Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZHANG Yunlei, XUE Ying, XU Binduo, ZHANG Chongliang, ZAN Xiaoxiao
- Evaluating the effect of input variables on quantifying the spatial distribution of croaker Johnius belangerii in Haizhou Bay, China
- Journal of Oceanology and Limnology, 39(4): 1570-1583
- http://dx.doi.org/10.1007/s00343-020-0193-4
Article History
- Received May. 14, 2020
- accepted in principle Jun. 28, 2020
- accepted for publication Sep. 23, 2020
2 Marine College, Shandong University, Weihai 264209, China;
3 Field Observation and Research Station of Haizhou Bay Fishery Ecosystem, Ministry of Education, Qingdao 266003, China
Identifying the habitat and spatial distribution of species is important for fishery management and marine biological conservation (Bailey and Thompson, 2009), especially for species that are susceptible to environmental variations (Sagarese et al., 2014; Peck et al., 2018). With fisheries management moving to ecosystem-based approaches (Pikitch et al., 2004; Dolan et al., 2016), spatial information for target species becomes even more important (Giannoulaki et al., 2013; Marzloff et al., 2016; Xue et al., 2018). Several data-driven multivariate statistical methods like multiple linear regression, generalized linear models (GLM), generalized additive model (GAM) and machine learning methods are usually used for analyzing the relationship between explanatory variables and habitats of species (Guisan et al., 2002; AhmadiNedushan et al., 2006; Jordaan et al., 2010; Li et al., 2017; Liu et al., 2019). Among them, GAM can determine the complex nonlinear relationship between response variables and input variables (Kitchens and Rooker, 2014), thus providing a flexible and robust method for predicting the spatial distribution of species (Schmiing et al., 2013; Zerbini et al., 2016).
The limited habitat range, low density, and low detection rate might result in a large number of zero catches (Martin et al., 2005; Virgili et al., 2017) and positive skewed distribution of species abundance. Excessive zeroes will increase the bias and uncertainty in estimation of model parameters (Martin et al., 2005), which is a challenge for determining the existence of the specie (Lee et al., 2006; Virgili et al., 2017). In addition, ecological data are often overdispersed with the variance being greater than the mean. Therefore, appropriate process of explanatory variables before habitat modelling is necessary to avoid biased predictions (Lee et al., 2006).
The spatial distribution of species can be predicted in habitat models based on environmental variables (Guisan and Thuiller 2005; Gormley et al., 2011; Virgili et al., 2017). Abiotic variables, such as spatial variables and environmental variables, are often used as explanatory variables to predict the spatiotemporal distribution of species. However, biotic variables such as prey data are also one of the important factors driving the predator's distribution and cannot be ignored in species distribution models (SDMs) (Bi et al., 2011; Murase et al., 2013; Arrizabalaga et al., 2015; Zerbini et al., 2016). Including prey data has been proved capable of improving the predictive skills of SDMs (Cormon et al., 2014; Vezza et al., 2015; Xue et al., 2018). However, prey data have rarely been considered due to the lack of information of non-commercial prey species, which may lead to incomplete and biased predictions of predator's distribution (Johnson et al., 2013). Understanding the relationship between species distribution and related abiotic and biotic variables can provide critical information to predict abundance and identify suitable habitats of marine organisms (Murase et al., 2013; Vezza et al., 2015).
Multicollinearity between variables needs to be assessed and addressed before habitat modeling (Xue et al., 2018). However, removing some variables by statistical methods such as residual regression, ordinary least squares, and ridge regression, may lead to important information missing. In this case, the principal component analysis (PCA) is preferred to reduce the dimension of variables with the principal components (PCs), which can effectively retain significant information and avoid collinearity (Ahmadi-Nedushan et al., 2006; Buisson et al., 2008). In addition, incorporating spatial autocovariate (SAC) of target species into habitat models can also improve the performance of SDMs, especially when significant SAC was detected in the residuals (Wu et al., 2009).
The aim of our study was to evaluate the effect of input variables on the performance of habitat models. Belanger's croaker (Johnius belangerii) collected from the bottom trawl surveys in Haizhou Bay with a large number of zero catches was selected as target species. We evaluated the effects of different types of input variables (i.e., abiotic variables, abiotic and biotic variables, abiotic variables and SAC, abiotic and biotic variables and SAC, and PCA-based variables) on the performance of GAM, and their ability to deal with the over-dispersed dataset were also evaluated. The process of input variables selection is critical in habitat modelling, and optimal combination of input variables is expected to improve the predictive performance of habitat models and deepen our understanding of species spatiotemporal distributions.
