Institute of Oceanology, Chinese Academy of Sciences
Article Information
- SUN Shao'e, SHA Zhongli, WANG Yanrong
- Mitochondrial phylogenomics reveal the origin and adaptive evolution of the deep-sea caridean shrimps (Decapoda: Caridea)
- Journal of Oceanology and Limnology, 39(5): 1948-1960
- http://dx.doi.org/10.1007/s00343-020-0266-4
Article History
- Received Jul. 15, 2020
- accepted in principle Oct. 26, 2020
- accepted for publication Dec. 17, 2020
2 Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology (Qingdao), Qingdao 266237, China;
3 Center for Ocean Mega-Science, Chinese Academy of Sciences, Qingdao 266071, China;
4 University of Chinese Academy of Sciences, Beijing 100049, China
Deep-sea ecosystems usually refer to the waters and sediments of the ocean below 200 m, representing the largest and most remote biome of the world (Herring, 2002). With the development and improvement of technology, the deep-sea exploration has discovered diverse habitats and ecosystems, including seamounts, ridges, deep-water coral reefs, hydrothermal vents, cold seeps, trenches, and so on (Corinaldesi, 2015). Compared with shallow water species, deep-sea species survive in harsh environmental conditions, such as low temperatures, low oxygen level, scarce food, a lack of sunlight, and high pressure (Sanders and Hessler, 1969). Despite recent major advances in biological research, only 5% of the deep-sea has been explored in detail so far, and less than 0.001% deep-sea organisms has been sampled and described in detail (Danovaro et al., 2014). It is meaningful and important to elucidate life origin by exploring the origin and adaptive genetic mechanisms of the large deep-sea organisms.
As complete organelle genome, mitochondrial genome (mitogenome) shows several advantages, such as relatively high nucleotide substitution rates, lack of extensive recombination, and conserved gene content. The mitogenomes have been proved to be a useful tool for phylogenetic and phylogeographic analyses of animal taxa (Moritz and Brown, 1987; Boore, 1999; Curole and Kocher, 1999). The mitochondrial genomes provide more phylogenetic signals, making more accurate results than analyses of one or few mitochondrial DNA (mtDNA) markers. Mitogenomic analyses are useful for phylogenetic reconstruction in different invertebrate groups, such as molluscs (e.g., Mikkelsen et al., 2018; Kong et al., 2020), insects (e.g., Yuan et al., 2015), and annelidans (e.g., Li et al., 2015). Many phylogenetic studies using the complete mitochondrial genome are available for crustaceans (Shi et al., 2012; Shen et al., 2013; Ji et al., 2014). In addition, mitochondria provide 95% cell energy through the oxidative phosphorylation (OXPHOS), playing a crucial role in aerobic respiration and energy metabolism (Das, 2006). The extreme environments of deep-sea may have affected the evolution of mitogenome, as well as the energy production of vent faunas, because several components of the electron transport chain are encoded by mitochondrial genes (Ki et al., 2009).
Caridean shrimps (Decapoda: Caridea) are the second most diverse group amongst the decapods, which are widespread from tropical to polar regions of the world, in both marine and freshwater habitats (Li et al., 2011). Furthermore, some caridean shrimps adapt to deep-sea environments. Species from the families Thoridae (Komai et al., 2019), Pandalidae (Yang et al., 2017), Oplophoridae (Cardoso and Young, 2005), Acanthephyridae (Li, 2015) inhabit in deep-sea seamounts. Species from the family Bathypalaemonellidae have been collected in a deep-sea coral reef habitat (Cardoso, 2010). The alvinocarid shrimps (Bresilioidea: Alvinocarididae) are vent-endemic species, which comprise the predominant faunal biomass of various hydrothermal ecosystems (Hernández-Ávila longirostris is a species of alvinocarid shrimp co-distributed in deep-sea hydrothermal vent and cold seep environments (Hui et al., 2018). Shrimps of the genus Lebbeus from the family Thoridae have also been reported from various deep-sea hydrothermal vents and cold seep sites (Komai et al., 2012; Chan and Komai, 2017). Caridean shrimps are good models to test the hypotheses regarding the origin and adaptive evolution of the deep-sea organisms. However, limited studies have focused on these matters because of the difficulty of sampling deep-sea species. Scientists have only focused their attention on habitats of deep-sea hydrothermal vents (Wang et al., 2017; Sun et al., 2018a, b, 2019a) and cold seeps (Hui et al., 2018; Xin et al., 2020), where life conditions are even more extreme.
In this study, we analyzed three complete mitogenomes of deep-sea hydrothermal vent alvinocarid shrimps and eight mitogenomes of deep-sea seamount caridean shrimps. Our specific aims were to clarify (1) where did the deep-sea (hydrothermal vent and seamount) caridean shrimps originate; (2) when did the caridean shrimps colonized deep-sea habitat; (3) how did the caridean shrimps adapted to the deep-sea environment in the perspective of mitochondrial genome.
2 MATERIAL AND METHOD 2.1 Sampling and DNA extractionIn this study, the deep-sea caridean shrimps were collected from both seamount and hydrothermal vent habitats. The hydrothermal vent alvinocaridid shrimps were captured from Manus Basin, Western Pacific. The seamount specimens of the other caridean families were captured from Yap Seamount, Western Pacific. Both the hydrothermal vent and seamount specimens were collected using the remotely operated vehicle (ROV) and then preserved in 95% ethanol until DNA extraction. Total genomic DNA of each species was isolated using the DNeasy tissue kit (Qiagen) according to the manufacturer's instructions.
2.2 Illumina sequencing, mitogenome assembly, and annotationThe mitogenomes were sequenced using Illumina-based whole genome shotgun sequencing. Sequencing libraries were generated using NEBNext® UltraTM DNA Library Prep Kit for Illumina (NEB, USA) following manufacturer's recommendations, which was sequenced on an Illumina HiSeq 2500 platform. The reads with average quality scores less than 20 were trimmed from further analysis. Clean data were then assembled using CLC Genomics Workbench v. 11.0.64 (http://www.clcbio.com/products/clcgenomics-workbench/) and SOAP denovo (Li et al., 2010) (k-mer=55). De novo assembled contigs were blasted against the NCBI nr database using "BLAST" tool implemented in the CLC Genomics Workbench to identify contigs of mitochondrial origin (E-value=1.0E-15). The putative "mitochondrial DNA" contigs were aligned with the available complete mitogenomes of the Caridea. In order to obtain a circular mtDNA, the contigs identified as mitogenome sequences were examined for repeats at the beginning and end of the sequence using SeqMan of the DNASTAR software package (http://www.dnastar.com).
ORF Finder (http://www.ncbi.nlm.nih.gov/gorf/gorf.html) and BLASTx were used to annotate the protein-coding genes (PCGs) using the invertebrate mitochondrial genetic code. The software ARWEN (Laslett and Canbäck, 2008), DOGMA (Wyman et al., 2004), and MITOS Web Server (Bernt et al., 2013) located the tRNA genes using the invertebrate mitochondrial genetic code. The rRNA genes were inferred by BLAST searches (http://www.ncbi.nlm.nih.gov/BLAST/) against the available caridean sequences.
