Institute of Oceanology, Chinese Academy of Sciences
Article Information
- XU Yu, ZHAN Zifeng, XU Kuidong
- Morphology and phylogeny of Chrysogorgia pinniformis sp. nov. and C. varians sp. nov., two golden corals from the Caroline seamounts in the tropical Western Pacific Ocean
- Journal of Oceanology and Limnology, 39(5): 1767-1789
- http://dx.doi.org/10.1007/s00343-021-0386-5
Article History
- Received Oct. 10, 2020
- accepted in principle Nov. 13, 2020
- accepted for publication Apr. 15, 2021
2 Southern Marine Science and Engineering Guangdong Laboratory (Zhuhai), Zhuhai 519082, China;
3 University of Chinese Academy of Sciences, Beijing 100049, China
The genus Chrysogorgia Duchassaing & Michelotti, 1864 has a wide latitudinal range, and has been found from Antarctica to the Denmark Straight in a bathymetric range of 10-4 492 m (Watling et al., 2011). Species of Chrysogorgia attach to hard substrates by a strongly calcified discoidal holdfast, or grow in soft bottoms with a calcareous root-like holdfast (Cairns, 2001; Watling et al., 2011; Pérez et al., 2016). The genus now contains 76 species (Xu et al., 2020b). Based on the appearance and shapes of rods or scales in the polyp body wall and tentacles, following the work of Wright and Studer (1889). Versluys (1902) established three groups that have been recognized for the identifiaction of Chrysogorgia species, including group A (Spiculosae), group B (Squamosae aberrantes), and group C (Squamosae typicae) (Versluys, 1902; Cairns, 2001, 2018; Xu et al., 2019). In addition, Cordeiro et al. (2015) established group D (Spiculosae aberrantes), but it was later turned out to be invalid and in fact a part of Versluys' group B due to the mix of sclerite types (Cordeiro et al., 2015; Untiedt et al., 2021). Moreover, Chrysogorgia has five common branching forms, namely: bottlebrush-shaped, planar, bi-flabellate, multi-flabellate, and tree-shaped colonies (Pante and Watling, 2012; Xu et al., 2020a).
Application of a short DNA barcode to discriminate Chrysogorgia species has been a challenge. Although the mitochondrial region mutS homolog (mtMutS), the mostly used barcode, was diagnostic at the genus level of Chrysogorgia, it failed to separate some closely related species of this genus (Xu et al., 2019, 2020a). Recently, 28S nuclear ribosomal gene (28S rDNA) was proved to yield higher species richness accuracy than mtMutS in some octocoral genera, e.g., Alcyonium (McFadden et al., 2014). To date, only one 28S rDNA sequence of Chrysogorgia is available in GenBank, and there is a pressing need to test it as a barcode for this genus with more new sequences.
In the present study, ten specimens of Chrysogorgia were collected during the survey of the benthic diversity on seamounts in the tropical Western Pacific Ocean. Based on morphological and phylogenetic analyses, we describe two new species, C. pinniformis sp. nov. and C. varians sp. nov. Their genetic distances at mtMutS and 28S rDNA as well as phylogenetic relationships within Chrysogorgia species are discussed.
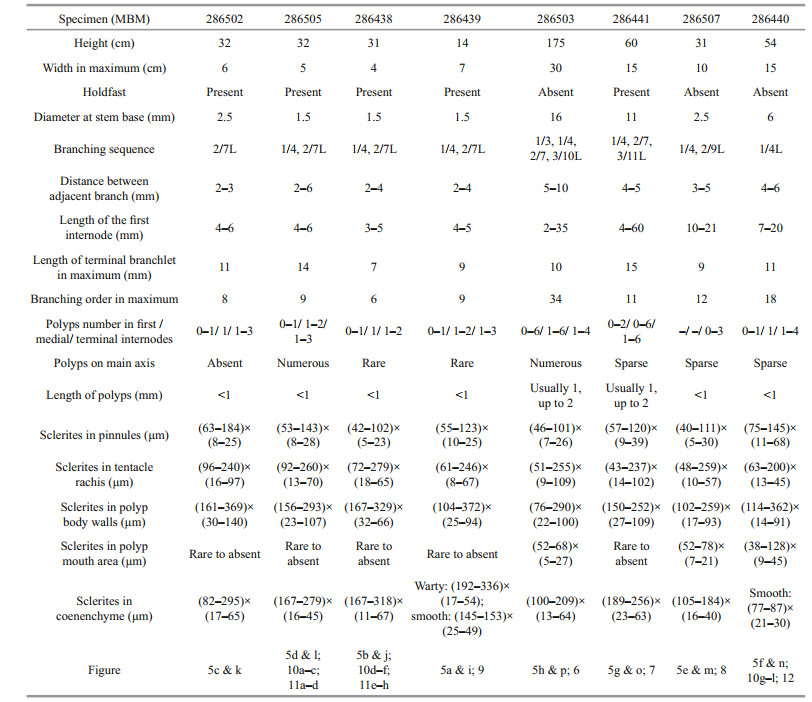
2 MATERIAL AND METHOD 2.1 Specimen collection and morphological examinationTen specimens were obtained by the ROV Faxian (Discovery in Chinese) from seamounts (tentatively named as M5-M7) on the Caroline Ridge in the tropical Western Pacific during the cruise of the R/V Kexue (Science in Chinese) in 2019 (Table 1; Fig. 1). They were photographed in situ before being sampled, photographed on board and then stored in 75% ethanol after collection. A few branches were stored at -80 ℃ for molecular study.
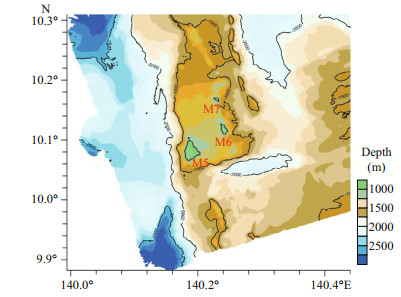
|
| Fig.1 Sampling site on the Caroline seamounts (M5-M7) in the Western Pacific Ocean |
A stereo dissecting microscope was used to examine the general morphology and anatomy. The preparation of sclerites for imaging followed Xu et al. (2020a). A TM3030Plus Scanning Electron Microscope (SEM) was used to obtain the SEM scans and the optimum magnification was chosen for each kind of sclerite. The morphological terminology used follows Bayer et al. (1983).
The type specimen of the new species has been deposited in the Marine Biological Museum of Chinese Academy of Sciences (MBMCAS) at Qingdao, China.
2.2 DNA extraction and sequencingGenomic DNA was extracted from the polyps of the ten specimens using DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany) following instructions. The mitochondrial region mutS homolog (mtMutS) and an approximately 750-nt fragment of the 28S nuclear ribosomal gene (28S rDNA) were selected for the phylogenetic analysis. To amplify mtMutS, the primers AnthoCorMSH (5'-AGGAGAATTATTCTA-AGTATGG-3'; Herrera et al., 2010) and Mut-3458R (5'-TSGAGCAAAAGCCACTCC-3'; Sánchez et al., 2003) were used. PCR reactions were conducted following Xu et al. (2020a). We sequenced 28S rDNA using primers 28S-Far and 28S-Rar (McFadden and van Ofwegen, 2013), and the same PCR protocol used for mtMutS. PCR purification and sequencing were performed by TsingKe Biological Technology (TsingKe Biotech, Beijing, China).
2.3 Genetic distance and phylogenetic analysesAll the Chrysogorgia sequences and the related chrysogorgiid genera and out-group species Chromoplexaura marki, Alaskagorgia aleutiana, and Plexaura kuna were downloaded from GenBank. The sequences containing sequencing errors (usually marked with "n" or "y") or from unclassified species (e.g. Chrysogorgia sp.) or without associated publications or from duplicate isolates were excluded from the molecular analyses. The sequences were aligned using MAFFT v.7 (Katoh and Standley, 2013) with the G-INS-i and Q-INS-i algorithms for the mtMutS and 28S rDNA regions, respectively. The nucleotide alignment was trimmed to equal length using BioEdit v7.0.5 (Hall, 1999). Genetic distances between species/populations were calculated with MEGA 6.0 using Kimura 2-parameter model (Tamura et al., 2013).
For the phylogenetic construction, when conspecific sequences were identical, one was chosen randomly for analysis. The best-fitted model GTR+G was selected with the Akaike information criterion in jModeltest2 (Darriba et al., 2012) for both the mtMutS and 28S rDNA alignments. Maximum likelihood (ML) analysis was performed using PhyML-3.1 (Guindon et al., 2010). With node support from a majority-rule consensus tree of 1 000 bootstrap replicates. Following Hillis and Bull (1993), the ML bootstraps < 70%, 70%-94%, and ≥95% were considered as low, moderate and high, respectively. Bayesian inference (BI) analysis was carried out using MrBayes v3.2.7a (Ronquist and Huelsenbeck, 2003). Posterior probability was estimated based on 10 000 000 Monte Carlo Markov Chain (MCMC) generations (×4 chains) sampling every 1 000 generations (burn-in=25%). Convergence of the MCMC was assessed using Tracer 1.4.1 (Rambaut and Drummond, 2007). Following Alfaro et al. (2003), the Bayesian posterior probabilities < 0.95 and ≥0.95 were considered as low and high, respectively.
