Institute of Oceanology, Chinese Academy of Sciences
Article Information
- LÜ Ting, ZHAN Zifeng, XU Kuidong
- Morphology and molecular phylogeny of three black corals (Antipatharia, Schizopathidae) from seamounts in the Western Pacific Ocean, with description of a new species
- Journal of Oceanology and Limnology, 39(5): 1740-1757
- http://dx.doi.org/10.1007/s00343-021-0455-9
Article History
- Received Nov. 24, 2020
- accepted in principle Feb. 3, 2021
- accepted for publication May. 11, 2021
2 Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology (Qingdao), Qingdao 266237, China;
3 University of Chinese Academy of Sciences, Beijing 100049, China
Members of the black coral order Antipatharia are widely distributed in oceans, ranging from shallow waters to deep seas, and more than 75% species occur in depths over 50 m (Cairns, 2007; Wagner et al., 2012). This order belongs to the anthozoan subclass Hexacorallia and encompasses 7 families, 46 genera, and 273 species (Opresko et al., 2020; Terrana et al., 2020). Within Antipatharia, members of the family Schizopathidae are mainly distributed in deep seas (Molodtsova and Opresko, 2017). The family consists of two subfamilies and 13 genera, with varying degrees of pinnulation (Opresko, 2002, 2005; MacIsaac et al., 2013; Molodtsova and Opresko, 2017).
To explore the biodiversity on seamounts in the tropical Western Pacific Ocean, we conducted a cruise in the region the Caroline Ridge in May 2019. By using the remotely operating vehicle (ROV), we obtained ten black coral specimens of the family Schizopathidae. Based on the morphological analyses, three specimens are identified as Umbellapathes parva sp. nov., four specimens are Telopathes cf. magna, and three specimens are Stauropathes cf. punctata. Meanwhile, their genetic distances and phylogenetic positions are evaluated using the region of the nuclear internal transcribed spacer (ITS1 and ITS2) and two mitochondrial fragments cox3-IGR-cox1 (COI) and trnW-IGR-nad2 (NAD2).
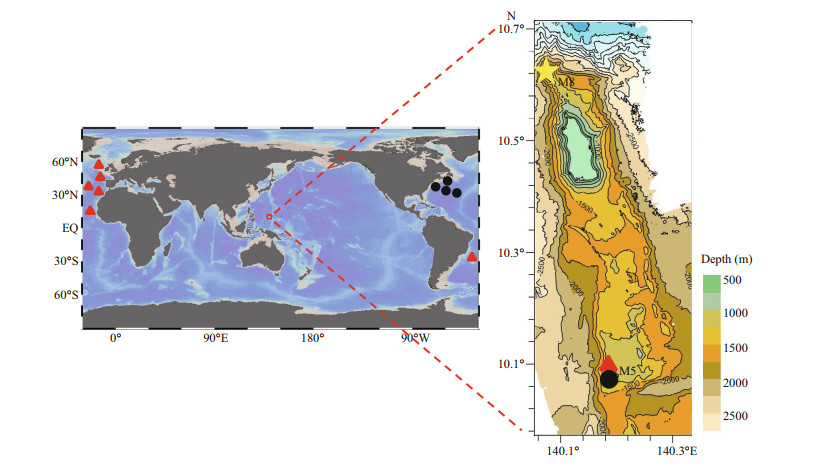
2 MATERIAL AND METHOD 2.1 Specimen collection and morphological examinationDuring the 2019 R/V Kexue (Science in Chinese) cruise, samples from two seamounts (tentatively named as Caroline Seamounts M5 and M8) on the Caroline Ridge in the tropical Western Pacific were obtained by the ROV Faxian (Discovery in Chinese) (Fig. 1). Before sampling, the specimens were photographed in situ. After collection, samples were photographed on the deck. The collected specimens were stored in 85% ethanol, and some branches of each specimen were stored at -80 ℃ for molecular research. All the sampled specimens were deposited in the Marine Biological Museum of Chinese Academy of Sciences (MBMCAS) at Institute of Oceanology, Chinese Academy of Sciences (IOCAS), Qingdao, China (Table 1).
|
| Fig.1 Geographic distribution of Umbellapathes parva sp. nov. (yellow star), Telopathes magna MacIsaac & Best, 2013 (black dots), and Stauropathes punctata (Roule, 1905) (red triangles) Red rectangle indicates the right figure, showing the sampling sites of the three species from the seamounts M5 and M8 on the Caroline Ridge in the tropical Western Pacific. |
Several fragments were taken from distal pinnules on upper part of corallum of each specimen for scanning electron microscopy (Horowitz et al., 2020). The larger tissues are carefully removed with forceps. The remaining smaller tissues were cleaned with commercial bleach for 1-2 min, and then transferred to 90% alcohol for 1 h (Wagner and Opresko, 2015). To study the microscopic skeletal structure of spines, they were air-dried, fixed to carbon double adhesive tape, coated with golden and observed with scanning electron microscope. All measurements were taken on the preserved material, following Wagner et al. (2010). Terminology follows Opresko et al. (2014).
2.2 DNA extraction and sequencingThe nuclear rDNA internal transcribed spacer (ITS) region spanning partial 18S rDNA, ITS1, 5.8S, ITS2, and partial 28S rDNA has been proved to successfully evaluate the phylogenetic relationships and establish new taxa for black coral species (Lapian et al., 2007; Bo et al., 2009, 2018; Wagner et al., 2010). Meanwhile, two frequently used markers, mitochondrial cox3-IGR-cox1 (COI) and trnW-IGR-nad2 (NAD2) were also selected for the phylogenetic analyses.
Genomic DNA was extracted from each specimen using the DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany) following the instructions. PCR amplifications for the ITS, COI, and NAD2 regions were conducted using primer pairs RA2 (5'-GTCCC-TGCCCTTTGTACACA-3') and ITS 2.2 (5'-CCTGG-TTAGTTTCTTTTCCTCCGC-3'; Wörheide, 1998), CO3 gen3360F (5'-CTTTGTGGCAACTGGGTTT-CATG-3') and CO1 gen4446R (5'-GATAACATTG-CATAAACCATCCCT-3'; Thoma et al., 2009), and TRPantiF (5'-GGAAGACCGTTAGCCTTC-3') and NAD2anti1040R (5'-CCAAATAAGAATAAGCCT-GAAG-3'; Thoma et al., 2009), respectively. PCR reactions were performed using I-5TM 2× High-Fidelity Master Mix DNA polymerase (TsingKe Biotech, Beijing, China). The amplification cycle conditions were as follow: denaturation at 98 ℃ for 2 min, followed by 32 cycles of denaturation at 98 ℃ for 20 s, annealing at 51-55 ℃ for 20 s, extension at 72 ℃ for 15 s, and a final extension step at 72 ℃ for 2 min. PCR purification and sequencing were performed by TsingKe Biological Technology (TsingKe Biotech, Beijing, China).
2.3 Genetic distance and phylogenetic analysesThe ITS, COI, and NAD2 sequences of Schizopathidae spp. and two Cladopathidae species as an out group were downloaded from GenBank (Table 1). The nucleotide sequences were aligned using MAFFT v.7 (Katoh and Standley, 2013) with the G-INS-i algorithm. The nucleotide alignment was trimmed to equal length using BioEdit v7.0.5 (Hall, 1999). Genetic distances of ITS1-5.8S-ITS2 between species/populations were calculated with MEGA 6.0 using Kimura 2-parameter model (Tamura et al., 2013).