2.3 Phylogenetic analysisPhylogeny of the Caridea was inferred based on 44 available complete mitogenomes. Available caridean mitogenomes were expanded with 11 newly determined complete sequence from the families Alvinocarididae, Thoridae, Pandalidae, Oplophoridae, Nematocarcinidae, Acanthephyridae, and Bathypalaemonellidae. Five penaeoid species were selected as outgroup taxa (Supplementary Table S1).
The phylogenetic relationships were reconstructed based on the concatenated nucleotide sequences of 13 PCGs. The nucleotide sequences for each PCG were aligned with MAFFT (Katoh et al., 2005) using default settings. Ambiguously aligned and variable areas were recognized using the program Gblocks (Talavera and Castresana, 2007) (default setting) and excluded from the analyses. A single alignment for phylogenetic analyses was conducted by concatenating all nucleotide sequences of the 13 PCGs. The best-fit nucleotide substitution models were selected by jModelTest (Posada, 2008).
Phylogenetic trees were built by both Maximum Likelihood (ML; Felsenstein, 1981) and Bayesian inference (BI; Huelsenbeck and Ronquist, 2001) methods. ML trees were constructed using RAxML Black-Box webserver (http://phylobench.vital-it.ch/raxml-bb/index.php; Stamatakis et al., 2008). The reliability of the tree topology was evaluated using bootstrap support with 1 000 replicates. RAxMLoptimized parameters of the GTR model for nucleotide sequences. BI was conducted with PhyloBayes MPI (Lartillot et al., 2013) with the CAT site-heterogeneous mixture model (Lartillot and Philippe, 2004, 2006; Lartillot et al., 2007). Two independent Markov chain Monte Carlo (MCMC) runs chains were run. Tracer v 1.6 (Rambaut et al., 2014) was used to assess stationarity and appropriate burn-in (1 000). Chains were considered to converge well when the maxdiff value was less than 0.1, rel_ diff value was less than 0.3, and the minimum effective sample size was greater than 50, respectively, which were measured by bpcomp and tracecomp (Lartillot et al., 2013).
2.4 Divergence time estimationBEAST 1.8.1 (Drummond et al., 2012) was applied to estimate divergence times of deep-sea clades of Caridea with the relaxed uncorrelated lognormal clocks, random starting trees, and the Yule speciation model. Two independent MCMC runs were executed for 10 million steps, sampling every 1 000 steps. Convergence of the chains was checked using the program Tracer v 1.6 (Rambaut et al., 2014) to ensure that effective sample sizes (ESSs) were above 200, and discard the initial 50% as burn-in. The remaining samples were combined and the maximum-clade-credibility tree was obtained using TreeAnnotator in BEAST.
2.5 Analysis of selective pressureCodeML implemented in the PAML package (Yang, 2007) was used to explore the patterns of natural selection and identify the positive selection sites in each of the thirteen PCGs. The overall mean ratio of nonsynonymous (Ka) to synonymous (Ks) substitution rates (ω) for 13 PCGs was calculated with the "one-ratio (M0)" model. The deep-sea caridean shrimps were labeled the foreground branch, and all other caridean shrimps were labeled the background branch. The "two-ratio (M2)" model was used to compute ω ratio for the foreground (ω1) and background (ω0) branches. Likelihood ratio tests (LRTs) were performed to test whether the M2 fits the data significantly better than the M0. The branch-site model (Zhang et al., 2005) was used to test for positive selection on the foreground branch. The Model A (MA) assumes either purifying selection (0≤ω0≤1) or neutrality (ω1=1) on all branches, and positive selection (ω2 > 1) along the foreground branch. The null model (MA0) is constrained that ω2=1. When the likelihood ratio tests were significant, we used the Bayes Empirical Bayes (BEB) method (Yang et al., 2005) to identify sites under positive selection by calculating the posterior probabilities.
3 RESULT 3.1 General features of the new mitochondrial genomesFrom the eleven new mitochondrial genomes (mitogenomes), 10 were complete and 1 was partial (Supplementary Table S2). The complete mitogenomes ranged in size from 15 618 to 17 168 bp, with length variations mainly in the A+T-rich regions. Most of these mitogenomes harboured a typical set of 37 genes and an identical gene arrangement as previously reported for other caridean mitogenomes. While the tRNA-Ala, tRNA-Val, and tRNA-Tyr genes were absent from Bathypalaemonella sp. mitogenome, and tRNA-Ile gene was absent from Paralebbeus jiaolongi mitogenome. In order to verify if the tRNA absence in these species is a methodological fail, we diagnosed the targeted tRNA sequences by means of the polymerase chain reaction (PCR). However, the tRNA-Ala, tRNA-Val, and tRNA-Tyr genes were still not found in Bathypalaemonella sp. mitogenome, and tRNA-Ile gene was still absent from P. jiaolongi mitogenome. The partial sequence of Lebbeus shinkaiae mitogenome lacks the region between nad5 and nad4. According to the annotation of the sequenced caridean mitogenomes, we inferred that partial sequences of the nad5 and nad4 genes, and the tRNA gene tRNA-His locate in the unamplified portion of L. shinkaiae mitogenome.
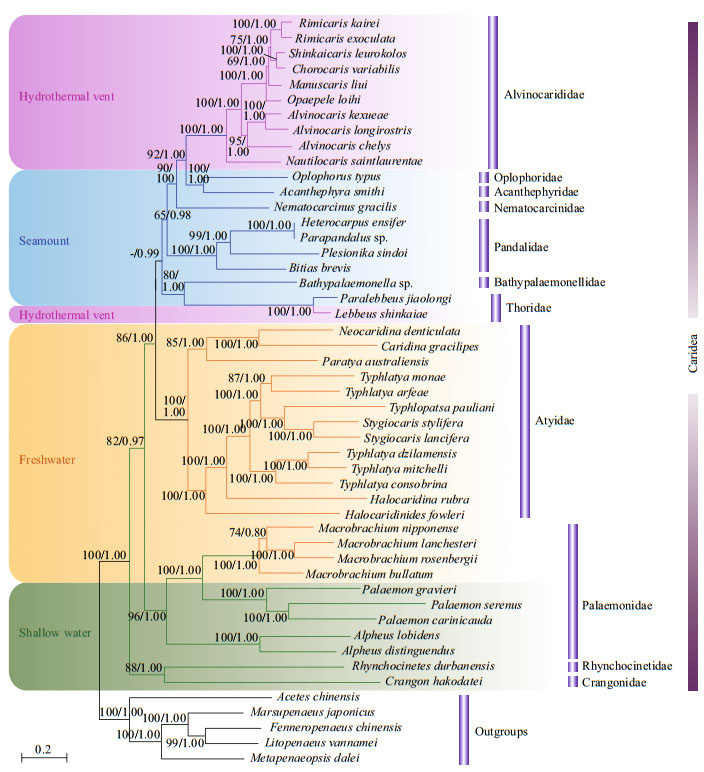
3.2 Phylogenetic reconstruction and divergence time estimationMaximum Likelihood (ML) and Bayesian inference (BI) inferred similar topological structures with varying levels of support. Therefore, we showed the nodal supports obtained from the two methods together on the ML tree (Fig. 1). At the first level, the families Rhynchocinetidae and Crangonidae from shallow water were separated from all other taxa and placed as a basal clade of the tree. The other caridean families included in the analysis separated into three clades. The first clade was solely comprised of Palaemonidae species from both shallow water and freshwater, and the second clade contained only Atyidae species from freshwater. The third major clade included all the remaining families (Thoridae, Bathypalaemonellidae, Pandalidae, Nematocarcinidae, Acanthephyridae, Oplophoridae, and Alvinocarididae) from seamount and hydrothermal vent environments. The hydrothermal vent alvinocaridid shrimps were situated at a more evolved position within the infraorder Caridea. They have the closest relationship with the seamount caridean shrimps (Fig. 1).