3 RESULT 3.1 TaxonomyClass Anthozoa Ehrenberg, 1834 Subclass Octocorallia Haeckel, 1866 Order Alcyonacea Lamouroux, 1812 Suborder Calcaxonia Grasshoff, 1999 Family Chrysogorgiidae Verrill, 1883
Genus Chrysogorgia Duchassaing & Michelotti, 1864
Chrysogorgia pinniformis sp. nov. (Figs. 2-4)
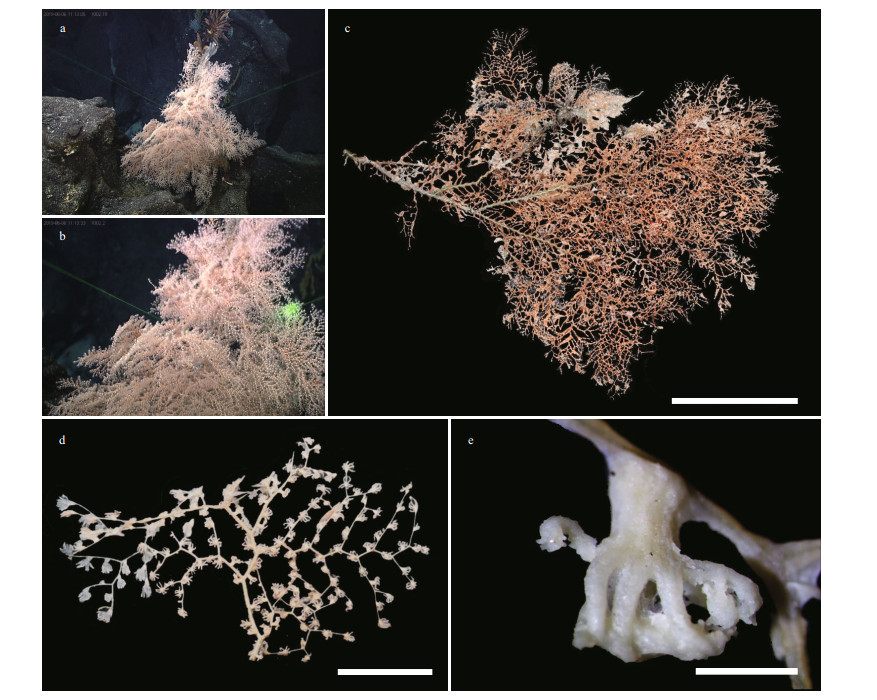
|
| Fig.2 The external morphology and polyps of Chrysogorgia pinniformis sp. nov. a. the holotype in situ; b. close-up of branches and polyps in situ; c. the holotype after collection; d. a terminal branchlet under a light microscope; e. a single non-terminal polyp under a light microscope. Scale bar: 20 cm (c); 5 cm (d); 1 mm (e). |
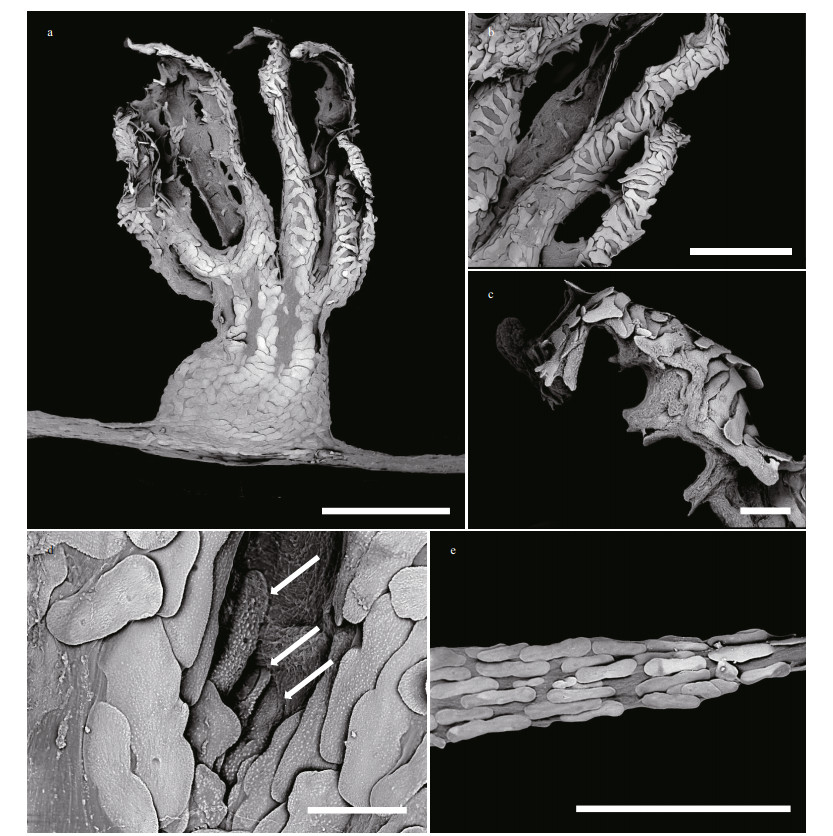
|
| Fig.3 Polyp and coenenchyme of Chrysogorgia pinniformis sp. nov. under SEM a. a single non-terminal polyp under SEM; b. tentacular part under SEM; c. tentacular rachis and proximal part of pinnule under SEM; d. sclerites in mouth area of polyps under SEM (the arrow); e. coenenchyme under SEM. Scale bar: 1 mm (a); 500 μm (b, e); 100 μm (c, d). |

|
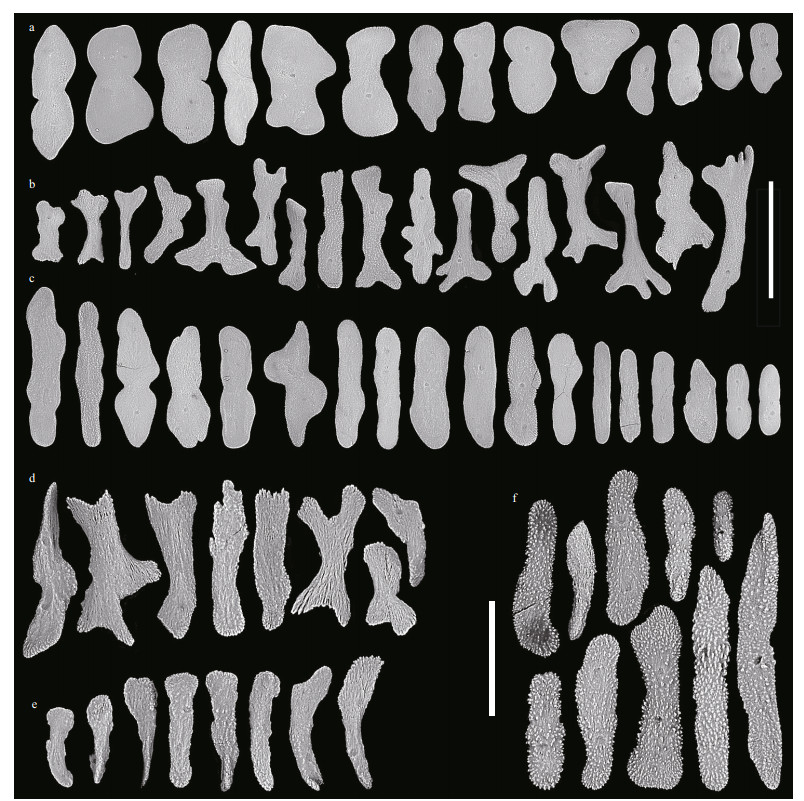
| Fig.4 Sclerites of Chrysogorgia pinniformis sp. nov. a. sclerites in polyp body wall; b. sclerites in the back of tentacle rachis; c. sclerites in coenenchyme; d. sclerites in the joint part between tentacle rachis and pinnules; e. sclerites in pinnules; f. sclerites in mouth area of polyps. Scale bar: 200 μm (a, b, c; all in the same scale); 100 μm (d, e, f; all in the same scale). |
urn: lsid: zoobank.org: act: 74FE8B57-1038-49CC-83E2-CF05BB27C544
Material examined Holotype: MBM286504, station FX-Dive 218 (10°07'48"N, 140°14'23"E), M6 seamount, 1 002 m, 6 June 2019.
Diagnosis Chrysogorgia in Versluys' group C (Squamosae typicae) with a bushy colony. Large branches pinnately branched and forming multiple fans with its small branches regularly and quasi-dichotomous branched. Polyps nearly perpendicular to the branches, with tentacles expanded usually without contraction. Scales in the tentacle rachis transversely and obliquely arranged, branched with various shapes. Scales in pinnules longitudinally arranged, small and cuneate. Scales in polyp body wall longitudinally and transversely arranged, with a medial contraction. Scales and rods in mouth area of polyps longitudinally arranged, with many small warts on surface. Scales in coenenchyme elongate and longitudinally arranged.