Phylogenetic analyses were performed on the ITS, mitochondrial regions (COI-NAD2), and concatenated ITS, COI, and NAD2 (ITS-COI-NAD2) datasets. When conspecific sequences were identical, only one was chosen for analysis. The taxa included in the COI-NAD2 trees are fewer than the ITS and ITS-COI-NAD2 ones (20 vs. 26 for both the two latter datasets; Table 1). The best-fit models, GTR+G for COI-NAD2, and GTR+I+G for ITS and ITS-COI-NAD2 were selected by SMS with the Akaike information criterion in PhyML-3.0 (see http://www.atgc-montpellier.fr/phyml/; Lefort et al., 2017) on the online ATGC bioinformatic platform. Maximum likelihood (ML) analysis was carried out using PhyML-3.0 (Guindon et al., 2010), with node support from a majority-rule consensus tree of 1 000 bootstrap replicates. Following Hillis and Bull (1993), the ML bootstraps < 70%, 70%-94%, and ≥95% were considered as low, moderate, and high, respectively. Bayesian inference (BI) analysis was performed using MrBayes v3.2.7a (Ronquist and Huelsenbeck, 2003). Posterior probability was estimated based on 10 000 000 Monte Carlo Markov Chain (MCMC) generations (×4 chains) sampling every 1 000 generations (burn-in=25%). Convergence of the MCMC was assessed using Tracer 1.4.1 (Drummond and Rambaut, 2007). Following Alfaro et al. (2003), the Bayesian posterior probabilities < 0.95 and ≥0.95 were considered as low and high, respectively.
3 RESULT 3.1 TaxonomyClass Anthozoa Ehrenberg, 1831
Order Antipatharia Milne-Edwards & Haime, 1857
Family Schizopathidae Brook, 1889
Subfamily Parantipathinae Roule, 1905
Genus Umbellapathes Opresko, 2005
Type species: Umbellapathes helioanthes Opresko, 2005
Umbellapathes parva sp. nov. (Figs. 2 & 3; Table 2)
Material examined Holotype: MBM286412, station FX-Dive 226 (10°38'08"N, 140°04'08"E), M8 seamount, depth 1 766 m, 14 June 2019. Paratype: MBM286410, station FX-Dive 224 (10°37'36"N, 140°05'25"E), M8 seamount, depth 1 488 m, 12 June 2019; MBM286411, station FX-Dive 224 (10°37'38"N, 140°05'27"E), M8 seamount, depth 1 509 m, 12 June 2019.

|
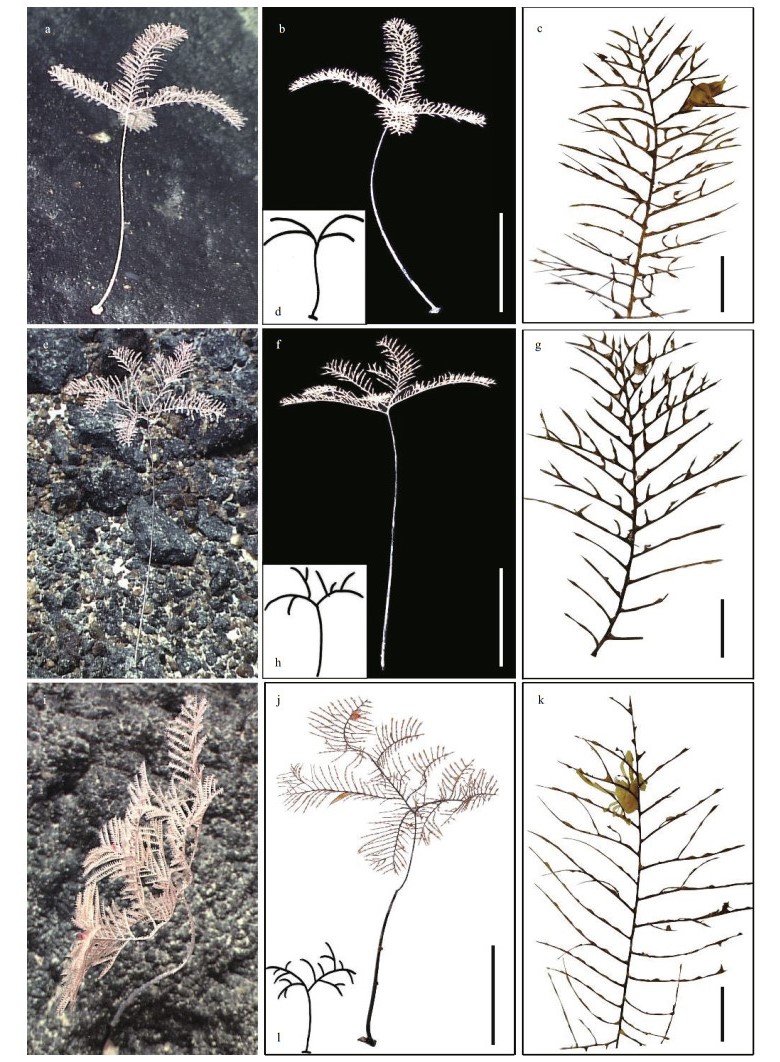
| Fig.2 The external morphology of Umbellapathes parva sp. nov. The holotype specimen MBM286412 (i-l) and paratype specimens MBM286410 (a-d) and MBM286411 (e-h) in situ (a, e, i) and after collection (b, c, f, g, j, k), showing the arrangement of pinnulated branches (d, h, l) and pinnules (c, g, k). Scale bars: 10 cm (b, f, j); 1 cm (c, g, k). |
|
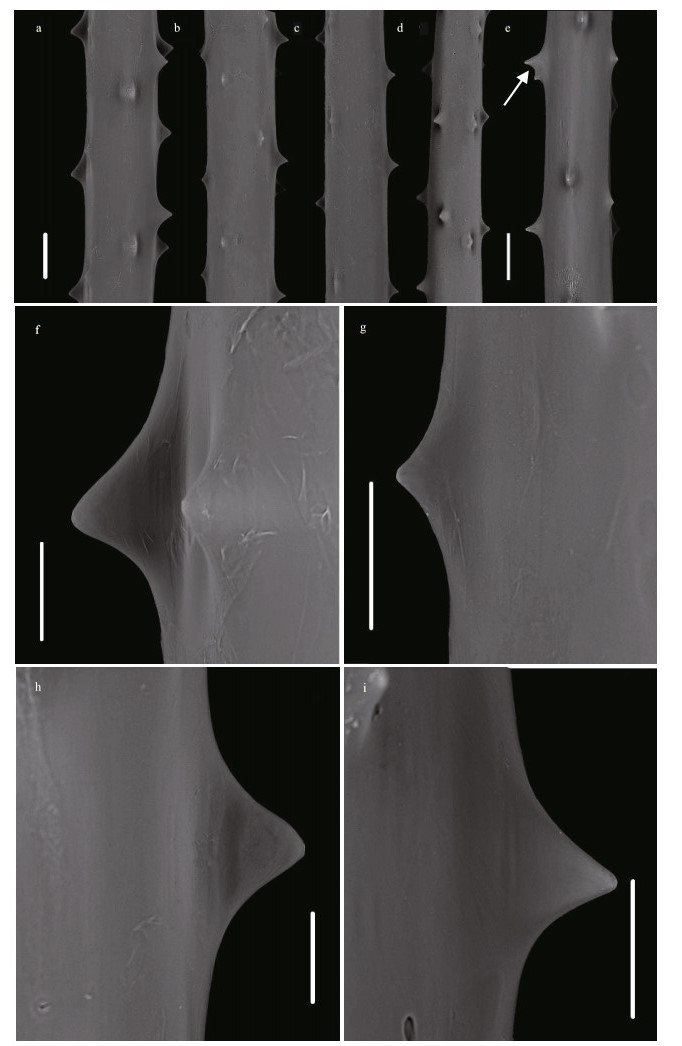
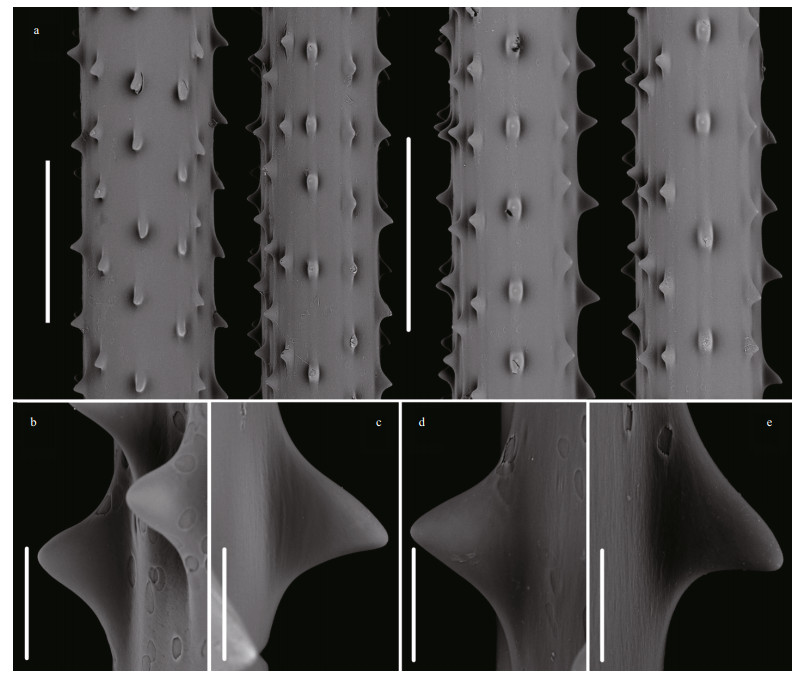
| Fig.3 Spines of the holotype of Umbellapathes parva sp. nov. under Scanning Electron Microscope (SEM) a-b. spines on the middle part of primary pinnules; c-d. spines on the middle part of secondary pinnules; e. a forked spine (arrow); f. a polypar spine with the measured maximum height of 0.050 mm; g. an Abpolypar spine with the measured minimum height of 0.017 mm; h. an abpolypar spine with the height of 0.044 mm; i. a polypar spine with the height of 0.041 mm. Scale bars: 0.1 mm (a-e); 0.05 mm (f-i). |
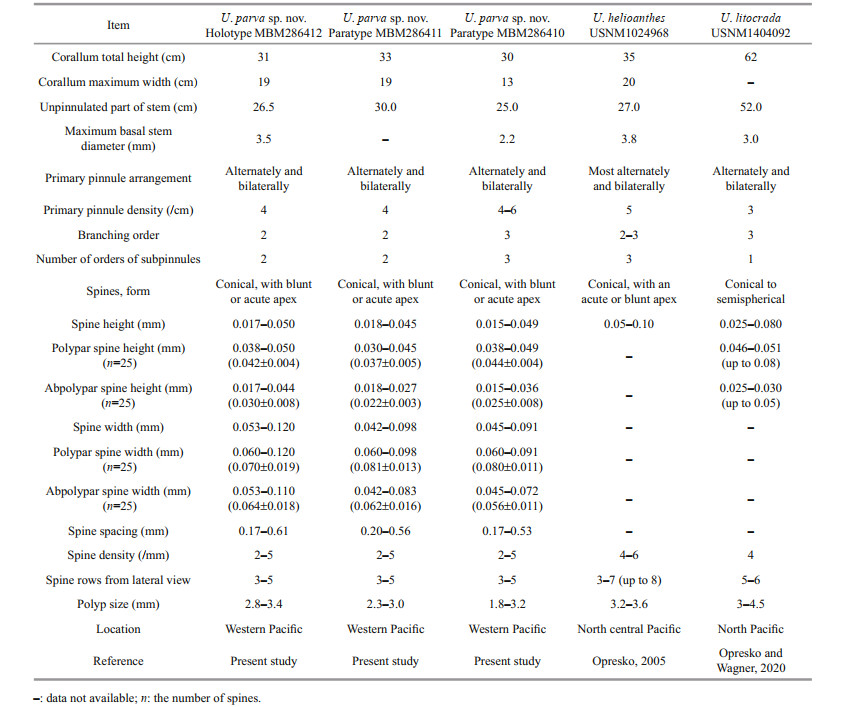
Diagnosis Corallum monopodial and pinnulate, up to three orders of branches arising from the primary pinnules of stem. Unpinnulated portion of stem long and slightly curved. Primary pinnules on branches and stem arranged in two lateral or anterolateral rows and in alternating pattern. Secondary pinnules arranged uniserially on front side of primary pinnules. Spines smooth, conical, rarely forked, with blunt or acute apex. Polypar spines 0.038-0.050-mm high and abpolypar spines 0.017-0.044-mm high. Spines in 3-5 rows from lateral view, with 2-5 spines per millimeter in each row. Polyps 1.8-3.4 mm in transverse diameter.
