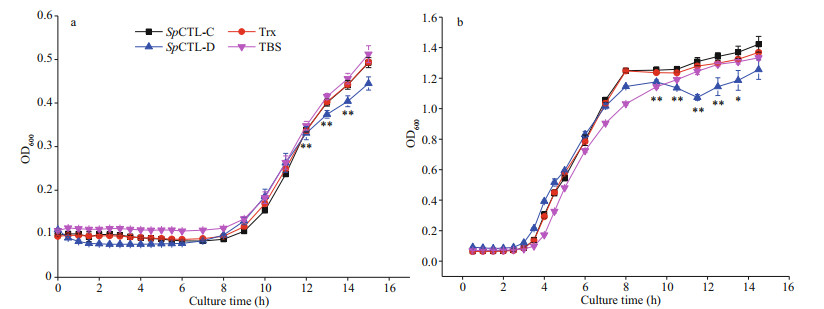
Comparing with the negative control and the blank control, rSpCTL-D had a certain inhibitory effect on the growth of V. fluvialis and M. luteus, while rSpCTL-C had no obvious inhibitory activity (Fig. 6).
|
| Fig.6 Antimicrobial activity analysis of rSpCTL-C and r SpCTL-D against V. fluvialis (a) and M. luteus (b) V. fluvialis and M. luteus were washed and suspended into same volume of rTrx, rSpCTL-C, and rSpCTL-D. The OD600 was recorded every 30 min to get the growth curve of different microbes. The results are shown as the mean of three individual measurements ± SE (n=3). ** indicates an extremely significant difference (P < 0.01) and * indicates a significant difference (P < 0.05). |
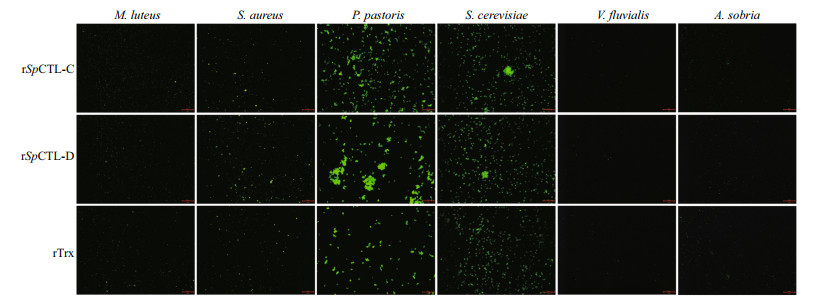
By incubating various microbes with rSpCTL-C and rSpCTL-D, the agglutination activity of rSpCTL-C and rSpCTL-D were observed. Both rSpCTL-C and rSpCTL-D showed agglutination activity towards fungi P. pastoris and S. cerevisiae and Gram-positive bacteria S. aureus (Fig. 7), but no agglutination was observed in other microorganisms (Fig. 7), indicating that rSpCTL-C and rSpCTL-D can selectively bind and agglutinate S. cerevisiae and P. pastoris.
|
| Fig.7 Bacteria agglutination of rSpCTL-C and r SpCTL-D on FITC-labeled V. fluvialis, A. sobria, S. aureus, M. luteus, S. cerevisiae, and P. pastoris rTrx protein was used as the negative control and TBS was used as the blank control. After incubating for 30 min in the dark, the agglutination was observed under the fluorescence microscope. Scale bars: 50 μm. |
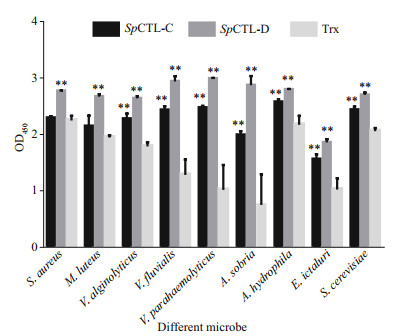
The microbe-binding spectrum of rSpCTL-C and rSpCTL-D was detected by ELISA. Comparing with negative control, rSpCTL-D can bind to all nine tested microorganisms, while rSpCTL-C can bind to seven microorganisms except two Gram-positive bacteria (S. aureus and M. luteus). This result indicates that rSpCTL-C and rSpCTL-D have broad-spectrum "non-self" binding activity, and the two proteins have certain selectivity for microbial recognition and binding. The binding activity of rSpCTL-C and rSpCTL-D to various microorganisms was significantly higher than that of the negative control group (P < 0.01) (Fig. 8).