Description Specimen of the holotype large, about 69-cm tall and 53-cm wide, including three main large branches (Fig. 2b & c). Stem with a golden metallic luster, about 8 mm in diameter at base. The colony is made up of multiple, irregularly arranged fans. Each major branch zigzags and produces small lateral branches in an alternate pinnate manner. The lateral branches re-divide in a dichotomous or quasi-dichotomous manner, with distance between adjacent branches 3-6 mm and parallel branches 4-14 mm (Fig. 2d). Branchlets diverge in an alternating opposite fashion and straight up to 45-mm long and bear up to 10 polyps, after which they terminate or continue to branch in a dichotomous fashion. Polyps about 2.0- 2.5-mm long and 1-mm wide at base, sometimes thin and translucent, well spaced on branchlets (Figs. 2d-e & 3a). Polyps usually perpendicular to branchlets, arranged up to three in the branchlet internodes and up to eight in the larger branch internodes. No polyps present in the smallest internodes but a polyp present at each node. Tentacular part usually spread with eight obvious tentacles without contraction after collection, up to 2-mm long (Figs. 2e & 3a).
Sclerites with same forms and arrangements in non-terminal and terminal polyps in this species. Scales in polyp body wall longitudinally and obliquely arranged, smooth with an obvious medial contraction, measuring (54-231) μm×(29-142) μm (length×width, the same below, Figs. 3a & 4a). Scales in aboral region of tentacle rachis transversely and obliquely arranged, elongate, and branched with various shapes and relatively coarse surface, measuring (75-298) μm× (15-145) μm (Figs. 3b & 4b). Scales in pinnules longitudinally arranged, small and cuneate, measuring (74-232) μm×(15-58) μm (Figs. 3c & 4e). Scales in joint part between tentacle rachis and pinnules longitudinally arranged, usually branched, with one or both ends forked, sometimes extending onto the proximal part of the pinnule, measuring (92-151) μm× (35-61) μm (Figs. 3c & 4d). Scales and rods in mouth area on the outside of polyp body radially arranged, elongate and thick, both with many small conical and densely arranged warts on surface, measuring (70-238) μm×(15-52) μm (Figs. 3d & 4f). Scales in coenenchyme longitudinally arranged, elongate and often smooth, occasionally with a medial contraction, measuring (72-293) μm×(13-113) μm (Figs. 3e & 4c). Sclerites of some polyps sparse or rare.
Type locality A seamount (tentatively named as M6) located on the Caroline Ridge with water depth of 1 692 m.
Etymology The Latin adjective pinniformis (pinniform) refers to the pinnate branches of this colony.
Distribution and habitat Found from an unnamed seamount located on the Caroline Ridge in the tropical Western Pacific Ocean, where the water depth was 1 692 m, the water temperature about 4.85 ℃ and the salinity about 36.7. Colony very bushy in situ and attached to a rocky substrate (Fig. 2a & b).
Remarks Chrysogorgia pinniformis sp. nov. belongs to Versluys' group C (Squamosae typicae) and is similar to C. electra Bayer & Stefani, 1988, C. scintillans Bayer & Stefani, 1988, and C. binata Xu, Zhan & Xu, 2019 by having some fan-shaped structure, but differs from them by having regular scales in polyp body wall (vs. irregular in all), and warty scales and rods present in the polyp (vs. absent in all). It differs from the species with a bottlebrush-shaped colony (clockwise or counterclockwise) in the Squamosae typicae group by being a bushy colony composed of multiple, irregularly arranged fans. Chrysogorgia pinniformis sp. nov. is similar to C. japonica (Wright & Studer, 1889) (unknown colony shape) by some dichotomous branching manner, but differs by scales in tentacle rachis (transversely and obliquely arranged, branched with various shapes vs. longitudinally arranged, rod-like with blunt ends, sometimes with a short fork at one end), and sclerites in coenenchyme (elongate scales vs. broad spindles with lancet-shaped bodies). Within the genus Chrysogorgia, only the new species and Chrysogorgia pinnata Cairns, 2007 possess pinnate branching. However, Chrysogorgia pinnata belongs to the Versluys' group A (Spiculosae).
Chrysogorgia varians sp. nov. (Figs. 5-13; Tables 2 & 3)

|
| Fig.5 The external morphology of Chrysogorgia varians sp. nov. The specimens MBM286439 (a, i), MBM286438 (b, j), MBM286502 (c, k), MBM286505 (d, l), MBM286507 (e, m), MBM286440 (f, n), MBM286441 (g, o), MBM286503 (h, p) in situ (a-h) and after collection (i-p). Laser dots spaced at 33 cm used for measuring scale. Scale bar: 50 cm (p), 20 cm (i-o, all in the same scale). |
|
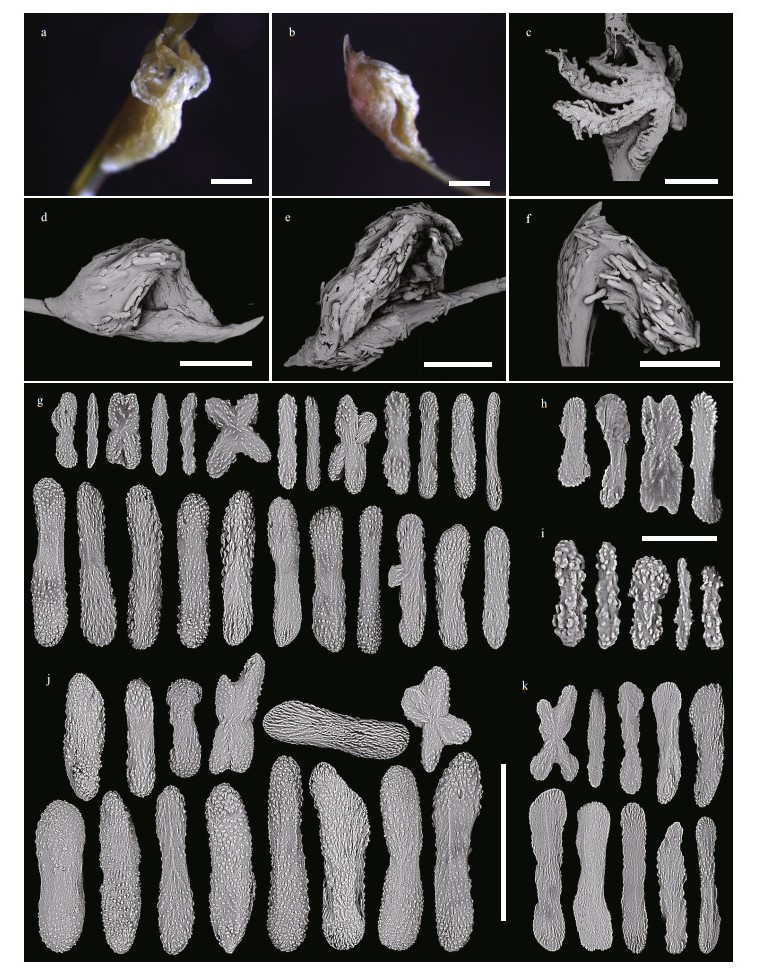
| Fig.6 Polyps and sclerites of C. varians sp. nov. (MBM286503) a. a single non-terminal polyp under a light microscope; b. a single terminal polyp under a light microscope; c. a single non-terminal polyp under SEM; d-f. a single terminal polyp under SEM; g. sclerites in tentacle rachis; h. sclerites in pinnules; i. sclerites in polyp mouth; j. sclerites in polyp body wall; k. sclerites in coenenchyme at the base of polyps. Scale bar: 500 μm (a-f), 200 μm (g, j, k in the same scale); 50 μm (h, i in the same scale) |