Description of holotype Colony was slightly pink in situ and became yellowish-brown after being preserved in alcohol. The corallum is about 31-cm high and 19-cm wide, with the basal attachment plate about 1.5 cm×2 cm (Fig. 2i; Table 2). The unpinnulated stalk is about 26.5-cm long, 3.5 mm×1.8 mm in diameter near the base, and 1 mm in diameter just below the first branch (Fig. 2j). There are up to three orders of branches arising from the primary pinnules of stem. The primary pinnules on branch and stem arranged in two lateral or anterolateral rows and in alternating pattern. The stem pinnules develop into pinnulated branches and form discoidal crown (Fig. 2l). The lengths of branches and pinnulated branches range from 5 cm to 17 cm. The stem is slightly curved and the pinnulated portion of stem grows upwards and then bent backwards.
Primary pinnules on larger branches and pinnulated branches are arranged biserially and alternately (Fig. 2k). They are spaced 5-7 mm on the same side of the axis, about 4 pinnules per centimeter for both sides. The length of primary pinnules ranges from 0.7 cm to 4 cm. The distal angle between primary pinnules and pinnulated branches ranges from 60° to 70°. The internal angle between the two anterolateral rows of primary pinnules is in the range of 110°-130°.
The subpinnules are arranged uniserially and inclined to the distal ends. There are 1-2 secondary pinnules on each primary pinnules, 0.5-1.5 cm in length and spaced 5-8-mm apart along the pinnules (Fig. 2k). The distal angle formed by the secondary pinnules and the primary pinnules ranges from 30° to 60°.
Spines on pinnules and branches are simple, smooth, rarely forked, with blunt or acute apex, subequal in size along the axis on the same side (Fig. 3; Table 2). The polypar spines on the middle part of the pinnules are 0.038-0.050-mm (0.042 mm 0.004 mm, n=25) high and 0.060-0.12-mm (0.070 mm 0.019 mm, n=25) wide at the base, and the abpolypar spines are much smaller, 0.017-0.044-mm (0.030 mm 0.008 mm, n=25) high and 0.053-0.11-mm (0.064 mm 0.018 mm, n=25) wide at the base (Fig. 3). On the basal part of the pinnules, the polypar spines are 0.036-0.045-mm tall and the abpolypar spines are mostly 0.018-0.027-mm tall. Along the distal parts of the pinnules, the polypar spines are 0.03-0.045-mm tall, and the abpolypar spines are 0.023-0.038-mm tall. Spines are arranged in 3-5 longitudinal rows from the lateral view. Larger polypar spines are usually in 4 rows and smaller abpolypar spines in 3 rows, when viewed from transection. The distance between adjacent spines in each row is 0.17-0.61 mm, about 2-5 spines per millimeter in each row.
Polyps were not well preserved in ethanol-fixed specimens, about 2.8-3.4 mm in transverse diameter. The polyps in situ were arranged in a single row along each pinnule.
Paratype The two paratype specimens showed similar morphology as the holotype (Fig. 2a-g). However, there are some minor differences. In the paratype MBM286410 and MBM286411, the arrangement of primary pinnules is closer and the degrees of pinnulation is relatively simple than in the holotype. Instead of forming discoidal crown, there are several elongated pinnulated branches growing upwards and then backwards in the paratype specimens. Additionally, the number of subpinnules in the holotype is lower than that in the paratype specimens and tertiary pinnules only occur in the paratype MBM286410 (Fig. 2c; Table 2). These differences are minor and not constant.
Type locality A seamount (tentatively named as M8) located on the Caroline Ridge in the Western Pacific Ocean (10°38'08"N, 140°04'08"E) with depth of 1 766 m.
Etymology The Latin adjective parvus (small) refers to the small spines of the species.
Distribution and habitat Found only from the M8 seamount with depth of 1 488-1 766 m in the tropical Western Pacific Ocean (Fig. 1), where the species attached on rocky substrate.