|
| Fig.1 Phylogenetic trees derived from ML and BI analysis of caridean species based on nucleotide sequences of 13 PCGs The first number at each node is the bootstrap probability of ML analysis and the second number is Bayesian posterior probability. The pink, blue, yellow, and green branches indicate shrimps from hydrothermal vent, seamount, freshwater, and shallow water, respectively. |
The divergence times and the 95% highest posterior density (HPD) intervals are presented for each node (Fig. 2). The time to the most recent common ancestor (MRCA) of seamount and hydrothermal vent caridean shrimps was estimated at 147.75 Ma (95% HPD: 111.44–185.73). The MRCA of the hydrothermal vent alvinocaridid shrimps were estimated at 61.39 Ma (95% HPD: 24.31–102.01) at the Cretaceous-Tertiary boundary. In the family Thoridae, divergence between hydrothermal vent shrimp L. shinkaiae and its seamount relative P. jiaolongi was estimated to have occurred 20.09 Ma (95% HPD: 4.22–40.50). The divergence of most of the modern deep-sea species are in the late Cretaceous/early Tertiary.

|
| Fig.2 Caridea divergence time estimated using the Bayesian relaxed-molecular clock method The 95% confidence intervals for each node are shown in light purple bars. Calibrated nodes are indicated by arrows. |
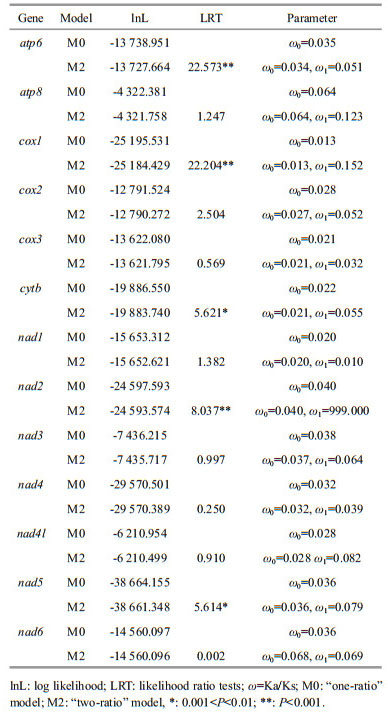
The one-ratio (M0) model analyses of 13 PCGs showed that the ω values for each gene were significantly less than 1 (Table 1), implying that these genes have experienced constrained selective pressure to maintain their function. The result showed that cox1 gene (0.013) experienced the strongest purifying Pressure, while atp8 gene (0.064) experienced the weakest purifying pressure. Based on the two-ratio (M2) model, we found that all the PCGs of deep-sea caridean shrimps showed a higher ω value than other caridean shrimps, except for nad1, which had a slightly reduced ω value in alvinocaridid shrimps branch. The LRT tests showed that the M2 model fits the data significantly better than the M0 model at five genes (atp6, cox1, cytb, nad2, and nad5), suggesting a divergence in selective pressure between deep-sea and other caridean shrimps.
|
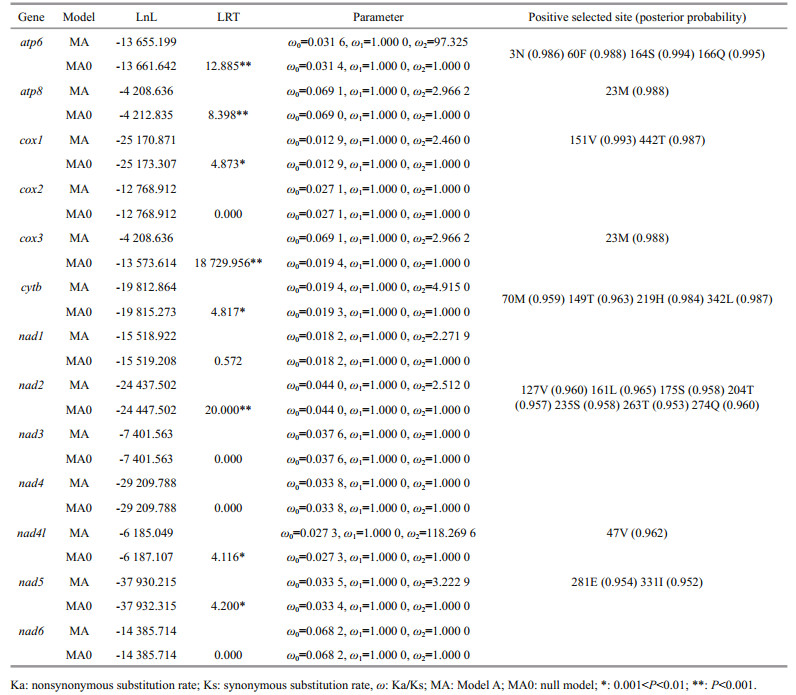
The branch-site model was performed to detect positive selection in individual codons. Eight genes (i.e., atp6, atp8, cox1, cox3, cytb, nad2, nad4l, and nad5) were found to be under positive selection in the deep-sea shrimp lineages, where LRTs of the branch-site model (MA vs. MA0) were statistically significant (Table 2). In these eight genes, total 22 positive selection sites showed BEB values > 0.95 using branch-site models.
|
The evolutionary origin of the deep-sea fauna has been debated for a long time. The study of different taxa has revealed a variety of different colonization patterns based on the zoogeographic data. The analysis of the distribution and taxonomy of deep-sea molluscan, echinoderms, and asellote isopods suggested that all the faunas are probably shaped by invasions from adjacent shallow-water regions (Allen, 1979; Jablonski and Bottjer, 1990; Raupach et al., 2009; Woolley et al., 2016). In the cases of stylasterid corals, it has been suggested that abyssal forms are most primitive, while those in the shallow waters are more recently evolved (Lindner et al., 2008). The microfossil analyses of Foraminifera from both deep-sea and shallow Antarctic indicate that migration of foraminiferal species may has occurred in both directions (Lipps and Hickman, 1982; Hayward, 2001).