|
| Fig.8 The microbe binding spectrum of rSpCTL-C and rSpCTL-D Results were obtained with ELISA. Plates that coated with microbes were incubated with rSpCTL-C and rSpCTL-D and rTrx. After incubation with rabbit anti-6 × His tag (HRP) polyclonal antibody (diluted to 1꞉3 000, Abcam), the interaction between recombinant proteins and microbes were detected following the instructions of TMB Chromogenic Reagent kit (Sangon) at 450 nm. Extremely difference (P < 0.01) is indicated with two asterisks. |
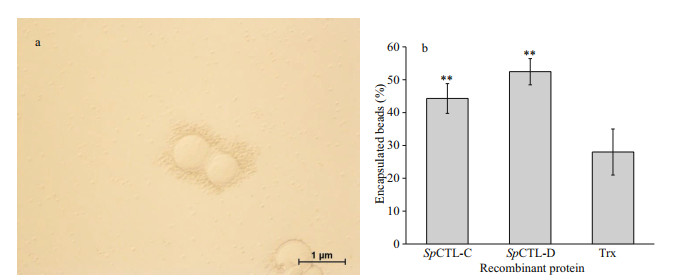
The encapsulation enhancement activity of rSpCTL-C and rSpCTL-D was carried out by incubating agarose gel beads with hemocytes. After microscopic examination and statistical analysis, the encapsulation percentage of rSpCTL-C and rSpCTL-D were 44.29% and 52.44% respectively (Fig. 9a–b), which were significantly higher than 28% of rTrx (P < 0.01). It indicated that both rSpCTL-C and rSpCTL-D had certain encapsulation-promoting activity, and rSpCTL-D had stronger encapsulation-promoting activity.
|
| Fig.9 The enhancement of hemocytes encapsulation by rSpCTL-C and rSpCTL-D a. amylose resin coated with rSpCTL-C, rSpCTL-D, or rTrx were encapsulated by S. paramamosain hemocytes; b. the percentage of beads encapsulated by hemocytes after 6-h incubation (n=3). Extremely significant difference (P < 0.01) is indicated with two asterisks. |
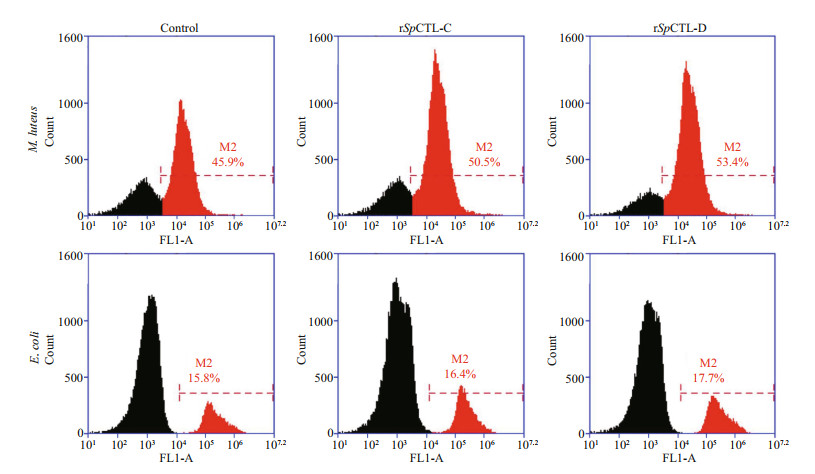
The phagocytosis rates of rSpCTL-C and rSpCTL-D proteins to M. luteus were 50.5% and 53.4% respectively, which were higher than those of the negative control group. The phagocytosis rates to E. coli were 16.4% and 17.7% respectively (Fig. 10). The results showed that rSpCTL-C and rSpCTL-D had better phagocytosis-promoting effect on M. luteus than the negative control, and had no obvious phagocytosis-promoting activity on E. coli.