Remarks Considering the pinnulation degree and basal stem diameter, the three specimens of U. parva sp. nov. showed different growth stages, from simple pinnulation colony (younger colony) to complex pinnulation colony (adult). The specimen MBM286410 is likely a younger colony with 4 pinnulated branches and narrow basal stem (Fig. 2b & d), the specimen MBM286411 is likely in the intermediate growth stage with 8 pinnulated branches and narrow basal stem (Fig. 2f & h), and the specimen MBM286412 is possible an adult with 16 pinnulated branches and wider basal stem (Fig. 2j & l). The spines are composed of two types, and the polypar spines are much larger than the abpolypar spines. However, both the size of the polypar spines and that of the abpolypar spines showed little variation within and among the three specimens (Table 2), and thus they can be served as differing features.
Umbellapathes parva sp. nov. belongs to the genus Umbellapathes characterized by a long unpinnulated stem and pinnulated branches developing from primary pinnules. Umbellapathes contains only two species, U. helioanthes, Opresko, 2005 and U. litocrada Opresko & Wagner, 2020. Both of them were discovered from North Pacific (Opresko, 2005; Opresko et al., 2020). The new species is most similar to Umbellapathes helioanthes Opresko, 2005, particularly in the arrangement of subpinnules and size of polyps (Opresko, 2005). Opresko (2005) did not give separate measurements of the polypar and abpolypar spines. Nonetheless, U. parva sp. nov. can be distinguished from U. helioanthes by its much smaller spines (0.017-0.044-mm high in abpolypar spines and 0.038-0.050-mm high in polypar spines vs. 0.05-0.1-mm high; Table 2). U. parva sp. nov. is easily distinguished from U. litocrada Opresko & Wagner, 2020 by the present of secondary pinnules (present vs. absent), the shape of spines (conical vs. conical to semispherical; Table 2).
Subfamily Schizopathinae Brook, 1889
Genus Telopathes MacIsaac & Best, 2013
Type species: Telopathes magna MacIsaac & Best, 2013
Telopathes cf. magna MacIsaac & Best, 2013 (Figs. 4 & 5; Table 3)
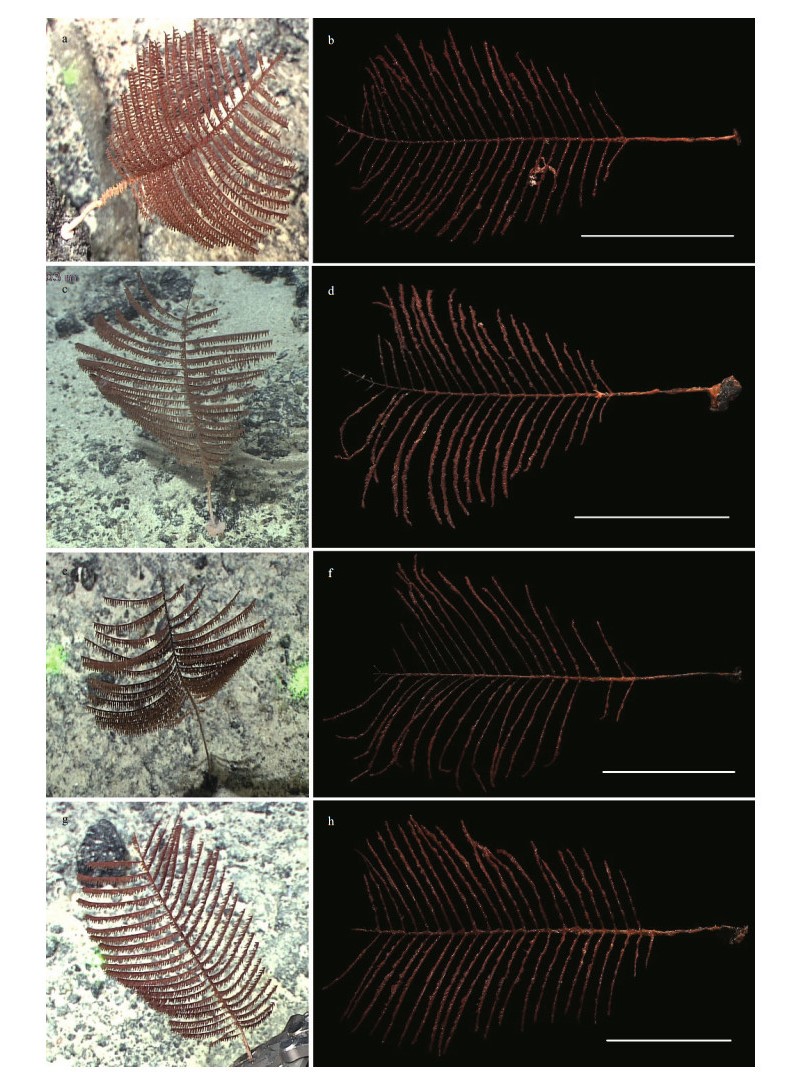

|
| Fig.4 The external morphology of Telopathes cf. magna MacIsaac & Best, 2013 The specimens MBM286483 (a, b), MBM286482 (c, d), MBM286481 (e, f), and MBM286480 (g, h) in situ (a, c, e, g) and after collection (b, d, f, h). Scale bars: 10 cm. |

|
| Fig.5 Spines of Telopathes cf. magna MacIsaac & Best, 2013 (MBM286483) under SEM a. spines on the pinnules; b, d. abpolypar spines; c, e. polypar spines. Scale bars: 0.5 mm (a); 0.05 mm (b-e). |
Telopathes magna MacIsaac & Best, 2013: 237- 258; figs. 2-7.
Material examined MBM286483, station FX-Dive 209 (10°04'40"N, 140°12'13"E), M5 seamount, depth 1 016 m, 27 May, 2019; MBM286482, station FX-Dive 210 (10°04'39"N, 140°12'07"E), M5 seamount, depth 937 m, 28 May, 2019; MBM286481, station FX-Dive 210 (10°04'39"N, 140°12'07"E), M5 seamount, depth 945 m, 28 May, 2019; MBM286480, station FX-Dive 210 (10°04'39"N, 140°12'07"E), M5 seamount, depth 944 m, 28 May, 2019.