|
| Fig.7 Polyps and sclerites of C. varians sp. nov. (MBM286441) a. two polyps under a light microscope; b. a single non-terminal polyp under SEM; c. a single terminal polyp under SEM; d. sclerites in polyps; e. sclerites in pinnules; f. sclerites in coenenchyme at the base of polyps. Scale bar: 500 μm (a-c), 200 μm (d, f in the same scale); 100 μm (e). |

|
| Fig.8 Polyps and sclerites of Chrysogorgia varians sp. nov. (MBM286507) a. a single non-terminal polyp under a light microscope; b. a single non-terminal polyp under SEM; c. a single terminal polyp under SEM; d. sclerites in polyps; e. sclerites in coenenchyme at the base of polyps; f. sclerites in pinnules; g. sclerites in the polyp mouth. Scale bar: 500 μm (a-c), 100 μm (d, e in the same scale); 50 μm (f, g). |
|
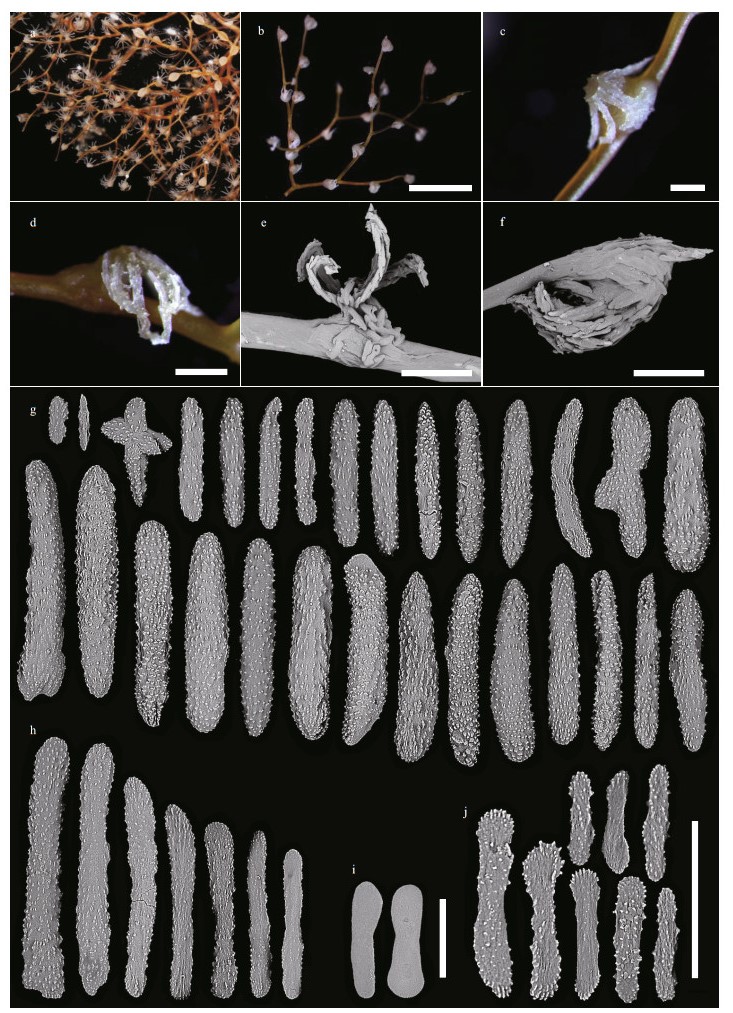
| Fig.9 Polyps and sclerites of Chrysogorgia varians sp. nov. (MBM286439) a. branches and polyps after collection; b. a branch after fixation; c & d. a single non-terminal polyp under a light microscope; e. a single non-terminal polyp under SEM; f. a single terminal polyp under SEM; g. sclerites in polyps; h. sclerites in coenenchyme at the base of polyps; i. sclerites in coenenchyme between inter-polyps; j. sclerites in pinnules. Scale bar: 5 mm (b); 500 μm (c-f), 100 μm (g-i, j; g-i in the same scale). |
|
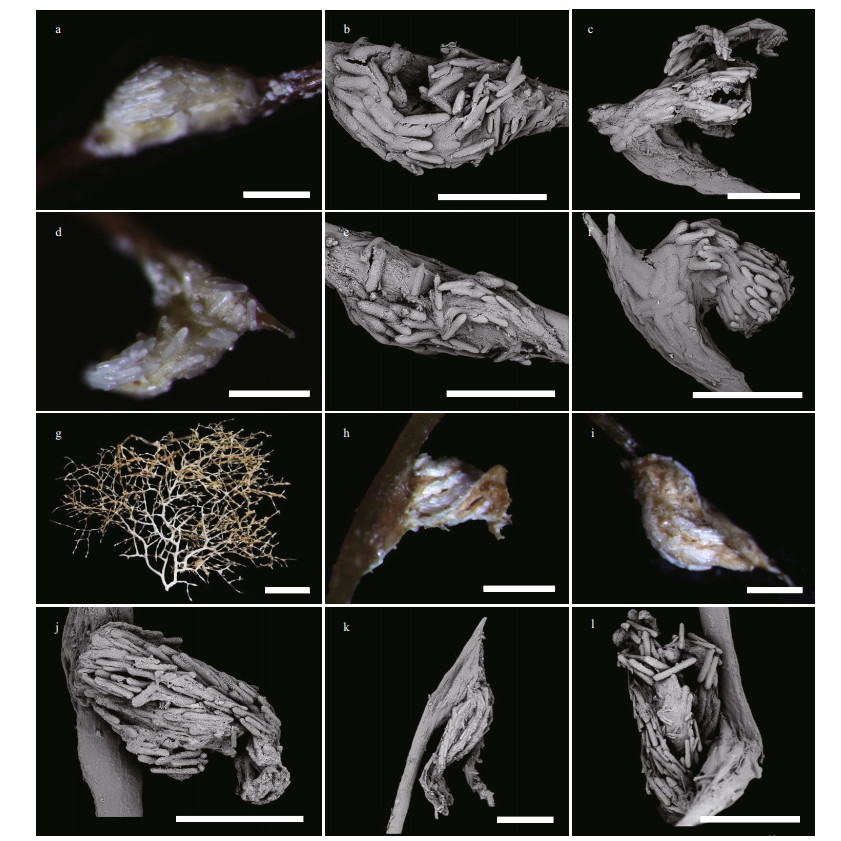
| Fig.10 Branches and polyps of Chrysogorgia varians sp. nov. a, d, h, i. a single polyp under a light microscope; b, e, j. a non-terminal polyp under SEM; c, f, k, l. a terminal polyp under SEM; g. an irregular branch with anastomose; specimens MBM286505 (a-c), MBM286438 (d-f), and specimen MBM286440 (g-l). Scale bar: 2 cm (g); 500 μm (a-f, h-l). |

|
| Fig.11 Sclerites of Chrysogorgia varians sp. nov. a & e. sclerites in pinnules; b & f. sclerites in tentacle rachis; c & g. sclerites in the polyp body wall; d & h. sclerites in coenenchyme at the base of polyps; MBM286505 (a-d) and MBM286438 (e-h), respectively. Scale bar: 200 μm (b-c, f-h in the same scale); 100 μm (a, e in the same scale). |
|
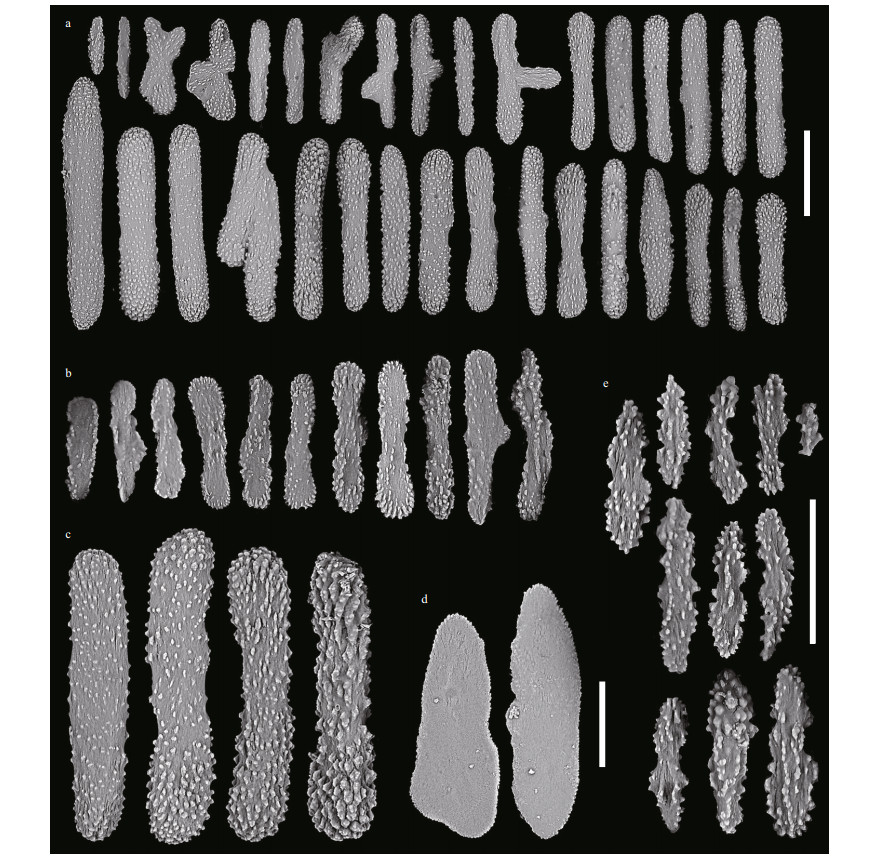
| Fig.12 Sclerites of Chrysogorgia varians sp. nov. (MBM286440) a. sclerites in polyps; b. sclerites in pinnules; c. four typical rods with different warts; d. sclerites in coenenchyme between inter-polyps; e. sclerites in polyp mouth. Scale bar: 200 μm (a); 100 μm (b-d; b-d in the same scale), 50 μm (e). |
|
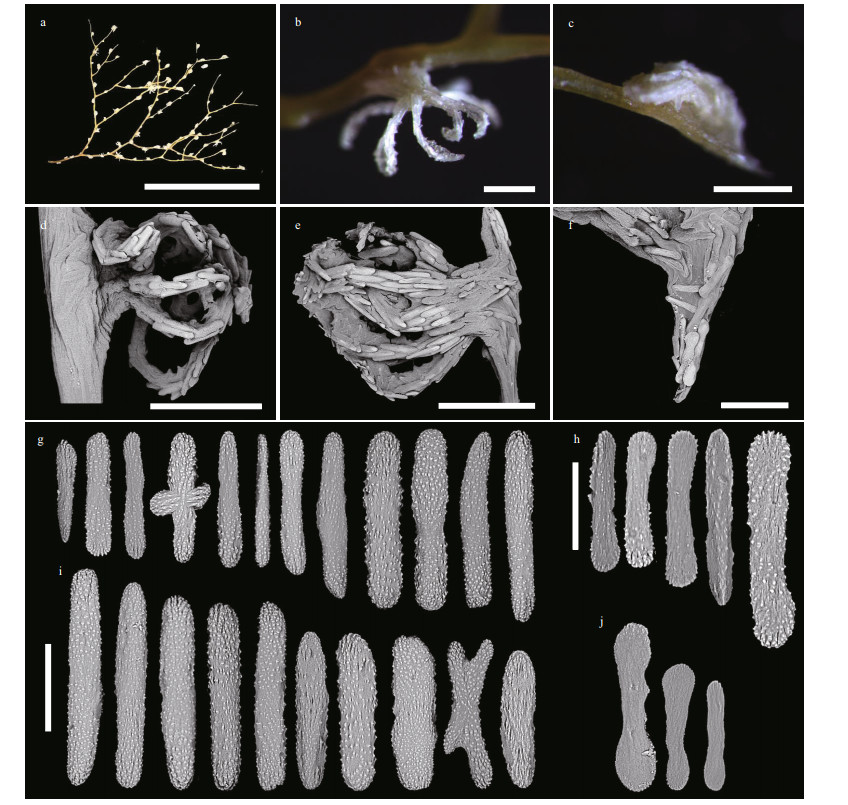
| Fig.13 Polyps and sclerites of Chrysogorgia varians sp. nov. (MBM286506) a. a branch after fixation; b. a single non-terminal polyp under a light microscope; c. a single terminal polyp under a light microscope; d. a single non-terminal polyp under SEM; e. a single terminal polyp under SEM; f. sclerites in coenenchyme at the base of a terminal polyp; g. sclerites in tentacle rachis; h. sclerites in pinnules; i. sclerites in polyp body wall; j. sclerites in coenenchyme at the base of polyps. Scale bar: 5 cm (a); 500 μm (b-e), 200 μm (f); 100 μm (g, i, and j in the same scale); 50 μm (h) |
|
urn: lsid: zoobank.org: act: 96111211-8D7B-40EF-BFF4-1A2D78DE5247
Material examined Holotype: MBM286503, station FX-Dive 217 (10°06'49"N, 140°15'00"E), M6 140°11'57"E), M5 seamount, 884 m, 28 May 2019; MBM286507, station FX-Dive 211 (10°03'N, 140°10'E), M5 seamount, 1384-1 482 m, 29 May 2019; MBM286438, station FX-Dive 213 (10°04'28"N, 140°11'36"E), M5 seamount, 899 m, 31 May 2019; MBM286439, station FX-Dive 215 (10°04'44"N, 140°11'04"E), M5 seamount, 865 m, 2 June 2019; MBM286440, station FX-Dive 216 (10°05'23"N, 140°11'12"E), M5 seamount, 902 m, 4 June 2019; MBM286441, station FX-Dive 223 (10°04'38"N, 140°15'08"E), M7 seamount, 1 071 m, 11 June 2019.
Diagnosis Chrysogorgia in Versluys' group A (Spiculosae) with a bottlebrush-shaped colony and a varied branching sequence in different growth stages. Branches subdivided dichotomously, occasionally anastomose in some old branches. Polyps are small with tentacles extended or not visible. Rods in aboral face of tentacle rachis longitudinally arranged, regular with many small conical or ridge-like warts on surface and two rounded ends. Scales in pinnules and the end of tentacles usually longitudinally arranged, slender and often having many warts and ridges, and conspicuously toothed margins at the rounded ends, occasionally nearly smooth and with irregular shape. Rods in polyp body wall same as those of tentacle rachis, longitudinally or obliquely arranged, some of them relatively larger than those in tentacles. Rods near the polyp mouth area rare to absent, small and usually with elongate and ridge-like warts. Scales in coenenchyme at the base of polyps slender and a little coarser with more or less conical warts on surface, and in inter-polyps rare to absent, occasionally smooth with two rounded ends, all of them longitudinally and obliquely arranged.