|
| Fig.10 Promoting phagocytic activity of rSpCTL-C and rSpCTL-D hemocyte phagocytotic analysis Analysis of hemocytes in flow cytometry and the red side showed the hemocytes which endocytosed the FITC-labeled M. luteus and E. coli. |
Lectins are a kind of important pattern recognition receptors (PRRs), which mediate the recognition of non-autoantigens by binding to specific carbohydrate residues on the surface of pathogen cells. Lectins usually contains a CRD structure, which was made up of 120–130 amino acids. The primary structure determines the tertiary structure, although the lectins have different amino acid sequence, they contain a similar three-dimensional space structure, which was associated with the binding of carbohydrates (Taylor, 1993; Drickamer, 1999). Studies have shown that the carbohydrate recognition active site of C-type lectins is located in CRD. There are four Ca2+ binding sites in CRD, and Ca2+ binding site 2 is the key site for CRD to recognize and bind carbohydrates (Zelensky and Gready, 2005; Zhu et al., 2016). The key motif Glu-Pro-Asn (EPN) or Gln-Pro-Asp (QPD) are very conservative at this site. Usually the C-type lectin with EPN motif could recognize and bind to mannose while other C-type lectin with QPD motif could bind to galactose (Drickamer, 1999). However, the study of C-type lectins in invertebrates found that Ca2+ binding site 2 is diverse (Zelensky and Gready, 2003). Accordingly, the key motifs that determine the specificity of carbohydrate binding in SpCTL-C and SpCTL-D are Gln-Pro-Ala (QPA) and Met-Pro-Ala (MPA), respectively, which were newly found key motifs.
As the member of C-type lectin, both rSpCTL-C and rSpCTL-D have extensive PAMPs binding spectrum and bacterial binding spectrum, which can bind to PGN extracted from Gram-positive bacteria, LPS from Gram-negative bacteria, and β-glucan from yeast. At the same time, rSpCTL-D can bind to Gram-positive bacteria S. aureus, M. luteus; gram-negative bacteria E. tarda, V. fluvialis, V. parahaemolyticus, A. sobria, A. hydrophila, and V. alginolyticus; fungus S. cerevisiae. However, rSpCTL-C can bind to all the tested microorganisms except two kinds of Gram-positive bacteria S. aureus and M. luteus. Some previous studies have shown that C-type lectin can bind PAMPs and have the ability to bind bacteria. For example, the C-type lectin, PtCTL-9 in P. trituberculatus, displayed a dose-dependent binding ability for above PAMPs (Kang et al., 2021). While SpCTL-B could bind S. aureus, β-hemolytic Streptococcus, E. coli, V. alginolyticus, and A. hydrophila, but it had a weaker affinity for other tested microorganisms (V. parahemolyticus and S. cerevisiae) (Wei et al., 2018). Accordingly, different C-type lectins can bind to different bacteria, indicating that binding is selective, and different lectins have different functions.
It has been found that C-type lectins of marine organisms are mainly distributed in immune tissue organs, such as hepatopancreas, hemolymph and some tissues or organs directly contacted with the outside world. Previous studies have shown that ScCTL-1 was expressed in almost all tissues in Sinonovacula constricta, with the highest expression level in the hepatopancreas (Lan et al., 2020). In Marsupenaeus japonicus, CTLD was mainly expressed in the hepatopancreas (Wang et al., 2017). The results of Northern hybridization showed that C-type lectin (Fclectin) gene in Chinese shrimp Fenneropenaeus chinensis had obvious hybridization bands only in the blood cells (Liu et al., 2007). In this study, SpCTL-C and SpCTL-D were mainly expressed in the hepatopancreas and muscle, and the expression level in hepatopancreas was higher than that in the muscle, which proved SpCTL-C and SpCTL-D might be connected with immune defense in S. paramamosain.
Studies have shown that some C-type lectins in invertebrates can inhibit the growth of microorganisms and even kill them. In this study, rSpCTL-D had a certain inhibitory effect on V. fluvialis and M. luteus, while rSpCTL-C had no inhibitory effect on the growth of the tested microorganisms. M. japonicus MjHeCL maintains the expression of antimicrobial peptides to inhibit the proliferation of microflora in hemolymph (Wang et al., 2014). Argopecten irradians AiCTL-7 can inhibit the growth of E. coli at the concentration of 100 μg/mL. Therefore, different kinds of microorganisms can be inhibited by different C-type lectins, and further studies are required to understand the molecular mechanisms behind this process.