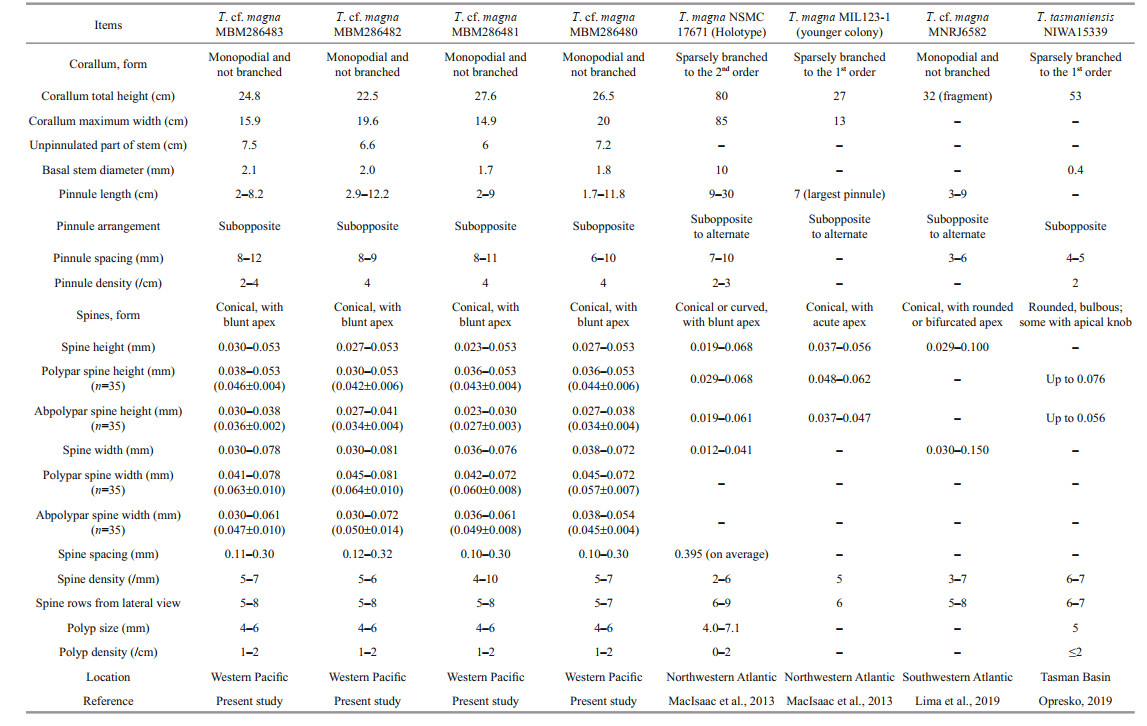
Description Colony was brownish in situ and became dark brown after being preserved in alcohol. The monopodial corallum is about 24.8-cm high and 15.9-cm wide, and the unpinnulated section of lowermost stem is about 7.5-cm long (Fig. 4a-b; Table 3). The basal attachment plate is 0.6 cm across, and the stem is about 2.1 mm in diameter near the basal attachment plate and 0.4 mm in diameter near the top of stem. The colony consists of 24 pairs of bilateral and subopposite pinnules. The length of pinnules ranges from 2 cm to 8.2 cm. Pinnules increase in length from the lowermost pairs of pinnules, reach maximum length at about the mid-pair, then decrease in length to the shortest at the top of colony. Pinnule pairs are spaced 8-12-mm apart (mostly 8 mm), with distance decreasing from the lowest pairs toward the apex of colony. In each pair, the pinnule on the right side (viewing the front of the corallum) is 1-mm higher than the pinnule on the left. The distal angle between the stem and pinnules ranges from 60° to 70°. The interior angle between subopposite pairs is in the range of 100°-150°.
Spines on pinnules are simple, smooth, conical and nearly equal-sized on the same side of axis (Fig. 5a-e). The polypar spines are 0.038-0.053-mm (0.046 mm 0.004 mm, n=35) high and 0.041-0.078-mm (0.063 mm 0.010 mm, n=35) wide at the base, and the abpolypar spines are 0.030-0.038-mm(0.036 mm 0.002 mm, n=35)-mm high and 0.030-0.061-mm (0.047 mm 0.010 mm, n=35)-mm wide at the base. Spines are arranged in 5-8 rows from the lateral view. Larger polypar spines are usually in 6 rows and smaller abpolypar spines in 4 rows, when viewed from transection. Spines are spaced 0.11-0.30-mm apart, about 5-7 spines per mm in each row. Polyps are not well preserved in ethanol-fixed specimens, about 4-6 mm in transverse diameter (Table 3).
Distribution and habitat The species was first found from the Northwestern Atlantic, i.e., on the continental slope off Nova Scotia, New England Seamounts, and Corner Rise Seamounts with depth of 1 073-1 983 m (MacIsaac et al., 2013; Fig. 1). In the tropical Western Pacific, it was found on rocky substrate of the M5 seamount with depth of 937-1 016 m. The in-situ water temperatures were 5.0-5.1 ℃ and the salinity approximately 36.0-36.7.
Remarks The four Pacific specimens match with the holotype of T. magna in having the relatively long, simple, bilateral pinnules and simple, conical spines and large polyps. The main difference is that our specimens have a smaller colony size and lack of branching (MacIsaac et al., 2013). Other minor differences include the arrangement of pinnules (subopposite to alternate vs. only subopposite) and the color of living colony (brownish vs. bright orange), and the spine density is slightly more crowded in the specimen MBM286481 than that in the holotype (4-10 vs. 2-6 per millimeter). However, MacIsaac et al. (2013) reported that the small/young colonies of T. magna might lack branching. The overall corallum morphology of the four specimens is more similar to that of the juvenile colony (see Fig. 6b in MacIsaac et al. (2013)) rather than the adult holotype of T. magna. Furthermore, there is no or only one-nucleotide variation of ITS-5.8S-ITS2 between the present specimens and the previously published ones (see the molecular result), and there is no genetic variation of NAD2 between them. The four specimens likely represent certain juvenile/young-colony stages of T. magna. Nevertheless, considering that all the reported samples were discovered in the Northwestern Atlantic, we identified the four Pacific specimens as T. cf. magna. More specimens, especially the adult ones from the Pacific are needed to confirm their conspecific assignment or reject it.
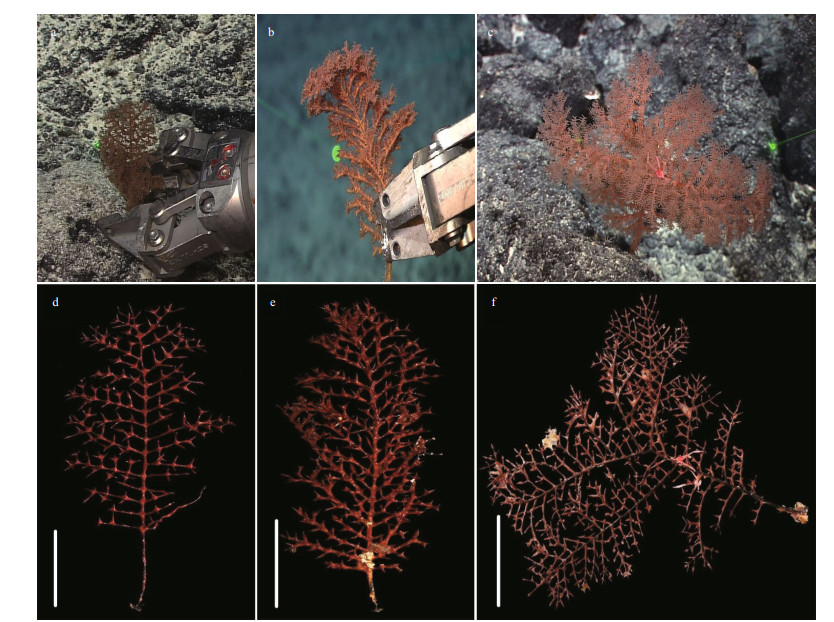

|
| Fig.6 The external morphology of Stauropathes cf. punctata (Roule, 1905) The specimens MBM286474 (a, d), MBM286475 (b, e), and MBM286476 (c, f) in situ (a-c) and after collection (d-f). MBM286474 and MBM286475 are younger colonies and MBM286476 is an adult colony. Scale bars: 5 cm (d, e); 10 cm (f). |
Subfamily Schizopathinae Roule, 1905
Genus Stauropathes Opresko, 2002
Type species: Stauropathes staurocrada Opresko, 2002
Stauropathes cf. punctata (Roule, 1905) (Figs. 6, 7; Table 4)
|
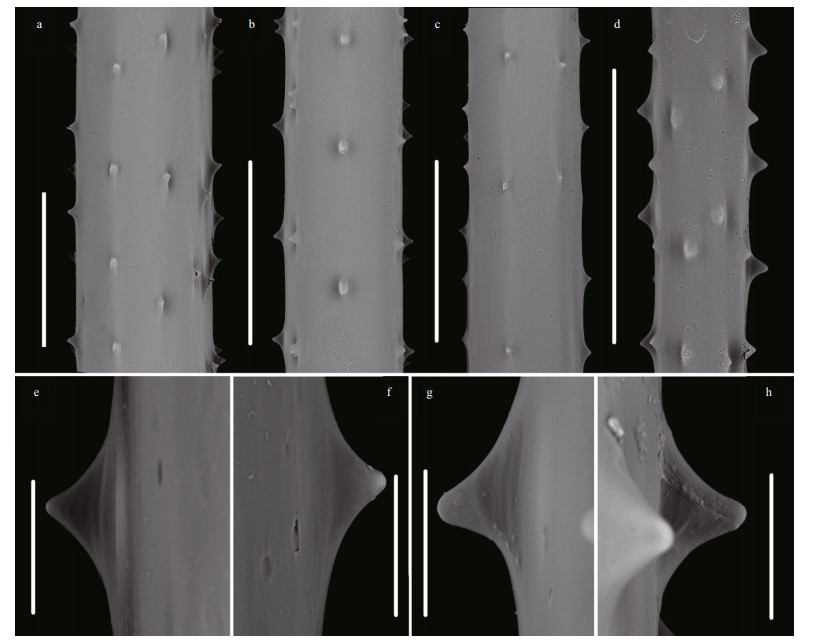
| Fig.7 Spines on the pinnules of Stauropathes cf. punctata (Roule, 1905) under SEM a, b. spines of MBM286476; c, d. spines of MBM286474; e, h. polypar spines; f, g. abpolypar spines. Scale bars. 0.5 mm (a-d); 0.05 mm (e-h). |
(?) Tylopathes hirta-Roule 1902: 81-82; pl.4: 1.