In this context, our phylogenetic analyses showed that the deep sea might be initially colonized with caridean shrimps by the migration events from the shallow water. Transitions from shallow (onshore) to deep (offshore) environments has already been documented in many other decapod crustaceans (e.g. Chan et al., 2009; Palero et al., 2009; Tsang et al., 2009; Tsoi et al., 2011; Yang et al., 2012, 2015). These studies suggested that the invasion of deep-sea habitats might drive the early diversifications in diverse groups of decapods (Yang et al., 2012).
Furthermore, our investigation revealed that the Alvinocarididae and Thoridae species may underwent a second range expansion from the seamount to the hydrothermal vent habitats. These observations suggested that the vent-endemic species perhaps migrated from the surrounding deep-sea environments, i.e. seamount, instead of the remnants of ancient hydrothermal species, which agree with the extinction/repopulation hypothesis of vent taxa (Jacobs and Lindberg, 1998). The extinction/repopulation hypothesis posited that the global deep-water anoxic/dysoxic events during the Late Cretaceous and Early Tertiary resulted in the extinction of nearly all contemporary vent species, and the hydrothermal vent species later migrated from non-hydrothermal environments (Jacobs and Lindberg, 1998). The hydrothermal vents are the typical extreme deep-sea environment, which present some of the most physically and chemically challenging habitats to evolutionary invasion (Van Dover, 2000). The hydrothermal vent environment is full with plentiful chemoautotrophic bacteria, producing a rich food source. Simultaneously, this environment has the least predation pressure as predators must also have specialized adaptation to these extreme habitats, and very few appear to have evolved these adaptations (Fisher et al., 2007). Thus, the hydrothermal vents also act as environmental filters that promote the evolution and distribution of species with specialized adaptation and limits of tolerance to oxygen, temperature, and sulfide (Tunnicliffe et al., 2003; Takai et al., 2006; Fisher et al., 2007). The widespread persistence of hydrothermal vent environments in earth's geologic history (Shock et al., 1995) may have provided an important element enabling the independent colonization by alvinocaridid shrimps.
4.2 When did caridean shrimps invade deep sea?Although the time to the MRCA of deep-sea caridean shrimps was in the late Jurassic, the MRCA of hydrothermal vent alvinocarid shrimps and the divergence time of the most modern seamount species were in the late Cretaceous/early Tertiary. These times are comparable to the estimates of origin and radiation in other deep-sea taxa, e.g. radiation of hydrothermal vent barnacles at ca. 68 Ma (Herrera et al., 2015); origin of siboglinid tubeworms at ca. 60 Ma (Chevaldonné et al., 2002); origin of bythograeid crabs at 48.4–55.9 Ma (Yang et al., 2013); radiation of mytilid mussels at ca. 45 Ma (Lorion et al., 2013). The period of the Late Cretaceous/Early Tertiary was marked by the continuous separation of continents (Parker and Gealey, 1985). During this period, the back-arc systems of Southeast Asia and northeast Australia were opened, forming the main back-arc basins of the Western Pacific. This tectonic and oceanographic transition provided habitats for the caridean species colonization in the deep sea. Moreover, the global deep-water anoxic/dysoxic events occurred in the late Cretaceous/early Tertiary were thought to lead to a massive extinction in deep-sea benthic organisms (Jacobs and Lindberg, 1998), providing opportunities for invasion of the modern taxa (Jacobs and Lindberg, 1998). In addition, this period coincided with a series of other remarkable events including global warming, profound changes in carbon cycling, as well as the greenhouse gas concentrations (Norris et al., 2001). These events may have been attributed to increasing the disturbance in shallow marine environments, such as the evolution of new predators (Vermeij, 1987) and more effective competitors (Vermeij, 1995), promoting the offshore retreat. The global warming was followed by a declined deep-water temperature beginning in the Middle (50–48 Ma) to Late (40–36 Ma) Eocene, associated with the initial glaciation of Antarctica (Zachos et al., 2001). The low water temperatures may decreased the metabolic rate of carideans larvae, increasing their longevity and enhancing their dispersal capability (Lorion et al., 2013). This may be an advantage for carideans diversifying in deep-sea habitats.
4.3 Positive selection of mitochondrial genes associated with deep-sea adaptation in caridean shrimpsOnce shrimps invaded into deep-sea habitats, they began to face drastic environmental fluctuations, including low water temperature, high pressure, dark, low oxygen concentration, and so on (Katayama et al., 2012). Energy metabolism is an important aspect in adapt to different environments, which can be altered due to environmental conditions to best match energetic demands (da Silva-Castiglioni et al., 2010, 2011; Guo et al., 2018). Mitochondrion is the main site of energy production, providing about 95% of the adenosine triphosphate (ATP) needed for the basic activities of life through the OXPHOS (Das, 2006). The mitogenome can be characterized by its adaptations to the extreme living environments (Castellana et al., 2011). Previous researchers have found that one major adaptation of decapod crustaceans to harsh conditions is positive selection on mitochondrial genes involved in hypoxia response and energy metabolis, such as cave shrimps (Guo et al., 2018), hydrothermal vent squat lobsters (Sun et al., 2019a, b), hydrothermal vent alvinocaridid shrimps and crabs (Sun et al., 2019a).
In the present study, twenty-two residues, which were located in eight mitochondrial OXPHOS genes along the lineage of deep-sea caridean shrimps, were identified as positively selected sites. These mutations may have functional implications, as they are in components of the electron transport chain. At complex IV (cytochrome c oxidase), about 95% of the molecular oxygen (O2) is consumed to form water (Ferguson-Miller et al., 2012; Koopman et al., 2013). Therefore, hypoxia is a major inhibitor for cytochrome c oxidase (Cooper and Brown, 2008). Previous study reported that the mutation of the structure and/or activity of cytochrome c oxidase of the respiratory chain might help to hypoxia adaptation (Luo et al., 2008). Therefore, it is reasonable to assume that the positive selections detected in cox1 and cox3 perhaps enhanced the ability of caridean shrimps to resist the hypoxia environment in deep sea. The nicotinamide adenine dinucleotide (NADH) dehydrogenase complex acts as a proton pump and mutations in this complex may influence the efficiency of proton-pumping, and then affect the metabolic efficiency (da Fonseca et al., 2008). ATP synthase is the last enzyme complex in the respiratory, which participates in the oxidative phosphorylation after electrons have been transported and a proton gradient has been formed (Mishmar et al., 2003; Zhou et al., 2014; Zhang et al., 2017). When environmental oxygen is reduced, the ATPase activity will increase (Martinez-Cruz et al., 2011). According, mutations in atp6 and atp8 genes could influence the ATPase activity. In this study, most positive selected residues were identified in the NADH dehydrogenase complex, followed by ATP synthase. Thus, it is reasonable to assume that positive selections in the NADH dehydrogenase complex and ATP synthase perhaps enhanced the ability of deep-sea caridean shrimps to resist the hypoxic deep-sea environment. These results provided a necessary molecular mechanism underlying the adaptation of the carideans to deep-sea environments at mitochondrial level, and revealed a modified and adapted energy metabolism in deep-sea species under extreme environments.