Description Specimen of the holotype having a large bottlebrush-shaped colony, about 175-cm tall and 30-cm wide, with the holdfast not recovered (Fig. 5h & p). Stem about 16 mm in diameter at base. Branching sequence irregular, including 1/3L, 1/4L, 2/7L, and 3/10L. Distance between adjacent branch about 5-10 mm, and the first internode of branch up to 35-mm long. Branches subdivided dichotomously, up to 34 orders, occasionally irregular in some old branches. Polyps small, usually 1-mm long, sometimes up to 2 mm, with tentacles extended or not visible and forming a conical shape (Fig. 6a-f). Sometimes the oral region is angled, upwards or towards the periphery, rarely downwards (Fig. 6c). Polyps usually zero to six on first internode, up to six in the other internodes. Polyps on the main axis are small and numerous, and nematozooids are absent.
Rods in aboral face of tentacle rachis longitudinally arranged, regular with many small warts on surface and two rounded ends, rarely with one sharp end, some of them crosses, measuring (51-255) μm×(9-109) μm (Fig. 6g). These warts usually conical and densely arranged, sometimes large and ridge-like, and longitudinally or transversely arranged on the sclerites surface. Scales in pinnules and the end of tentacles usually longitudinally arranged, sparse, small and slender, often having many warts and ridges and conspicuously toothed margins at the rounded ends, occasionally nearly smooth and with irregular shape, measuring (46-101) μm×(7-26) μm (Fig. 6h). The warts on the surface of the pinnule's scales commonly elongate and ridge-like, sometimes in rows. Rods in polyp body wall same as those of tentacles rachis, longitudinally or obliquely arranged, some of them relatively larger and stout with more blunt ends, measuring (76-290) μm×(22-100) μm (Fig. 6j). Rods in the polyp mouth area rare to absent, small with many warts on surface, measuring (52-68) μm×(5-27) μm (Fig. 6i). These warts usually elongate and ridge-like, sometimes cylindrical or conical, arranged relatively densely and sometimes in rows. Scales usually present in coenenchyme at the base of terminal polyp, while rare to absent in other polyps, longitudinally and obliquely arranged. Scales in coenenchyme at the base of polyps slender and a little coarser with more or less conical warts on surface, and in inter-polyps rare to absent, some of them almost smooth with two rounded ends, measuring (100-209) μm×(13-64) μm (Fig. 6k).
Variation of paratype The specimens of paratype have relatively complete colony except the specimen MBM286506. For their morphological measurements, see Table 2. There are some differences compared with the holotype, such as varied branching sequence, polyp's size, and numbers in internodes, and the length of the first internode (Table 2). Some another differences among them include: the polyps of specimens MBM286502, MBM286505, and MBM286438 usually form a conical or columnar shape (Fig. 10a & d); the branches of the specimen MBM286440 are more irregular and occasionally anastomose in some old branches (Fig. 10g); the specimen MBM286507 was possibly dying when it was collected with a few polyps and almost exposed branches (Fig. 5m); the numbers of the sclerites in pinnules are more sparse in the holotype; the sclerites in polyp mouth area are rare in specimens MBM286502, MBM286505, MBM286438, MBM286439, and MBM286441; the sclerites in the coenenchyme at the base of terminal polyps are more slender and coarse in specimen MBM286502, MBM286505, MBM286438, and MBM286439 (Figs. 9 & 11); the sclerites in polyps are more slender in specimen MBM286440 (Fig. 12).
Type locality A seamount (tentatively named as M6) located on the Caroline Ridge with water depth of 1 145 m.
Distribution and habitat Found from the Caroline seamounts (tentatively named as M5-M7) with water range of 832-1 482 m. Colonies attached to rocky substrate. The water temperatures were about 4.13-5.27 ℃ and the salinity about 36.6-36.8.
Etymology The Latin adjective varians (varied) refers to the varied branching sequence in different growth stages of this species.
Remarks Chrysogorgia varians sp. nov. belongs to Versluys' group A (Spiculosae) with a bottlebrush-shaped colony. It is characterized by warty rods and elongate scales in the tentacles, scales often with many warts and ridges and conspicuously toothed margins at the rounded ends in the pinnules, and small rods with elongate and ridge-like warts in the polyp mouth area. The nine specimens collected from the Caroline seamounts all have a bottlebrush-shaped colony and the same sclerite forms (Figs. 5-13). Thus, we identified them as the same species. As mentioned above, there are some differences among the nine specimens of C. varians sp. nov. However, these character differences are not constant and may be caused by different growth stage or environment or inadequate measurement, and we thus treated as the intraspecific variation.