Tylopathes? punctata Roule, 1905: 26, 28, 35, 81, 93, 95. Fig. 1; Opresko, 2002: 417, 420.
Antipathes punctata Silberfeld 1909: 8; Land & Opresko, 2001: 109; Grasshoff, 1985b: 303, 306; Tyler & Zibrowius, 1992: 217.
Stauropathes punctata Opresko, 2002: 417, 420; Molodtsova, 2006: 148; Molodtsova, 2014: 6; Lima et al., 2019: 41.
Material examined MBM286474, station FX-Dive 209 (10°04'42"N, 140°12'11"E), M5 seamount, depth 1 087 m, 27 May, 2019; MBM286475, station FX-Dive 210 (10°04'39"N, 140°12'07"E), M5 seamount, depth 942 m, 28 May, 2019; MBM286476, station FX-Dive 212 (10°03'19"N, 140°10'45"E), M5 seamount, depth 1 353 m, 30 May, 2019.
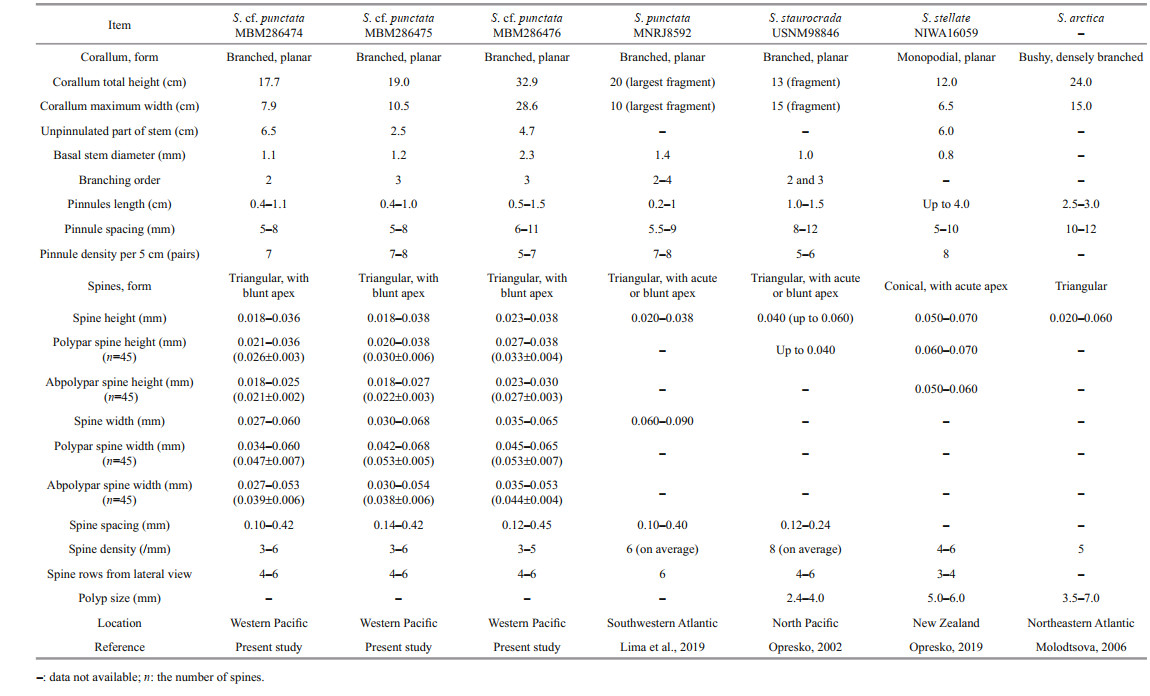
Description Colony reddish brown in situ and became brown after being preserved in alcohol. The corallum is branched to third order and generally in a single plane, about 32.9-cm high and 28.6-cm wide (Fig. 6c & f; Table 4). The stem extends to the top of corallum and the unpinnulated section of stem is about 4.7-cm long. The basal attachment plate is about 1.7 cm in diameter and the stem about 2.3 mm near the basal attachment plate. The pinnulated part of stem consists of 12 pairs of subopposite pinnules and branches, which are 2-19-cm long and 8-12-mm spaced along the stem. The pinnules are 0.5-1.5-cm long and 6-11-mm spaced on the branches. The pinnules spacing generally decreases from the lowest pairs towards the apex of branches. Distal angle formed by stem and pinnules is in the range of 65°-90°. Internal angle between the subopposite pairs ranges from 100° to 140°.
Spines on pinnules are simple, smooth, conical, and nearly equal-sized on the same side of axis (Fig. 7). The polypar spines are 0.027-0.038-mm (0.033 mm 0.004 mm, n=45) in height and 0.045-0.065-mm (0.053 mm 0.007 mm, n=45) in width, and the abpolypar spines are 0.023-0.030-mm (0.027 mm 0.003 mm, n=45) in height and 0.035-0.053-mm (0.044 mm 0.04 mm, n=45) in width. Spines are arranged in 4-6 rows from the lateral view (Fig. 7). Polypar spines are usually in 6 rows and abpolypar spines in 4 rows, when viewed from transection. There are 3-5 spines per mm in each row, where they are 0.12-0.45-mm spaced. Polyps are not well preserved in ethanol-fixed specimens.
Distribution and habitat The species Stauropathes punctata was found from Santa Cruz das Flores Island, the Azores, Portugal, and Cabo Verde Archipelago in the Northeastern Atlantic with depth of 1 300 m (Roule, 1905; Molodtsova, 2006) and from the Rio Grande Rise in the Southwestern Atlantic with depth of 1 600 m (Lima et al., 2019; Fig. 1). In the tropical Western Pacific, the species occurred on the rocky substrate of the M5 seamount, where the depths were 942-1 353 m, water temperatures about 3.9-5.0 ℃ and salinity approximately 36.5-36.7.
Remarks To date, there are four species in the genus Stauropathes: S. arctica (Lütken, 1872), S. punctata (Roule, 1905), S. staurocrada Opresko, 2002, and S. stellate Opresko, 2019. The three specimens from the Caroline seamounts are highly similar to the description of S. punctata by Lima et al. (2019) as well as the incomplete original description by Roule (1905). All of them have monopodial and planar corallum with two rows pinnules arranged in subopposite pairs. Moreover, the ITS, COI, and NAD2 sequences of the three specimens are identical. Considering all the records of the species were discovered from Atlantic, we identified the three Pacific specimens as S. cf. punctata.
Opresko (2002) separated S. punctata from S. staurocrada solely by the pinnule spacing (6-8 mm vs. 8-12 mm). Considering the corallum size, branching degree and basal stem diameter, the specimen MBM286476 with large and complex branching colony is likely an adult colony, while the specimens MBM286474 and MBM286475 with small and sparsely branched colonies are younger colonies (Table 3). The spine size is nearly identical between the younger colony and adult colony and can be used for species identification. By contrast, the pinnule spacing is highly variable (5-8-mm apart in the younger colony vs. 6-11-mm apart in the adult colony), and thus we consider it as conspecific variation. The data show that the ranges of the pinnule spacing are highly variable in conspecific specimens and overlap between S. cf. punctata and S. staurocrada (5-11 mm vs. 8-12 mm), indicating that pinnule spacing may be not a good feature to separate Stauropathes species. In addition, Stauropathes cf. punctata differs from S. arctica and S. stellate by the corallum mainly in a single plane and the smaller spines (Table 4).