5 CONCLUSIONIn this study, we newly determined 11 mitogenomes of deep-sea caridaen shrimps. The phylogenetic relationship, divergence times, and adaptive evolution of deep-sea caridean shrimps were explored. The origin analysis indicated that deep sea was initially colonized with caridean shrimps from shallow water at 147.75 Ma, and the hydrothermal vent shrimps underwent a second range expansion from deep sea to vent ecosystems at 61.39 Ma (shrimps of Alvinocarididae) and 20.09 Ma (shrimps of Thoridae), which support the extinction/repopulation hypothesis of vent taxa. The divergence of most of the modern deep-sea species are in the late Cretaceous/early Tertiary. Total twenty-two positively selected sites were identified in eight mitochondrial genes of deep-sea species, indicating their deep-sea adaption. This investigation strengthen our understanding of origin and adaptive evolution in the mitogenome of deep-sea shrimps.
6 DATA AVAILABILITY STATEMENTThe authors declare that all data supporting the findings of this study are available within the appendix sections.
7 ACKNOWLEDGMENTWe thank the crews of R/V Kexue for their help in sample collection.
Electronic supplementary materialSupplementary material (Supplementary Tables S1–S2) is available in the online version of this article at https://doi.org/10.1007/s00343-020-0266-4.
Allen J A. 1979. The adaptations and radiation of deep-sea bivalves. Sarsia, 64(1-2): 19-27.
DOI:10.1080/00364827.1979.10411357 |
Bernt M, Donath A, Jühling F, Externbrink F, Florentz C, Fritzsch G, Pütz J, Middendorf M, Stadler F P. 2013. MITOS: improved de novo metazoan mitochondrial genome annotation. Molecular Phylogenetics and Evolution, 69(2): 313-319.
DOI:10.1016/j.ympev.2012.08.023 |
Boore J L. 1999. Animal mitochondrial genomes. Nucleic Acids Research, 27(8): 1 767-1 780.
DOI:10.1093/nar/27.8.1767 |
Cardoso I A. 2010. First record of family Bathypalaemonellidae (Caridea: Decapoda) on Brazilian deep-sea coral reefs. Marine Biodiversity Records, 3: e108.
DOI:10.1017/S1755267210000941 |
Cardoso I, Young P. 2005. Deep-sea oplophoridae (Crustacea Caridea) from the southwestern Brazil. Zootaxa, 1031(1): 1-76.
DOI:10.11646/zootaxa.1031.1.1 |
Castellana S, Vicario S, Saccone C. 2011. Evolutionary patterns of the mitochondrial genome in metazoa: exploring the role of mutation and selection in mitochondrial protein‐coding genes. Genome Biology and Evolution, 3: 1 067-1 079.
DOI:10.1093/gbe/evr040 |
Chan T Y, Ho K C, Li C P, Chu K H. 2009. Origin and diversification of the clawed lobster genus Metanephrops (Crustacea: Decapoda: Nephropidae). Molecular Phylogenetics and Evolution, 50(3): 411-422.
DOI:10.1016/j.ympev.2008.11.020 |
Chan T Y, Komai T. 2017. A new shrimp species of the genus Lebbeus White, 1847 (Crustacea: Deacpoda: Caridea: Thoridae) from a deep-sea cold seep site off Southwestern Taiwan. Zootaxa, 4238(3): 426-432.
DOI:10.11646/zootaxa.4238.3.9 |
Chevaldonné P, Jollivet D, Desbruyeres D, Lutz R A, Vrijenhoek R C. 2002. Sister-species of eastern Pacific hydrothermal vent worms (Ampharetidae, Alvinellidae, Vestimentifera) provide new mitochondrial COI clock calibration. Cahiers de Biologie Marine, 43: 367-370.
|
Cooper C E, Brown G C. 2008. The inhibition of mitochondrial cytochrome oxidase by the gases carbon monoxide, nitric oxide, hydrogen cyanide and hydrogen sulfide: chemical mechanism and physiological significance. Journal of Bioenergetics and Biomembranes, 40(5): 533-539.
DOI:10.1007/s10863-008-9166-6 |
Corinaldesi C. 2015. New perspectives in benthic deep-sea microbial ecology. Frontiers in Marine Science, 2: 17.
DOI:10.3389/fmars.2015.00017 |
Curole J P, Kocher T D. 1999. Mitogenomics: digging deeper with complete mitochondrial genomes. Trends in Ecology & Evolution, 14(10): 394-398.
DOI:10.1016/S0169-5347(99)01660-2 |
da Silva-Castiglioni D, Oliveira G T, Buckup L. 2010. Metabolic responses of Parastacus defossus and Parastacus brasiliensis (Crustacea, Decapoda, Parastacidae) to hypoxia. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 156(4): 436-444.
DOI:10.1016/j.cbpa.2010.03.025 |
da Silva-Castiglioni D, Oliveira G T, Buckup L. 2011. Metabolic responses in two species of crayfish (Parastacus defossus and Parastacus brasiliensis) to post-hypoxia recovery. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 159(3): 332-338.
DOI:10.1016/j.cbpa.2011.03.030 |
Danovaro R, Snelgrove P V R, Tyler P. 2014. Challenging the paradigms of deep-sea ecology. Trends in Ecology & Evolution, 29(8): 465-475.
DOI:10.1016/j.tree.2014.06.002 |
Das J. 2006. The role of mitochondrial respiration in physiological and evolutionary adaptation. BioEssays, 28(9): 890-901.
DOI:10.1002/bies.20463 |
Drummond A J, Suchard M A, Xie D, Rambaut A. 2012. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Molecular Biology and Evolution, 29(8): 1 969-1 973.
DOI:10.1093/molbev/mss075 |
Felsenstein J. 1981. Evolutionary trees from DNA sequences: a maximum likelihood approach. Journal of Molecular Evolution, 17(6): 368-376.
DOI:10.1007/BF01734359 |
Ferguson-Miller S, Hiser C, Liu J. 2012. Gating and regulation of the cytochrome c oxidase proton pump. Biochimica et Biophysica Acta (BBA)-Bioenergetics, 1817(4): 489-494.
DOI:10.1016/j.bbabio.2011.11.018 |
Fisher C R, Takai K, Le Bris N. 2007. Hydrothermal vent ecosystems. Oceanography, 20(1): 14-23.