The most important feature of the C-type lectin family is that it can bind to a variety of cell surfaces by recognizing carbohydrate molecules on the cell surface and agglutinate cells with the participation of Ca2+ (Weis et al., 1998). rSpCTL-C and rSpCTL-D have specific agglutination effects on S. aureus, S. cerevisiae, and P. pastoris. Both of them could not agglutinate the tested Gram-negative bacteria. The studies on C-type lectins of other crustaceans showed that they all had different agglutination activities. In the presence of Ca2+, Rimicaris exoculate rReCTL-1 can agglutinate S. aureus and S. cerevisiae, and rReCTL-2 can agglutinate M. luteus, Vibrio vulnificus, and V. parahaemolyticus (Wang et al., 2020). S. paramamosain SpCTL-B has extensive microbial agglutination activity against Gram-positive /Gram-negative bacteria (Wei et al., 2018). The results show that C-type lectin, as a PRR, recognizes foreign microorganisms by carbohydrate domain and mediates cell-to-cell agglutination. CRD with different key motifs has specific agglutination activity.
Phagocytosis is an evolutionarily conserved immune mechanism for the clearance of pathogens and apoptosis. The whole process is completed by a single blood cell and can be roughly divided into four stages: chemotaxis, adhesion, endocytosis, and killing digestion (Coteur et al., 2002; Soares-Da-Silva et al., 2002). Previous studies have shown that C-type lectin can not only recognize and bind PAMPs and pathogenic microorganisms as PRR, but also promote the phagocytosis or encapsulation of blood cells to foreign invaders, which is the main way to remove the foreign microorganisms and plays an important role in innate immunity (Ling and Yu, 2006; Wang et al., 2017). In the present study, both recombinant proteins rSpCTL-C and rSpCTL-D can promote the phagocytosis of M. luteus by blood cells. At the same time, both rSpCTL-C and rSpCTL-D could promote the encapsulation activity of blood cells to agarose particles, and the encapsulation activity of rSpCTL-D was stronger. Similar results have been found in studies of other species; PtCTL-9 from swimming crab P. trituberculatus can significantly improve the encapsulation activity of blood cells to large invasive substances (Kang et al., 2021). MjGCTL from M. japonicus was observed to recruit hemocytes to encapsulate agarose particles (Alenton et al., 2017). In summary, SpCTL-C and SpCTL-D not only participates in the recognition and binding of the immune process, but also acts as an opsonin to promote the phagocytosis and encapsulation of foreign substances, suggesting they can act as collectin and selectin.
5 CONCLUSIONIn conclusion, this study shows that rSpCTL-C and rSpCTL-D can combine with PAMPs and a variety of pathogenic microorganisms, and remove some microorganisms by agglutination, and they can also promote blood cell phagocytosis and encapsulation to exert immune function. Therefore, rSpCTL-C and rSpCTL-D play an important role in the innate immunity of S. paramamosain.
6 DATA AVAILABILITY STATEMENTData that support the findings of this study are available from the corresponding author upon reasonable request.
Alenton R R R, Koiwai K, Miyaguchi K, Kondo H, Hirono I. 2017. Pathogen recognition of a novel C-type lectin from Marsupenaeus japonicus reveals the divergent sugar-binding specificity of QAP motif. Scientific Reports, 7: 45818.
DOI:10.1038/srep45818 |
Brubaker S W, Bonham K S, Zanoni I, Kagan J C. 2015. Innate immune pattern recognition: a cell biological perspective. Annual Review of Immunology, 33(1): 257-290.
DOI:10.1146/annurev-immunol-032414-112240 |
Cambi A, Koopman M, Figdor C G. 2005. How C-type lectins detect pathogens. Cellular Microbiology, 7(4): 481-488.
DOI:10.1111/j.1462-5822.2005.00506.x |
Cao J S, Lin Z Y, Zhang Y L, Zhang X Y, Li S K, Zhang N, Zou W H, Li Y Y, Chen J H, Wang X Y. 2013. Cloning and characterization of the SpLRR cDNA from green mud crab, Scylla paramamosain. Fish & Shellfish Immunology, 34(1): 129-135.