3.2 Genetic distance and phylogenetic analysesThe ITS, COI, and NAD2 sequences of the three species were deposited in GenBank (Table 1). The alignment datasets of ITS1-5.8S-ITS2, ITS (comprising partial 18S and 28S rDNA), COI-NAD2, and ITS-COI-NAD2 comprised 653, 796, 1 379, and 2 175 nucleotide positions, respectively. The intraspecific distances of ITS1-5.8S-ITS2 were calculated based on Telopathes magna, Telopathes cf. magna, and Umbellapathesparva sp. nov. populations/ specimens, and the distances were in range of 0−0.17% (Supplementary Table S1). The distances between Telopathes magna and Telopathes cf. magna were in range of 0−0.17%, and those between Umbellapathes parva sp. nov. and the closely related species Alternatipathes bipinnata were in range of 1.72%−1.89% (Supplementary Table S1).
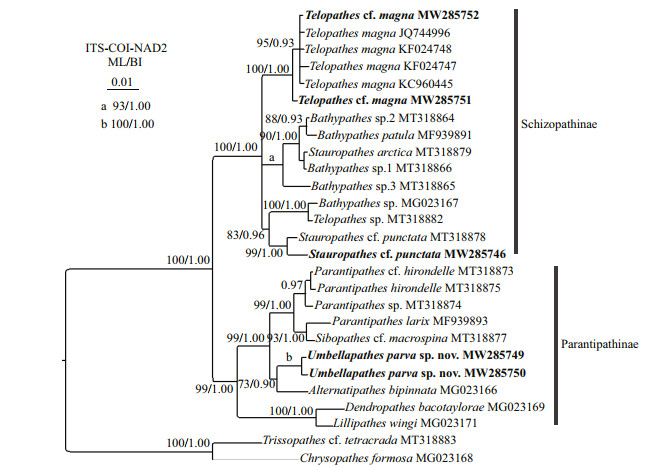
The phylogenetic reconstruction of ITS showed nearly identical results as that of the concatenated ITS-COI-NAD2, but with lower resolution, and the COI-NAD2 trees (Supplementary Fig.S1) were also highly similar to those of ITS-COI-NAD2. Hence, only the ITS-COI-NAD2 results are showed here. The ML and BI analyses yielded an identical topology, and thus only the BI tree with both support values are shown (Fig. 8). All the Schizopathidae species were separated into two main clades corresponding well to the two subfamilies Schizopathinae and Parantipathinae with full node support. The species of Schizopathinae were separated into three sister subclades: (1) all the Telopathes magna populations formed a monophyletic subclade with full node support; (2) Stauropathes punctata, S. cf. punctata, Bathypathes sp. MG023167, and Telopathes sp. MT318882 formed a subclade; (3) Stauropathes arctica MT318879 nested into a subclade including Bathypathes patula and Bathypathes sp. 1−3. Within Parantipathinae, Umbellapathes parva sp. nov. clustered with Alternatipathes bipinnata with low support (ML 74%, BI 0.90), followed by the Parantipathes subclade (including Sibopathes cf. macrospina), and the group Dendrobathypathes sp. and Lillipathes wingi branched basally with high support (ML 97%; BI 1.00).
|
| Fig.8 Maximum likelihood (ML) tree inferred from the ITS-COI-NAD2 sequences of Schizopathidae and Cladopathidae. The Bayesian inference (BI) tree is identical with the ML tree in topology. Node support is as follows: ML bootstrap /BI posterior probability. The ML bootstrap < 70% or BI posterior probability < 0.90 is not shown. Newly sequenced species are in bold. The GenBank accession numbers of the ITS sequences were listed next to the species names. The Bayesian inference (BI) tree is identical with the ML tree in topology. Node support is as follows: ML bootstrap /BI posterior probability. The ML bootstrap < 70% or BI posterior probability < 0.90 is not shown. Newly sequenced species are in bold. The GenBank accession numbers of the ITS sequences were listed next to the species names. |
In the present study, we added the sequences of Telopathes cf. magna and Umbellapathes parva sp. nov. and the congeneric sequences of Stauropathes for the phylogenetic analyses. Within the Parantipathinae clade, Umbellapathes parva sp. nov. formed a sister subclade with Alternatipathes bipinnata (Fig. 8), which is consistent with their morphological data. Umbellapathes parva sp. nov. is similar to A. bipinnata in growth form of colony, and both of them have a long unpinnulated stalk and lowermost pinnules developing into pinnulated branches (Opresko, 2005). There is even no or only one-nucleotide variation of COI between them. However, Umbellapathes parva sp. nov. is easily distinguished from the latter by the present of secondary pinnules (vs. absent). Furthermore, the relatively far distances (1.72%−1.89%) of ITS1-5.8S-ITS2 supported their separating. In addition, Umbellapathes parva sp. nov. is most similar to U. helioanthes except for the spine characteristics and molecular evidence would be useful to support identifying it as a new species. However, no sequence of U. helioanthes and U. litocrada is available in the GenBank. Future research including the congener sequences are needed to confirm the establishment of the new species.
All the trees showed Telopathes, Stauropathes, and Bathypathes had close relationships (Fig. 8; Supplementary Fig.S1). There was no genetic variation of ITS1-5.8S-ITS2 between Stauropathes arctica MT318879 and Bathypathes sp.1 MT318866, and only four of 17 700 nucleotide differences were found between their mitogenomes, suggesting they belong to the same genus (Supplementary Table S1).
Similarly, the distances between Telopathes sp. MT318882 and Bathypathes sp. MG023167 was 0.17%, indicating they may also belong to the same genus. Due to no available morphological information (not shown in the associated reference Barrett et al., 2020), these sequences need further verification. Excluding the uncertain sequences, the monophylies of Telopathes, Stauropathes, and Bathypathes are not rejected. Nonetheless, sequence data from more classified species of these genera are needed to confirm their monophylies.
5 CONCLUSIONWe report three black corals, Umbellapathes parva sp. nov., Telopathes cf. magna, and Stauropathes cf. punctata, from seamounts in the tropical Western Pacific. The present phylogenetic analyses indicate that Telopathes, Stauropathes, and Bathypathes have close relationships, and Umbellapathes formed a sister clade with Alternatipathes.
6 DATA AVAILABILITY STATEMENTThe specimens described in this study are available at the MBMCAS at IOCAS, Qingdao, China. Voucher IDs: Umbellapathes parva sp. nov.: MBM286410, MBM286411, MBM286412; Telopathes cf. magna: MBM286480, MBM286481, MBM286482, MBM286483; Stauropathes cf. punctata: MBM286474, MBM286475, MBM286476. The ITS sequences that support the findings of this study have been deposited in NCBI GenBank with the accession codes MW285749, MW285750 (Umbellapathes parva sp. nov.), MW285751, MW285752 (Telopathes cf. magna), MW285746, MW285748, MW285747 (Stauropathes cf. punctata). The COI sequences that support the findings of this study have been deposited in NCBI GenBank with the accession codes MW285726, MW285727, MW285728 (Umbellapathes parva sp. nov.), MW285729, MW285730, MW285731, MW285732 (Telopathes cf. magna), MW285733, MW285734, MW285735 (Stauropathes cf. punctata). The NAD2 sequences that support the findings of this study have been deposited in NCBI GenBank with the accession codes MW285743, MW285744, MW285745 (Umbellapathes parva sp. nov.), MW285739, MW285740, MW285741, MW285742 (Telopathes cf. magna), MW285736, MW285737, MW285738 (Stauropathes cf. punctata).
7 ACKNOWLEDGMENTWe thank the assistance of the crew members of R/V Kexue and ROV Faxian for sample collection. Special thanks to Mr. Shaoqing WANG in IOCAS for photographing on board.
Electronic supplementary materialSupplementary material (Supplementary Table S1 & Fig.S1) is available in the online version of this article at https://doi.org/10.1007/s00343-021-0455-9.