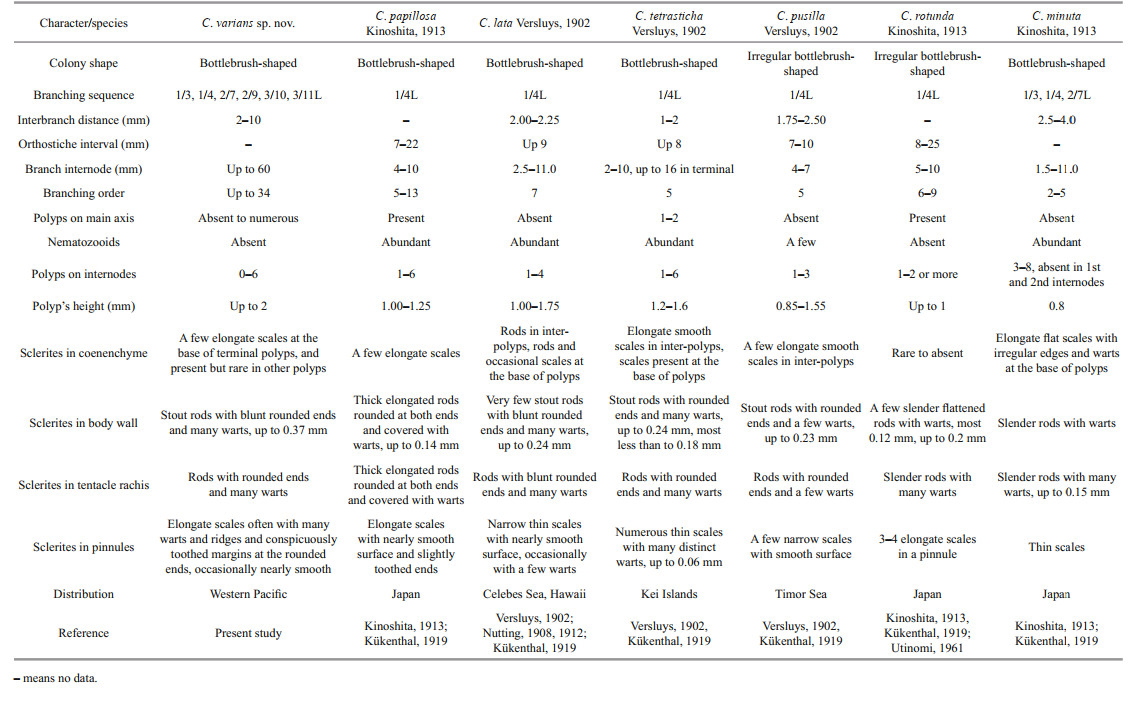
The sclerite forms and branching sequences of C. varians sp. nov. are similar to C. papillosa Kinoshita, 1913 and C. minuta Kinoshita, 1913 but it differs from C. papillosa by scales in pinnules often with many warts and ridges on surface (vs. nearly smooth), large stout rods in polyp body wall (up to 0.37 mm vs. small, thick and elongated rods, up to 0.14 mm), small warty rods present in the mouth area of polyps (vs. absent) and nematozooids absent in polyps and branches (vs. abundant) (Kinoshita, 1913) (Table 3). Chrysogorgia varians sp. nov. can be separated with C. minuta by larger and wider rods in polyps (vs. relatively small and slender), elongate scales with less warts on surface at the base of polyps (vs. stout with many warts and irregular outlines) and small warty rods present in the mouth area of polyps (vs. absent) (Kinoshita, 1913) (Table 3). It is also similar to C. lata Versluys, 1902 and C. tetrasticha Versluys, 1902, but differs from the former by rare scales in coenenchyme between polyps (vs. rare rods), elongate scales in the tip of tentacles (vs. rods), and small warty rods present in the mouth area of polyps (vs. smooth scales), and from the latter by scales in pinnules (relatively large, often with many warts and ridges and conspicuously toothed margins at the rounded ends vs. small and without ridges and conspicuously toothed margins), and small warty rods present in the mouth area of polyps (vs. absent) (Table 3).
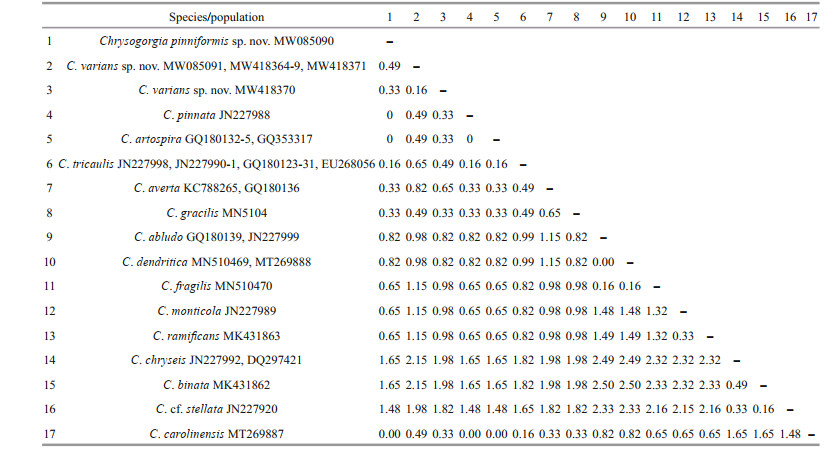
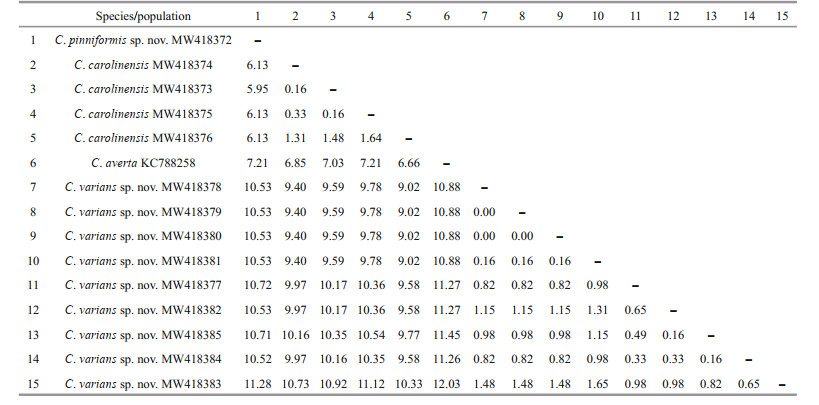
3.2 Genetic distance and phylogenetic analysesThe new sequences were deposited in GenBank (Table 1). The alignments comprised 623 and 647 nucleotide positions for the mtMutS and 28S rDNA regions, respectively. Based on the mtMutS aligned region, no intraspecific variability was observed, and the interspecific distances range from zero to 2.44% for the Chrysogorgia (Table 4). For mtMutS, the genetic distances between the new species C. pinniformis sp. nov. and 15 of its congeners are in the range of 0-1.65%, and those between C. varians sp. nov. and 15 of its congeners are in the range of 0.33%-2.15% (Table 4). Unlike the mtMutS database, only 28S rDNA sequences of four identified Chrysogorgia species are available, and the intraspecific data were obtained from only C. carolinensis and C. varians sp. nov. (Table 5). The intraspecific distances were in range of 0.16%-1.64% for C. carolinensis and 0-1.65% for C. varians sp. nov., while the interspecific distances ranged from 5.95% to 17.90% (Table 5).
|
|
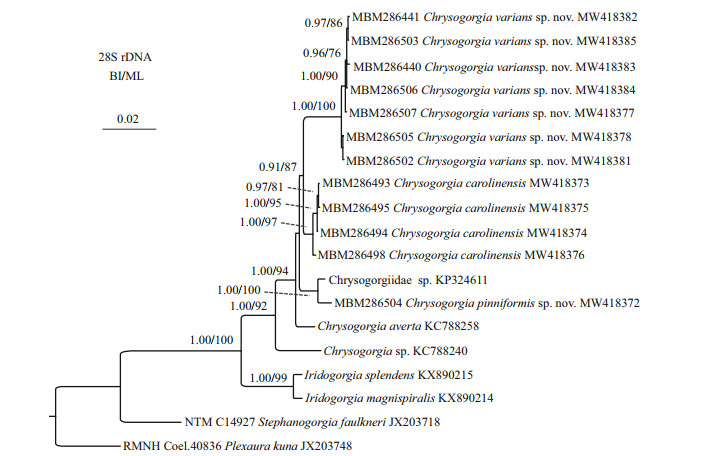
The ML tree is nearly identical with the BI tree in topology for both the mtMutS and 28S rDNA regions, and thus a single tree with both support values was shown for each of the markers (Fig. 14 & Supplementary Fig.S1). The two new species nested into the Chrysogorgia clade in both of the mtMutS and 28S rDNA trees with low to high support. In the mtMutS tree, all the Chrysogorgia species formed two main sister clades (Clade I and II) with moderate support, but most phylogenetic relationships within the two main clades were not resolved (Supplementary Fig.S1). In the 28S rDNA trees, all the Chrysogorgia species formed a monophyletic clade, and all the conspecific specimens clustered together with high support (ML 97%-100%; BI 1.00). Chrysogorgia varians sp. nov. and C. carolinensis formed a sister clade to the group C. pinniformis sp. nov. + Chrysogorgiidae sp. KP324611 with low support (Fig. 14).