DOI:10.1016/j.fsi.2012.10.018 |
Christophides G K, Zdobnov E, Barillas-Mury C, Birney E, Blandin S, Blass C, Brey P T, Collins F H, Danielli A, Dimopoulos G, Hetru C, Hoa N T, Hoffmann J A, Kanzok S M, Letunic I, Levashina E A, Loukeris T G, Lycett G, Meister S, Michel K, Moita L F, Müller H M, Osta M A, Paskewitz S M, Reichhart J M, Rzhetsky A, Troxler L, Vernick K D, Vlachou D, Volz J, Mering C V, Xu J N, Zheng L B, Bork P, Kafatos F C. 2002. Immunity-related genes and gene families in Anopheles gambiae. Science, 298(5591): 159-165.
DOI:10.1126/science.1077136 |
Coteur G, Warnau M, Jangoux M, Dubois P. 2002. Reactive oxygen species (ROS) production by amoebocytes of Asterias rubens (Echinodermata). Fish & Shellfish Immunology, 12(3): 187-200.
DOI:10.1006/fsim.2001.0366 |
Drickamer K, Taylor M E. 1993. Biology of animal lectins. Annual Review of Cell Biology, 9(1): 237-264.
DOI:10.1146/annurev.cb.09.110193.001321 |
Drickamer K. 1993. Evolution of Ca2+-dependent animal lectins. Progress in Nucleic Acid Research and Molecular Biology, 45: 207-232.
DOI:10.1016/S0079-6603(08)60870-3 |
Drickamer K. 1999. C-type lectin-like domains. Current Opinion in Structural Biology, 9(5): 585-590.
DOI:10.1016/S0959-440X(99)00009-3 |
Huang M M, Mu C K, Wu Y H, Ye F, Wang D, Sun C, Lv Z B, Han B N, Wang C L, Xu X W. 2017. The functional characterization and comparison of two single CRD containing C-type lectins with novel and typical key motifs from Portunus trituberculatus. Fish & Shellfish Immunology, 70: 398-407.
DOI:10.1016/j.fsi.2017.09.029 |
Kang T, Xia Y T, Dong T W, Zheng X Y, Yang S, Qian S C, Huang M M, Fei H. 2021. C-type lectin with a QPN motif from swimming crab Portunus trituberculatus displays broad nonself-recognition ability and functions as an opsonin. Developmental and Comparative Immunology, 120: 104066.
DOI:10.1016/j.dci.2021.104066 |
Lan T Y, Li Z, Peng M X, Niu D H, Li Y, Li J L. 2020. A four-CRD C-type lectin from razor clam Sinonovacula constricta mediates agglutination and phagocytosis. Gene, 728: 144287.
DOI:10.1016/j.gene.2019.144287 |
Ling E J, Yu X Q. 2006. Cellular encapsulation and melanization are enhanced by immulectins, pattern recognition receptors from the tobacco hornworm Manduca sexta. Developmental & Comparative Immunology, 30(3): 289-299.
DOI:10.1016/j.dci.2005.05.005 |
Liu Y C, Li F H, Dong B, Wang B, Luan W, Zhang X J, Zhang L S, Xiang J H. 2007. Molecular cloning, characterization and expression analysis of a putative C-type lectin (Fclectin) gene in Chinese shrimp Fenneropenaeus chinensis. Molecular Immunology, 44(4): 598-607.
DOI:10.1016/j.molimm.2006.01.015 |
Loker E S, Adema C M, Zhang S M, Kepler T B. 2004. Invertebrate immune systems—not homogeneous, not simple, not well understood. Immunological Reviews, 198(1): 10-24.
DOI:10.1111/j.0105-2896.2004.0117.x |
Patel K D, Cuvelier S L, Wiehler S. 2002. Selectins: critical mediators of leukocyte recruitment. Seminars in Immunology, 14(2): 73-81.
DOI:10.1006/smim.2001.0344 |
Rowley A F, Powell A. 2007. Invertebrate immune systems specific, quasi-specific, or nonspecific?. The Journal of immunology, 179(11): 7209-7214.
DOI:10.4049/jimmunol.179.11.7209 |
Soares-Da-Silva I M, Ribeiro J, Valongo C, Pinto R, Vilanova M, Bleher R, Machado J. 2002. Cytometric, morphologic and enzymatic characterisation of haemocytes in Anodonta cygnea. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 132(3): 541-553.