Alfaro M E, Zoller S, Lutzoni F. 2003. Bayes or Bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Molecular Biology and Evolution, 20(2): 255-266.
DOI:10.1093/molbev/msg028 |
Barrett N J, Hogan R I, Allcock A L, Molodtsova T, Hopkins K, Wheeler A J, Yesson C. 2020. Phylogenetics and mitogenome organisation in black corals (Anthozoa: Hexacorallia: Antipatharia): an order-wide survey inferred from complete mitochondrial genomes. Frontiers in Marine Science, 7: 440.
DOI:10.3389/fmars.2020.00440 |
Bo M, Barucca M, Biscotti M A, Brugler M R, Canapa A, Canese S, Iacono C L, Bavestrello G. 2018. Phylogenetic relationships of Mediterranean black corals(Cnidaria: Anthozoa: Hexacorallia) and implications for classification within the order Antipatharia. Invertebrate Systematics, 32(5): 1 102-1 110.
DOI:10.1071/IS17043 |
Bo M, Barucca M, Biscotti M A, Canapa A, Lapian H F N, Olmo E, Bavestrello G. 2009. Description of Pseudocirrhipathes (Cnidaria: Anthozoa: Hexacorallia: Antipathidae), a new genus of whip black corals from the Indo-Pacific. Italian Journal of Zoology, 76(4): 392-402.
DOI:10.1080/11250000802684104 |
Cairns S D. 2007. Deep-water corals: an overview with special reference to diversity and distribution of deep-water Scleractinian corals. Bulletin of Marine Science, 81(3): 311-322.
DOI:10.1002/aqc.809 |
Drummond A J, Rambaut A. 2007. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology, 7(1): 214.
DOI:10.1186/1471-2148-7-214 |
Guindon S, Dufayard J F, Lefort V, Anisimova M, Hordijk W, Gascuel O. 2010. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Systematic Biology, 59(3): 307-321.
DOI:10.1093/sysbio/syq010 |
Hall T A. 1999. BioEdit: a user friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41: 95-98.
|
Hillis D M, Bull J J. 1993. An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Systematic Biology, 42(2): 182-192.
DOI:10.1093/sysbio/42.2.182 |
Horowitz J, Brugler M R, Bridge T C L, Cowman P F. 2020. Morphological and molecular description of a new genus and species of black coral (Cnidaria: Anthozoa: Hexacorallia: Antipatharia: Antipathidae: Blastopathes) from Papua New Guinea. Zootaxa, 4821(3): 553-569.
DOI:10.11646/zootaxa.4821.3.7 |
Katoh K, Standley D M. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution, 30(4): 772-780.
DOI:10.1093/molbev/mst010 |
Lapian H F N, Barucca M, Bavestrello G, Biscotti M A, Bo M, Canapa A, Tazioli S, Olmo E. 2007. A systematic study of some black corals species (Antipatharia, Hexacorallia) based on rDNA internal transcribed spacers sequences. Marine Biology, 151(2): 785-792.
DOI:10.1007/s00227-006-0525-8 |
Lefort V, Longueville J E, Gascuel O. 2017. SMS: Smart Model Selection in PhyML. Molecular Biology and Evolution, 34(9): 2 422-2 424.
DOI:10.1093/molbev/msx149 |
Lima M A, Cordeiro R T S, Perez C D. 2019. Black Corals(Anthozoa: Antipatharia) from the southwestern Atlantic. Zootaxa, 4692(1): 1-67.
DOI:10.11646/zootaxa.4692.1.1 |
MacIsaac K G, Best M, Brugler M R, Kenchington E L R, Anstey L J, Jordan T. 2013. Telopathes magna gen. nov., spec. nov. (Cnidaria: Anthozoa: Antipatharia: Schizopathidae) from deep waters off Atlantic Canada and the first molecular phylogeny of the deep-sea family Schizopathidae. Zootaxa, 3700(2): 237-258.
DOI:10.11646/zootaxa.3700.2.3 |
Molodtsova T N, Opresko D M. 2017. Black corals (Anthozoa: Antipatharia) of the Clarion-Clipperton fracture zone. Marine Biodiversity, 47(2): 349-365.
DOI:10.1007/s12526-017-0659-6 |
Molodtsova T N. 2006. Black corals (Antipatharia: Anthozoa: Cnidaria) of the north-eastern Atlantic. In: Mironov A N, Gebruk A V, Southward A J eds. Biogeography of the Atlantic Seamounts. KMK Scientific Press, Moscow. p. 22-57.
|
Opresko D M, Goldman S L, Johnson R, Parra K, Nuttall M, Schmahl G P, Brugler M R. 2020. Morphological and molecular characterization of a new species of black coral from Elvers Bank, north-western Gulf of Mexico(Cnidaria: Anthozoa: Hexacorallia: Antipatharia: Aphanipathidae: Distichopathes). Journal of the Marine Biological Association of the United Kingdom, 100(4): 559-566.
DOI:10.1017/S002531542000051X |
Opresko D M, Tracey D M, Mackay E. 2014. Antipatharia(black corals) for the New Zealand region: a field guide of commonly sampled New Zealand black corals including illustrations highlighting technical terms and black coral morphology. New Zealand Aquatic Environment and Biodiversity Report, 131: 20.
|
Opresko D M. 2002. Revision of the Antipatharia (Cnidaria: Anthozoa). Part II. Schizopathidae. Zoologische Mededelingen, 76(17-24): 411-442.
|
Opresko D M. 2005. New genera and species of antipatharian corals (Cnidaria: Anthozoa) from the North Pacific. Zoologische Mededelingen, 79(2): 129-165.
|
Opresko D M. 2019. New species of black corals (Cnidaria: Anthozoa: Antipatharia) from the New Zealand region, part 2. New Zealand Journal of Zoology, 47(3): 149-186.
DOI:10.1080/03014223.2019.1650783 |
Ronquist F, Huelsenbeck J P. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19(12): 1 572-1 574.
DOI:10.1093/bioinformatics/btg180 |
Roule L. 1905. Description des Antipathaires et Cérianthaires recueillis par S.A.S. le Prince de Monaco dans l'Atlantique nord. (1886-1902). Résultats des Campagnes Scientifiques du Prince de Monaco, 30: 1-99.
DOI:10.5962/bhl.title.59328 |
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. 2013. MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30(12): 2 725-2 729.
DOI:10.1093/molbev/mst197 |
Terrana L, Bo M, Opresko D M, Eeckhaut I. 2020. Shallowwater black corals (Cnidaria: Anthozoa: Hexacorallia: Antipatharia) from SW Madagascar. Zootaxa, 4826(1): 1-62.
DOI:10.11646/zootaxa.4826.1.1 |
Thoma J N, Pante E, Brugler M R, France S C. 2009. Deep-sea octocorals and antipatharians show no evidence of seamount-scale endemism in the NW Atlantic. Marine Ecology Progress, 397: 25-35.
DOI:10.3354/meps08318 |
Wagner D, Brugler M R, Opresko D M, France S C, Montgomery A D, Toonen R J. 2010. Using morphometrics, in situ observations and genetic characters to distinguish among commercially valuable Hawaiian black coral species; a redescription of Antipathes grandis Verrill, 1928(Antipatharia: Antipathidae). Invertebrate Systematics, 24(3): 271-290.
DOI:10.1071/IS10004 |
Wagner D, Luck D G, Toonen R J. 2012. Chapter Two-The biology and ecology of black corals (Cnidaria: Anthozoa: Hexacorallia: Antipatharia). Advances in Marine Biology, 63: 67-132.
|
Wagner D, Opresko D M. 2015. Description of a new species of Leiopathes (Antipatharia: Leiopathidae) from the Hawaiian Islands. Zootaxa, 3974(2): 277-289.
DOI:10.11646/zootaxa.3974.2.11 |
Wörheide G. 1998. The reef cave dwelling ultraconservative coralline demosponge Astrosclera willeyana Lister, 1900 from the Indo-Pacific. Facies, 38(1): 1-88.
|
 2021, Vol. 39
2021, Vol. 39