DOI:10.5670/oceanog.2007.75 |
Fonseca R R D, Johnson W E, O'Brien S J, Ramos M J, Antunes A. 2008. The adaptive evolution of the mammalian mitochondrial genome. BMC Genomics, 9: 119.
DOI:10.1186/1471-2164-9-119 |
Guo H Y, Yang H, Tao Y T, Tang D, Wu Q, Wang Z F, Tang B P. 2018. Mitochondrial OXPHOS genes provides insights into genetics basis of hypoxia adaptation in anchialine cave shrimps. Genes & Genomics, 40(11): 1 169-1 180.
DOI:10.1007/s13258-018-0674-4 |
Hayward B W. 2001. Global deep-sea extinctions during the Pleistocene ice ages. Geology, 29(7): 599-602.
DOI:10.1130/0091-7613(2001)029<0599:GDSEDT>2.0.CO;2 |
Hernández-ávila I, Cambon-Bonavita M A, Pradillon F. 2015. Morphology of first zoeal stage of four genera of alvinocaridid shrimps from hydrothermal vents and cold seeps: implications for ecology, larval biology and phylogeny. PLoS One, 10(12): e0144657.
DOI:10.1371/journal.pone.0144657 |
Herrera S, Watanabe H, Shank T M. 2015. Evolutionary and biogeographical patterns of barnacles from deep-sea hydrothermal vents. Molecular Ecology, 24(3): 673-689.
DOI:10.1111/mec.13054 |
Herring P J. 2002. The Biology of the Deep Ocean. Oxford University Press, Oxford, UK.
|
Huelsenbeck J P, Ronquist F. 2001. Mrbayes: Bayesian inference of phylogenetic trees. Bioinformatics, 17(8): 754-755.
DOI:10.1093/bioinformatics/17.8.754 |
Hui M, Cheng J, Sha Z L. 2018. Adaptation to the deep-sea hydrothermal vents and cold seeps: insights from the transcriptomes of Alvinocaris longirostris in both environments. Deep Sea Research Part I: Oceanographic Research Papers, 135: 23-33.
DOI:10.1016/j.dsr.2018.03.014 |
Jablonski D, Bottjer D J. 1990. Onshore-offshore trends in marine invertebrate evolution. In: Ross R M, Allmon W D eds. Causes of Evolution. University of Chicago Press, Chicago. p. 21-75.
|
Jacobs D K, Lindberg D R. 1998. Oxygen and evolutionary patterns in the sea: onshore/offshore trends and recent recruitment of deep-sea faunas. Proceedings of the National Academy of Sciences of the United States of America, 95(16): 9 396-9 401.
DOI:10.1073/pnas.95.16.9396 |
Ji Y K, Wang A, Lu X L, Song D H, Jin Y H, Lu J J, Sun H Y. 2014. Mitochondrial genomes of two brachyuran crabs (Crustacea: Decapoda) and phylogenetic analysis. Journal of Crustacean Biology, 34(4): 494-503.
DOI:10.1163/1937240X-00002252 |
Katayama K, Ookura M, Yamasaki H, Shigeshima K, Fujimoto T, Fujiwara T. 2012. Effect of normal air pressure low oxygen concentration environments on resting metabolism. The Journal of Japan Academy of Health Sciences, 14(4): 199-204.
DOI:10.24531/jhsaiih.14.4_199 |
Katoh K, Kuma K I, Toh H, Miyata T. 2005. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Research, 33(2): 511-518.
DOI:10.1093/nar/gki198 |
Ki J S, Dahms H U, Hwang J S, Lee J S. 2009. The complete mitogenome of the hydrothermal vent crab Xenograpsus testudinatus (Decapoda, Brachyura) and comparison with brachyuran crabs. Comparative Biochemistry and Physiology Part D: Genomics and Proteomics, 4(4): 290-299.
DOI:10.1016/j.cbd.2009.07.002 |
Komai T, Chang S C, Chan T Y. 2019. A new deep-sea species of the caridean shrimp genus Lebbeus White, 1847 (Crustacea: Decapoda: Thoridae) from Southern Java, Indonesia. Raffles Bulletin of Zoology, 67: 150-159.
DOI:10.26107/RBZ-2019-0012 |
Komai T, Tsuchida S, Segonzac M. 2012. Records of species of the hippolytid genus Lebbeus White, 1847 (Crustacea: Decapoda: Caridea) from hydrothermal vents in the Pacific Ocean, with descriptions of three new species. Zootaxa, 3241(1): 35-63.
DOI:10.11646/zootaxa.3241.1.2 |
Kong L F, Li Y N, Kocot K M, Yang Y, Qi L, Li Q, Halanych K M. 2020. Mitogenomics reveals phylogenetic relationships of Arcoida (Mollusca, Bivalvia) and multiple independent expansions and contractions in mitochondrial genome size. Molecular Phylogenetics and Evolution, 150: 106857.
DOI:10.1016/j.ympev.2020.106857 |
Koopman W J H, Distelmaier F, Smeitink J A, Willems P H. 2013. OXPHOS mutations and neurodegeneration. The EMBO Journal, 32(1): 9-29.
DOI:10.1038/emboj.2012.300 |
Lartillot N, Brinkmann H, Philippe H. 2007. Suppression of long-branch attraction artefacts in the animal phylogeny using a site-heterogeneous model. BMC Evolutionary Biology, 7(Suppl 1): S4.
DOI:10.1186/1471-2148-7-S1-S4 |
Lartillot N, Philippe H. 2004. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Molecular Biology and Evolution, 21(6): 1 095-1 109.
DOI:10.1093/molbev/msh112 |
Lartillot N, Philippe H. 2006. Computing Bayes factors using thermodynamic integration. Systematic Biology, 55(2): 195-207.
DOI:10.1080/10635150500433722 |
Lartillot N, Rodrigue N, Stubbs D, Richer J. 2013. PhyloBayes MPI: phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Systematic Biology, 62(4): 611-615.
DOI:10.1093/sysbio/syt022 |
Laslett D, Canbäck B. 2008. ARWEN: a program to detect tRNA genes in metazoan mitochondrial nucleotide sequences. Bioinformatics, 24(2): 172-175.
DOI:10.1093/bioinformatics/btm573 |
Li C P, de Grave S, Chan T Y, Lei H C, Chu K H. 2011. Molecular systematics of caridean shrimps based on five nuclear genes: implications for superfamily classification. Zoologischer Anzeiger-A Journal of Comparative Zoology, 250(4): 270-279.
DOI:10.1016/j.jcz.2011.04.003 |
Li R Q, Zhu H M, Ruan J, Qian W B, Fang X D, Shi Z B, Li Y R, Li S T, Shan G, Kristiansen K, Li S G, Yang H M, Wang J, Wang J. 2010. De novo assembly of human genomes with massively parallel short read sequencing. Genome Research, 20(2): 265-272.