DOI:10.1016/S1095-6433(02)00039-9 |
Taylor M E. 1993. Carbohydrate-recognition proteins of macrophages and related cells. In: Horton M A ed. Macrophages and Related Cells. Blood Cell Biochemistry, Vol. 5. Springer, Boston, MA. p. 347-370.
|
Tran N T, Kong T T, Zhang M, Li S K. 2020. Pattern recognition receptors and their roles on the innate immune system of mud crab (Scylla paramamosain). Developmental & Comparative Immunology, 102: 103469.
DOI:10.1016/j.dci.2019.103469 |
Wang G Y, Lei Y T, Kang T, Li Z, Fei H, Zeng B X, Zhou P, Wang C S, Lv Z B, Huang M M, Xu X W. 2020. Two C-type lectins (ReCTL-1, ReCTL-2) from Rimicaris exoculata display broad nonself recognition spectrum with novel carbohydrate binding specificity. Fish & Shellfish Immunology, 96: 152-160.
DOI:10.1016/j.fsi.2019.11.068 |
Wang X W, Gao J, Xu Y H, Xu J D, Fan Z X, Zhao X F, Wang J X. 2017. Novel pattern recognition receptor protects shrimp by preventing bacterial colonization and promoting phagocytosis. The Journal of Immunology, 198(8): 3045-3057.
DOI:10.4049/jimmunol.1602002 |
Wang X W, Xu J D, Zhao X F, Vasta G R, Wang J X. 2014. A shrimp C-type lectin inhibits proliferation of the hemolymph microbiota by maintaining the expression of antimicrobial peptides. Journal of Biological Chemistry, 289(17): 11779-11790.
DOI:10.1074/jbc.M114.552307 |
Wang X W, Zhang X W, Xu W T, Zhao X F, Wang J X. 2009. A novel C-type lectin (FcLec4) facilitates the clearance of Vibrio anguillarum in vivo in Chinese white shrimp. Developmental & Comparative Immunology, 33(9): 1039-1047.
DOI:10.1016/j.dci.2009.05.004 |
Wei X Y, Wang L M, Sun W W, Zhang M, Ma H Y, Zhang Y L, Zhang X X, Li S K. 2018. C-type lectin B (SpCTL-B) regulates the expression of antimicrobial peptides and promotes phagocytosis in mud crab Scylla paramamosain. Developmental and Comparative Immunology, 84: 213-229.
DOI:10.1016/j.dci.2018.02.016 |
Weis W I, Taylor M E, Drickamer K. 1998. The C-type lectin superfamily in the immune system. Immunological Reviews, 163(1): 19-34.
DOI:10.1111/j.1600-065X.1998.tb01185.x |
Yu Y H, Yu Y C, Huang H Q, Feng K X, Pan M M, Yuan S C, Huang S F, Wu T, Guo L, Dong M L, Chen S W, Xu A L. 2007. A short-form C-type lectin from amphioxus acts as a direct microbial killing protein via interaction with peptidoglycan and glucan. The Journal of Immunology, 179(12): 8425-8434.
DOI:10.4049/jimmunol.179.12.8425 |
Zelensky A N, Gready J E. 2003. Comparative analysis of structural properties of the C-type-lectin-like domain (CTLD). Proteins: Structure, Function, and Bioinformatics, 52(3): 466-477.
DOI:10.1002/prot.10626 |
Zelensky A N, Gready J E. 2005. The C-type lectin-like domain superfamily. The FEBS Journal, 272(24): 6179-6217.
DOI:10.1111/j.1742-4658.2005.05031.x |
Zhang W J, Zhang Z Y, Mu C K, Li R H, Ye Y F, Zhang H, Song W W, Shi C, Liu L, Wang C L. 2019. Molecular cloning, characterization, and expression of a C-type lectin from Scylla paramamosain, which might be involved in the innate immune response. Fish & Shellfish Immunology, 93: 251-257.
DOI:10.1016/j.fsi.2019.07.035 |
Zhu Y T, Zhang X, Wang S C, Li W W, Wang Q. 2016. Antimicrobial functions of EsLecH, a C-type lectin, via JNK pathway in the Chinese mitten crab, Eriocheir sinensis. Developmental & Comparative Immunology, 61: 225-235.
DOI:10.1016/j.dci.2016.04.007 |
 2022, Vol. 40
2022, Vol. 40