|
| Fig.14 Bayesian inference (BI) tree constructed by 28S rDNA showing phylogenetic relationships among Chrysogorgia species The maximum likelihood (ML) tree is identical with the BI tree in topology. Node support is as follows: BI posterior probability /ML bootstrap. The ML bootstrap < 70% or BI posterior probability < 0.90 is not shown. The GenBank accession numbers of the 28S rDNA sequences were listed next to the species names. |
The assignment of the new species to the genus Chrysogorgia was supported by the morphology and molecular phylogenetic analysis. The mtMutS region is one of the most commonly used makers in an integrative identification of octocorals (McFadden et al., 2011; McFadden and van Ofwegen, 2013; Xu et al., 2019, 2020a, b). However, there is no barcoding gap for delimitating Chrysogorgia species, and, additionally, no genetic variability was observed among C. pinniformis sp. nov., C. carolinensis, C. pinnata, and C. artospira, indicating its very limited usefulness for species delimitation (Table 4). In contrast, the 28S rDNA showed higher level of genetic variation (e.g. interspecific distances 5.95- 12.03 vs. 0-2.5 at mtMutS). The 28S rDNA genetic distances among C. varians sp. nov. specimens were in the range of 0-1.65, similar to the intraspecific distances of C. carolinensis (0.16-1.64), supporting the conspecific assignment of the varians-like specimens. Although only 28S rDNA sequences of four Chrysogorgia species are available, the present distance analysis indicated the 28S rDNA region as a potential barcode for this genus. The genetic distances between C. varians sp. nov. and the congeners are relatively higher (in range of 0.33-0.49 at mtMutS), confirming its validity.
It is worth noting that the specimens of C. varians sp. nov. have different morphological features, such as the varied branching sequence, polyp's shape, size and numbers in internodes, and the length of the first internode (Table 2). Considering the diameter of stem base, branching order, and the colony size, these specimens showed a series of growth stages: the specimen MBM286438, MBM286439, MBM286502, and MBM286505 are likely juveniles with narrow stem base, less branching order and short colonial height, and MBM286507 is likely an intermediate state, while MBM286441, MBM286443, and MBM286503 are likely adults with the wider stem base, more branching order, and higher colonial height. Therefore, the branching sequence of C. varians sp. nov. often shows 1/4 and 2/7L in juveniles, and becomes more and more irregular (1/3, 1/4, 2/7, 2/9, 3/10, 3/11L) with continued growth. The lengths of the first internode are short (3-6 mm) in juvenile, and gradually becomes longer and longer from bottom to top which can up to 60 mm. The polyps are usually small (less than 1 mm) and few in number (1-3) in the internodes of the juveniles, while they can be up to 2 mm and up to 6 in the internodes of the old specimens. Furthermore, the numbers of sclerites in the pinnules, polyp mouth area, and the coenenchyme at the base of polyps are also different, but they both have same forms in the same part of the different specimens. Consequently, compared to the changing external morphological features, the sclerite forms show small variation in the growth stages and can be used as a main character to identify the species (Xu et al., 2020b).
Branching sequence as a relatively constant trait is often used as a diagnostic character for a species of Chrysogorgia (e.g. Cairns, 2001; Pante and Watling, 2012). It is known to be varied in some species, however, even within a single specimen due to the different growth stage and environment, like C. herdendorfi Cairns, 2001 (2/5R-3/8R), C. intermedia Versluys, 1902 (1/4R-1/7R), C. squamata (Verrill, 1883) (1/5R-1/7R), C. minuta Kinoshita, 1913 (1/3L, 1/4L, 2/7L), C. abludo Pante & Watling, 2012 (1/3L, 1/4L, irregular) and the species C. varians sp. nov. described here (Untiedt et al., 2021). It is almost certain that it varies in many other species, especially those established colony fragments, but has gone unnoticed. Furthermore, as mentioned above, the features like polyp's shape, size, and numbers in internodes, and the length of internode are also affected by the different growth stage and environment. Therefore, all of these features can be a secondary characteristic to help species identification, and the sclerites should be considered first (Xu et al., 2020b). In addition, it means there will be many synonyms in Chrysogorgia, which needs more new specimens and molecular data to support and verify, then revise this genus.
Including the new species here, the genus Chrysogorgia contains 78 species. Among them, 53 species are found only from the Pacific, 17 species from the Atlantic and seven only from the Indian Ocean, and Chrysogorgia flexilis (Wright & Studer, 1889) from both the Pacific and Indian Oceans (Cairns, 2001, 2018; Xu et al., 2019, 2020a, b). In the joining area of Indo-West Pacific, there are 42 species recorded, including 20 species (C. bracteata, C. geniculata, C. rigida, C. ramosa, C. axillaris, C. cupressa, C. tetrasticha, C. sibogae, C. pendula, C. lata, C. mixta, C. calypso, C. chryseis, C. anastomosans, C. orientalis, C. curvata, C. pentasticha, C. octagonos, C. intermedia, and C. pusilla) from the Malay Archipelago, 14 species (C. dispersa, C. pyramidalis, C. rotunda, C. papillosa, C. minuta, C. okinosensis, C. comans, C. sphaerica, C. debilis, C. pellucida, C. versluysi, C. excavata, C. japonica, and C. cavea) from the Japanese waters, and eight species (C. ramificans, C. binata, C. dendritica, C. fragilis, C. gracilis, C. carolinensis, C. pinniformis, and C. varians) that we recently described from the tropical Western Pacific (Watling et al., 2011, Xu et al., 2020b). The high species richness (42 of 78 in total) indicates that the Indo-West Pacific convergent region is a possible hotspot of deep-water Chrysogorgia species.
5 CONCLUSIONBased on the morphological and phylogenetic analyses, ten sampled specimens of Chrysogorgia in Caroline seamounts are recognized as two new species Chrysogorgia pinniformis sp. nov. and Chrysogorgia varians sp. nov. Chrysogorgia pinniformis sp. nov. is similar to C. pinnata Cairns, 2007 by the pinnate branching, but differs from the latter being in Versluys' group C, Squamosae typicae group (vs. Versluys' group A, Spiculosae) and has sclerites present in the mouth area of polyps. Chrysogorgia varians sp. nov. belongs to the Chrysogorgia group A (Spiculosae) and shows morphological variability during different growth period. Additionally, it is found that the mtMutS marker showed very limited usefulness for species delimitation of Chrysogorgia, while the 28S rDNA region may a potential barcode for this genus.
6 DATA AVAILABILITY STATEMENTThe specimens described in this study are available at the Marine Biological Museum of Chinese Academy of Sciences (MBMCAS) at Institute of Oceanology, Qingdao, China. Voucher IDs: Chrysogorgia pinniformis sp. nov.: MBM286504; Chrysogorgia varians sp. nov.: MBM286438, MBM286439, MBM286440, MBM286441, MBM286502, MBM286503, MBM286505, MBM286506, MBM286507. The mtMutS sequences that support the findings of this study have been deposited in NCBI GenBank with the accession codes MW085090 (Chrysogorgiapinniformis), MW085091, MW418365, MW418366, MW418367, MW418368, MW418369, MW418370, MW418371, MW418364 (Chrysogorgia varians sp. nov.) and MW418361, MW418362, MW418363 (Chrysogorgia carolinensis Xu, Zhan & Xu, 2020). The 28S rDNA sequences that support the findings of this study have been deposited in NCBI GenBank with the accession codes MW418372 (C. pinniformis sp. nov.), MW418381, MW418377, MW418379, MW418378, MW418380, MW418383, MW418385, MW418382, MW418384 (C. varians sp. nov.) and MW418373, MW418374, MW418375, MW418376 (C. carolinensis).
The new species registration of Chrysogorgia pinniformis sp. nov. and Chrysogorgia varians sp. nov. in Zoobank with LSID: urn: lsid: zoobank. org: act: 74FE8B57-1038-49CC-83E2-CF05BB27C544 and urn: lsid: zoobank.org: act: 96111211-8D7B-40EF-BFF4-1A2D78DE5247, respectively. The publication LSID: urn: lsid: zoobank.org: pub: 5C0EA878-ACF4-453A-9918-F4872AFA73E1.
7 ACKNOWLEDGMENTWe thank assistance of the crew of R/V Kexue (Science in Chinese) and ROV Faxian (Discovery in Chinese) for sample collection. Special thanks go to Mr. Shaoqing WANG for taking the photos onboard.
Electronic supplementary materialSupplementary material (Supplementary Fig.S1) is available in the online version of this article at http://doi.org/10.1007/s00343-021-0386-5.
Alfaro M E, Zoller S, Lutzoni F. 2003. Bayes or Bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Molecular Biology and Evolution, 20(2): 255-266.