DOI:10.1101/gr.097261.109 |
Li X Z. 2015. Report on two deep-water caridean shrimp species (Crustacea: Decapoda: Caridea: Alvinocarididae, Acanthephyridae) from the Northeastern South China Sea. Zootaxa, 3911(1): 130-138.
DOI:10.11646/zootaxa.3911.1.8 |
Li Y N, Kocot K M, Schander C, Santos S R, Thornhill D J, Halanych K M. 2015. Mitogenomics reveals phylogeny and repeated motifs in control regions of the deep-sea family Siboglinidae (Annelida). Molecular Phylogenetics and Evolution, 85: 221-229.
DOI:10.1016/j.ympev.2015.02.008 |
Lindner A, Cairns S D, Cunningham C W. 2008. From offshore to onshore: multiple origins of shallow-water corals from deep-sea ancestors. PLoS One, 3(6): e2429.
DOI:10.1371/journal.pone.0002429 |
Lipps J H, Hickman C S. 1982. Origin, age and evolution of Antarctic and deep-sea faunas. In Ernst W G, Morin J G eds. The Environment of the Deep Sea. Prentice Hall, Englewood Cliffs. p. 324-356.
|
Lorion J, Kiel S, Faure B, Kawato M, Ho S Y W, Marshall B, Tsuchida S, Miyazaki J I, Fujiwara Y. 2013. Adaptive radiation of chemosymbiotic deep-sea mussels. Proceedings of the Royal Society B: Biological Sciences, 280(1770): 20131243.
DOI:10.1098/rspb.2013.1243 |
Luo Y J, Gao W X, Gao Y Q, Tang S, Huang Q Y, Tan X L, Chen J, Huang T S. 2008. Mitochondrial genome analysis of Ochotona curzoniae and implication of cytochrome c oxidase in hypoxic adaptation. Mitochondrion, 8(5-6): 352-357.
DOI:10.1016/j.mito.2008.07.005 |
Martinez-Cruz O, Garcia-Carreño F, Robles-Romo A, Varela-Romero A, Muhlia-Almazan A. 2011. Catalytic subunits atpα and atpβ from the Pacific white shrimp Litopenaeus vannamei FOF1 ATP-synthase complex: cDNA sequences, phylogenies, and mRNA quantification during hypoxia. Journal of Bioenergetics and Biomembranes, 43(2): 119-133.
DOI:10.1007/s10863-011-9340-0 |
Mikkelsen N T, Kocot K M, Halanych K M. 2018. Mitogenomics reveals phylogenetic relationships of caudofoveate aplacophoran molluscs. Molecular Phylogenetics and Evolution, 127: 429-436.
DOI:10.1016/j.ympev.2018.04.031 |
Mishmar D, Ruiz-Pesini E, Golik P, Macaulay V, Clark A G, Hosseini S, Brandon M, Easley K, Chen E, Brown M D, Sukernik R I, Olckers A, Wallace D C. 2003. Natural selection shaped regional mtDNA variation in humans. Proceedings of the National Academy of Sciences of the United States of America, 100(1): 171-176.
DOI:10.1073/pnas.0136972100 |
Moritz C, Brown W M. 1987. Tandem duplications in animal mitochondrial DNAs: variation in incidence and gene content among lizards. Proceedings of the National Academy of Sciences of the United States of America, 84(20): 7 183-7 187.
DOI:10.1073/pnas.84.20.7183 |
Norris R D, Kroon D, Klaus A. 2001. Introduction: cretaceous-paleogene climatic evolution of the western North Atlantic, results from ODP Leg 171B, Blake Nose. Proceedings of the Ocean Drilling Program, Scientific Results, 171B.
|
Palero F, Crandall K A, Abelló P, Macpherson E, Pascual M. 2009. Phylogenetic relationships between spiny, slipper and coral lobsters (Crustacea, Decapoda, Achelata). Molecular Phylogenetics and Evolution, 50(1): 152-162.
DOI:10.1016/j.ympev.2008.10.003 |
Parker E S, Gealey W K. 1985. Plate tectonic evolution of the Western Pacific-Indian Ocean region. Energy, 10(3-4): 249-261.
DOI:10.1016/0360-5442(85)90045-3 |
Posada D. 2008. jModelTest: phylogenetic model averaging. Molecular Biology and Evolution, 25(7): 1 253-1 256.
DOI:10.1093/molbev/msn083 |
Rambaut A, Suchard M A, Xie D, Drummond A J. 2014. Tracer v1.6. Accessed at: http://beast.bio.ed.ac.uk/Traceron2020-12-15.
|
Raupach M J, Mayer C, Malyutina M, Wägele J W. 2009. Multiple origins of deep-sea Asellota (Crustacea: Isopoda) from shallow waters revealed by molecular data. Proceedings of the Royal Society B: Biological Sciences, 276(1658): 799-808.
DOI:10.1098/rspb.2008.1063 |
Sanders H L, Hessler R R. 1969. Ecology of the deep-sea benthos. Science, 163(3874): 1 419-1 424.
DOI:10.1126/science.163.3874.1419 |
Shen H, Braband A, Scholtz G. 2013. Mitogenomic analysis of decapod crustacean phylogeny corroborates traditional views on their relationships. Molecular Phylogenetics and Evolution, 66(3): 776-789.
DOI:10.1016/j.ympev.2012.11.002 |
Shi H F, Liu R Y, Sha Z L, Ma J P. 2012. Complete mitochondrial DNA sequence of Stenopus hispidus (Crustacea: Decapoda: Stenopodidea) and a novel tRNA gene cluster. Marine Genomics, 6: 7-15.
DOI:10.1016/j.margen.2011.11.002 |
Shock E L, McCollom T, Schulte M D. 1995. Geochemical constraints on chemolithoautotrophic reactions in hydrothermal systems. Origins of Life and Evolution of the Biosphere, 25(1-3): 141-159.
DOI:10.1007/BF01581579 |
Stamatakis A, Hoover P, Rougemont J. 2008. A rapid bootstrap algorithm for the RAxML Web servers. Systematic Biology, 57(5): 758-771.
DOI:10.1080/10635150802429642 |
Sun S E, Hui M, Wang M X, Sha Z L. 2018a. The complete mitochondrial genome of the alvinocaridid shrimp Shinkaicaris leurokolos (Decapoda, Caridea): Insight into the mitochondrial genetic basis of deep-sea hydrothermal vent adaptation in the shrimp. Comparative Biochemistry and Physiology Part D: Genomics and Proteomics, 25: 42-52.
DOI:10.1016/j.cbd.2017.11.002 |
Sun S E, Sha Z L, Wang Y R. 2018b. Phylogenetic position of Alvinocarididae (Crustacea: Decapoda: Caridea): new insights into the origin and evolutionary history of the hydrothermal vent alvinocarid shrimps. Deep Sea Research Part I: Oceanographic Research Papers, 141: 93-105.
DOI:10.1016/j.dsr.2018.10.001 |
Sun S E, Sha Z L, Wang Y R. 2019a. Divergence history and hydrothermal vent adaptation of decapod crustaceans: a mitogenomic perspective. PLoS One, 14(10): e0224373.
DOI:10.1371/journal.pone.0224373 |
Sun S E, Sha Z L, Wang Y R. 2019b. The complete mitochondrial genomes of two vent squat lobsters, Munidopsis lauensis and M. verrilli: novel gene arrangements and phylogenetic implications. Ecology and Evolution, 9(22): 12 390-12 407.
DOI:10.1002/ece3.5542 |
Takai K, Nakagawa S, Reysenbach A L, Hoek J. 2006. Microbial ecology of mid-ocean ridges and back-arc basins. In: Christie D M, Fisher C R, Lee S M, Givens S eds. Back-Arc Spreading Systems: Geological, Biological, Chemical, and Physical Interactions. American Geophysical Union, Washington DC. p. 185-213.
|
Talavera G, Castresana J. 2007. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Systematic Biology, 56(4): 564-577.
DOI:10.1080/10635150701472164 |
Tsang L M, Chan T Y, Cheung M K, Chu K H. 2009. Molecular evidence for the Southern Hemisphere origin and deep-sea diversification of spiny lobsters (Crustacea: Decapoda: Palinuridae). Molecular Phylogenetics and Evolution, 51(2): 304-311.
DOI:10.1016/j.ympev.2009.01.015 |
Tsoi K H, Chan T Y, Chu K H. 2011. Phylogenetic and biogeographic analysis of the spear lobsters Linuparus (Decapoda: Palinuridae), with the description of a new species. Zoologischer Anzeiger-A Journal of Comparative Zoology, 250(4): 302-315.
DOI:10.1016/j.jcz.2011.04.007 |
Tunnicliffe V, Juniper S K, Sibuet M. 2003. Reducing environments of the deep-sea floor. In: Tyler P A ed. Ecosystems of the World. Elsevier, Amsterdam, Netherlands. p. 81-110.
|
Van Dover C L. 2000. The Ecology of Deep-Sea Hydrothermal Vents. Princeton University Press, Princeton.
|
Vermeij G J. 1987. Evolution and Escalation: an Ecological History of Life. Princeton University Press, Princeton.
|
Vermeij G J. 1995. Economics, volcanoes, and Phanerozoic revolutions. Paleobiology, 21(2): 125-152.
DOI:10.1017/S0094837300013178 |
Wang Z F, Shi X J, Sun L X, Bai Y Z, Zhang D Z, Tang B P. 2017. Evolution of mitochondrial energy metabolism genes associated with hydrothermal vent adaption of alvinocaridid shrimps. Genes & Genomics, 39(12): 1 367-1 376.
DOI:10.1007/s13258-017-0600-1 |
Woolley S N C, Tittensor D P, Dunstan P K, Guillera-Arroita G, Lahoz-Monfort J J, Wintle B A, Worm B, O'Hara T D. 2016. Deep-sea diversity patterns are shaped by energy availability. Nature, 533(7603): 393-396.
DOI:10.1038/nature17937 |
Wyman S K, Jansen R K, Boore J L. 2004. Automatic annotation of organellar genomes with DOGMA. Bioinformatics, 20(17): 3 252-3 255.
DOI:10.1093/bioinformatics/bth352 |
Xin Q, Hui M, Li C L, Sha Z L. 2020. Eyes of differing colors in Alvinocaris longirostris from deep-sea chemosynthetic ecosystems: genetic and molecular evidence of its formation mechanism. Journal of Oceanology and Limnology.
DOI:10.1007/s00343-020-9312-5 |
Yang C H, Bracken-Grissom H, Kim D, Crandall K A, Chan T Y. 2012. Phylogenetic relationships, character evolution, and taxonomic implications within the slipper lobsters (Crustacea: Decapoda: Scyllaridae). Molecular Phylogenetics and Evolution, 62(1): 237-250.
DOI:10.1016/j.ympev.2011.09.019 |
Yang C H, Kumar A B, Chan T Y. 2017. Further records of the deep-sea pandalid shrimp Heterocarpus chani Li, 2006 (Crustacea, Decapoda, Caridea) from southern India. ZooKeys, 685: 151-159.
DOI:10.3897/zookeys.685.13398 |
Yang C H, Sha Z L, Chan T Y, Liu R Y. 2015. Molecular phylogeny of the deep-sea penaeid shrimp genus Parapenaeus (Crustacea: Decapoda: Dendrobranchiata). Zoologica Scripta, 44(3): 312-323.
DOI:10.1111/zsc.12097 |
Yang J S, Lu B, Chen D F, Yu Y Q, Yang F, Nagasawa H, Tsuchida S, Fujiwara Y, Yang W J. 2013. When did decapods invade hydrothermal vents? Clues from the Western Pacific and Indian Oceans. Molecular Biology and Evolution, 30(2): 305-309.
DOI:10.1093/molbev/mss224 |
Yang Z H, Wong W S W, Nielsen R. 2005. Bayes empirical Bayes inference of amino acid sites under positive selection. Molecular Biology and Evolution, 22(4): 1 107-1 118.
DOI:10.1093/molbev/msi097 |
Yang Z H. 2007. PAML 4: phylogenetic analysis by maximum likelihood. Molecular Biology and Evolution, 24(8): 1 586-1 591.
DOI:10.1093/molbev/msm088 |
Yuan M L, Zhang Q L, Guo Z L, Wang L, Shen Y Y. 2015. Comparative mitogenomic analysis of the superfamily Pentatomoidea (Insecta: Hemiptera: Heteroptera) and phylogenetic implications. BMC Genomics, 16(1): 460.
DOI:10.1186/s12864-015-1679-x |
Zachos J, Pagani M, Sloan L, Thomas E, Billups K. 2001. Trends, rhythms, and aberrations in global climate 65 Ma to present. Science, 292(5517): 686-693.
DOI:10.1126/science.1059412 |
Zhang B, Zhang Y H, Wang X, Zhang H X, Lin Q. 2017. The mitochondrial genome of a sea anemone Bolocera sp. exhibits novel genetic structures potentially involved in adaptation to the deep-sea environment. Ecology and Evolution, 7(13): 4 951-4 962.
DOI:10.1002/ece3.3067 |
Zhang J Z, Nielsen R, Yang Z H. 2005. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Molecular Biology and Evolution, 22(12): 2 472-2 479.
DOI:10.1093/molbev/msi237 |
Zhou T C, Shen X J, Irwin D M, Shen Y Y, Zhang Y P. 2014. Mitogenomic analyses propose positive selection in mitochondrial genes for high-altitude adaptation in galliform birds. Mitochondrion, 18: 70-75.
DOI:10.1016/j.mito.2014.07.012 |
 2021, Vol. 39
2021, Vol. 39