DOI:10.1093/molbev/msg028 |
Bayer F M, Grasshoff M, Verseveldt J. 1983. Illustrated Trilingual Glossary of Morphological and Anatomical Terms Applied to Octocorallia. E. J. Brill/Dr. W. Backhuys, Leiden. p. 75.
|
Cairns S D. 2001. Studies on western Atlantic Octocorallia (Coelenterata: Anthozoa) part 1: the genus Chrysogorgia Duchassaing & Michelotti, 1864. Proceedings of the Biological Society of Washington, 114(3): 746-787.
|
Cairns S D. 2007. Calcaxonian Octocorals (Cnidaria: Anthozoa) from the Eastern Pacific seamounts. Proceedings of the California Academy of Sciences, 58(25): 511-541.
|
Cairns S D. 2018. Deep-water octocorals (Cnidaria, Anthozoa) from the Galápagos and Cocos Islands. Part 1: suborder Calcaxonia. ZooKeys, 729: 1-46.
DOI:10.3897/zookeys.729.21779 |
Cordeiro R T S, Castro C B, Pérez C D. 2015. Deep-water octocorals (Cnidaria: Octocorallia) from Brazil: family Chrysogorgiidae Verrill, 1883. Zootaxa, 4058(1): 81-100.
DOI:10.11646/zootaxa.4058.1.4 |
Darriba D, Taboada G L, Doallo R, Posada D. 2012. jModelTest 2: more models, new heuristics and parallel computing. Nature Methods, 9(8): 772.
DOI:10.1038/nmeth.2109 |
Duchassaing P, Michelotti J. 1864. Supplément au mémoire sur les coralliaires des Antilles. Mémoires de l'Académie des Sciences de Turin, 2(23): 97-206.
|
Ehrenberg C G. 1834. Beiträge zur physiologischen Kenntniss der Corallenthiere im allgemeinen, und besonders des rothen Meeres, nebst einem Versuche zur physiologischen Systematik derselben. Abhandlungen der Königlichen Akademie der Wissenschaften zu Berlin, 1: 225-380.
|
Grasshoff M. 1999. The shallow water gorgonians of New Caledonia and adjacent islands (Coelenterata: Octocorallia). Senckenbergiana Biologica, 78: 1-245.
|
Guindon S, Dufayard J F, Lefort V, Anisimova M, Hordijk W, Gascuel O. 2010. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Systematic Biology, 59(3): 307-321.
DOI:10.1093/sysbio/syq010 |
Haeckel E. 1866. Generelle morphologie der Organismen, vol. 2. Verlag von Georg Reimer, Berlin.
|
Hall T A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41: 95-98.
|
Herrera S, Baco A, Sánchez J A. 2010. Molecular systematics of the bubblegum coral genera (Paragorgiidae, Octocorallia) and description of a new deep-sea species. Molecular Phylogenetics and Evolution, 55(1): 123-135.
DOI:10.1016/j.ympev.2009.12.007 |
Hillis D M, Bull J J. 1993. An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Systematic Biology, 42(2): 182-192.
DOI:10.1093/sysbio/42.2.182 |
Katoh K, Standley D M. 2013. MAFFT Multiple Sequence Alignment Software version 7: improvements in performance and usability. Molecular Biology and Evolution, 30(4): 772-780.
DOI:10.1093/molbev/mst010 |
Kinoshita K. 1913. Studien über einige Chrysogorgiiden Japans. Journal of the College of Science, Tokyo Imperial University, 33(2): 47.
|
Kükenthal W. 1919. Gorgonaria. Wissenschaftliche Ergennisse der Deutschen Tiefsee-Expedition auf dem Dampfer "Valdivia" 1898-1899, 13(2): 946.
|
Lamouroux J V F. 1812. Extrait d'un mémoire sur la classification des polypiers coralligènes non entièrement piérreux. Nouvelle Bulletin de la Societé Philomatique, Paris, 3(63): 181-188.
|
McFadden C S, Benayahu Y, Pante E, Thoma J N, Nevarez P A, France S C. 2011. Limitations of mitochondrial gene barcoding in Octocorallia. Molecular Ecology Resources, 11(1): 19-31.
DOI:10.1111/j.1755-0998.2010.02875.x |
McFadden C S, Brown A S, Brayton C, Hunt C B, van Ofwegen L P. 2014. Application of DNA barcoding in biodiversity studies of shallow-water octocorals: molecular proxies agree with morphological estimates of species richness in Palau. Coral Reefs, 33(2): 275-286.
DOI:10.1007/s00338-013-1123-0 |
McFadden C S, van Ofwegen L P. 2013. A second, cryptic species of the soft coral genus Incrustatus (Anthozoa: Octocorallia: Clavulariidae) from Tierra del Fuego, Argentina, revealed by DNA barcoding. Helgoland Marine Research, 67(1): 137-147.
DOI:10.1007/s10152-012-0310-7 |
Nutting C C. 1908. Descriptions of the Alcyonaria collected by the U.S. Bureau of Fisheries steamer Albatross in the vicinity of the Hawaiian Islands in 1902. Proceedings of the United States National Museum, 34: 543-601.
DOI:10.5479/si.00963801.34-1624.543 |
Nutting C C. 1912. Descriptions of the Alcyonaria collected by the U.S. Fisheries steamer "Albatross", mainly in Japanese waters, during 1906. Proceedings of the United States National Museum, 43: 104pp.
|
Pante E, Watling L. 2012. Chrysogorgia from the New England and Corner Seamounts: Atlantic-Pacific connections. Journal of the Marine Biological Association of the United Kingdom, 92(5): 911-927.
DOI:10.1017/S0025315411001354 |
Pérez C D, de Moura Neves B, Cordeiro R T, Williams G C, Cairns S D. 2016. Diversity and distribution of Octocorallia. In: Goffredo S, Dubinsky Z eds. The Cnidaria, Past, Present and Future: the World of Medusa and Her Sisters. Springer, Cham. p. 109–123. https://doi.org/10.1007/978-3-319-31305-4_8.
|
Rambaut A, Drummond A J. 2007. Tracer v1.4. Available from http://beast.bio.ed.ac.uk/Tracer.
|
Ronquist F, Huelsenbeck J P. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19(12): 1572-1574.
DOI:10.1093/bioinformatics/btg180 |
Sánchez J A, Lasker H R, Taylor D J. 2003. Phylogenetic analyses among octocorals (Cnidaria): mitochondrial and nuclear DNA sequences (lsu-rRNA, 16S and ssu-rRNA, 18S) support two convergent clades of branching gorgonians. Molecular Phylogenetics and Evolution, 29(1): 31-42.
DOI:10.1016/S1055-7903(03)00090-3 |
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30(12): 2725-2729.
DOI:10.1093/molbev/mst197 |
Untiedt C B, Quattrini A M, McFadden C S, Alderslade P A, Pante E, Burridge C P. 2021. Phylogenetic relationships within Chrysogorgia (Alcyonacea: Octocorallia), a morphologically diverse genus of octocoral, revealed using a target enrichment approach. Frontiers in Marine Science, 7: 599984.
DOI:10.3389/fmars.2020.599984 |
Utinomi H. 1961. Noteworthy Octocorals collected off the southwest coast of Kii Peninsula, middle Japan. Part 2, Telestacea, Gorgonacea and Pennatulacea. Publications of the Seto Marine Biological Laboratory, 9(1): 197-228.
DOI:10.5134/174656 |
Verrill A E. 1883. Report on the Anthozoa, and on some additional species dredged by the "Blake" in 1877-1879, and by the U.S. Fish Commission steamer "Fish 126 Hawk" in 1880-82. Bulletin of the Museum of Comparative Zoology, 11: 1-72.
|
Versluys J. 1902. Die gorgoniden der siboga-expedition. 1. Die chrysogorgiiden. Siboga Expeditie, 13: 1-120.
|
Watling L, France S C, Pante E, Simpson A. 2011. Chapter two-biology of deep-water octocorals. Advances in Marine Biology, 60: 41-122.
DOI:10.1016/B978-0-12-385529-9.00002-0 |
Wright E P, Studer T. 1889. Report on the Alcyonaria collected by H.M.S. Challenger during the years 1873-76. Report on the Scientific Results of the Voyage of H.M.S. Challenger during the years 1873-76, Zoology, 31(64): 1-314.
|
Xu Y, Li Y, Zhan Z F, Xu K D. 2019. Morphology and phylogenetic analysis of two new deep-sea species of Chrysogorgia (Cnidaria, Octocorallia, Chrysogorgiidae) from Kocebu Guyot (Magellan seamounts) in the Pacific Ocean. Zookeys, 881: 91-107.
DOI:10.3897/zookeys.881.34759 |
Xu Y, Zhan Z F, Xu K D. 2020a. Morphology and molecular phylogeny of three new deep-sea species of Chrysogorgia (Cnidaria, Octocorallia) from seamounts in the tropical Western Pacific Ocean. Peer J, 8: e8832.
DOI:10.7717/peerj.8832 |
Xu Y, Zhan Z F, Xu K D. 2020b. Morphology and phylogenetic analysis of five deep-sea golden gorgonians (Cnidaria, Octocorallia, Chrysogorgiidae) in the Western Pacific Ocean, with the description of a new species. ZooKeys, 989: 1-37.
DOI:10.3897/zookeys.989.53104 |
 2021, Vol. 39
2021, Vol. 39