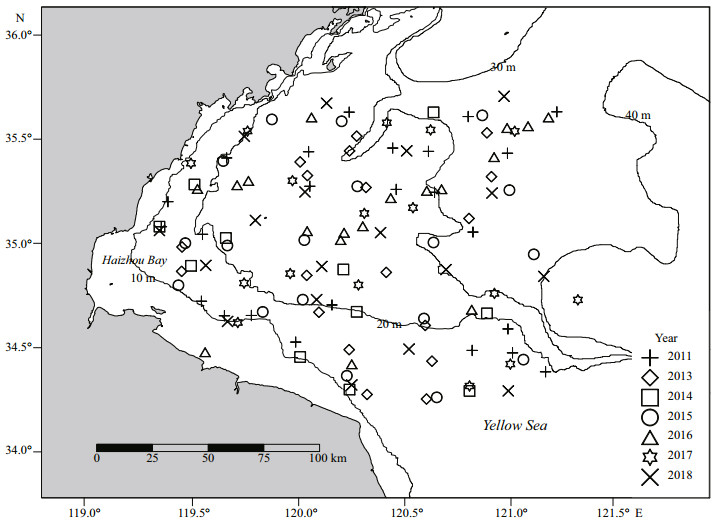
2 MATERIAL AND METHOD 2.1 Data collectionBelanger's croakers (Johnius belangerii) were collected from bottom trawl surveys in Haizhou Bay and the adjacent areas on spring (May) in 2011 and 2013–2018 (Fig. 1). The survey area spans 119°20′E– 121°10′E, 34°20′N–35°40′N. Stratified random sampling was used for investigation (Xu et al., 2015). In accordance with the differences in geological, oceanographic and biological characteristics of the survey area, a certain number of stations were randomly selected in each stratification (Xu et al., 2015). Twenty-four stations were selected in 2011, and the sampling stations were optimized to 18 after 2013 (Xu et al., 2015; Zhang et al., 2020). The bottom trawl was towed at 2–3 knots for about 1 h, with a trawl mesh size of 17 mm and net of 12 m width (Zhang et al., 2020). Tows were only conducted during daytime (6∶00–18∶00). At each survey station, GPS was used to record the sampling location. At the same times, CTD system (XR-420) was used to measure environmental data including temperature, salinity, and depth. The distance between sampling station and the nearest coastline was determined geographically (Zhang et al., 2020). To account for spatial autocorrelation, we included a measure of the surrounding mean abundance of J. belangerii in the Voronoi diagram (Muramatsu et al., 2018) as a SAC (Fig. 2). The survey data were standardized by 2 knots and 1 hour, and the catch number per unit area (inds./ km2) was used as the abundance index (Liu et al., 2019; Zhang et al., 2020).
|
| Fig.1 The survey areas in the Haizhou Bay and the adjacent areas in 2011 and 2013–2018 |
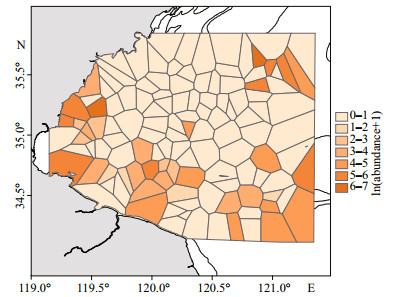
|
| Fig.2 The Voronoi diagram of J. belangerii abundance in the study areas in the springs of 2011 and 2013–2018 |
The stomach contents of J. belangerii were analyzed based on 166 stomach samples. In laboratory, prey items were identified to the lowest possible taxonomic level and calculated weight for each content (Paquin et al., 2014). The index of relative importance (IRI) and the percentage of relative importance index (%IRI) were calculated to describe the relative contribution of prey species in the diet (Guedes et al., 2015).
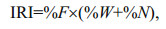 (1)
(1)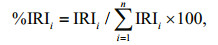 (2)
(2)where %W is the percent of weight of prey species, %N is percent of number, %F is frequency of occurrence, and n is the total number of prey items.
2.3 Candidate explanatory variablesThe abiotic explanatory variables included bottom temperature, bottom salinity, depth, distance offshore, and year. The dominant prey species of J. belangerii were chosen as biotic variables in the modeling. The prey species were synchronously collected in each survey station by bottom trawl surveys in Haizhou Bay. The mean abundance of J. belangerii in the Voronoi diagram was chosen as SAC. In this study, five types of input variables were used for modeling, including (1) abiotic variables only, (2) abiotic and biotic variables, (3) SAC and abiotic variables, (4) SAC and abiotic and biotic variables, and (5) PCAbased SAC, abiotic, and biotic variables. Before modeling, highly correlated explanatory variables were excluded from model by variance inflation factors (VIF) to avoid collinearity (Li et al., 2017; Luan et al., 2018). Previous studies showed that the variable whose 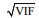
GAM is the generalization of linear regression model and additive in the predictor effects (Zhang et al., 2020). GAM is expressed as:
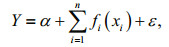 (3)
(3)where Y is abundance of species, α is the intercept term, fi() is a smooth function of explanatory variables spline, xi are explanatory variables, n is the number of predictors, and ε is the residual error term (Hastie and Tibshirani, 1986; Zhang et al., 2020).
For the PCA based GAM, we applied PCA to analyze the uncorrelated explanatory variables. The PCA informed GAM can be described as follows:
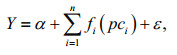 (4)
(4)where pci are principal components. In order to avoid overfitting, all the smoothers were constrained with 4 knots. The models were fitted with tw function (Tweedie distribution) using mgcv package of R software (Wood, 2006). The principal components were chosen with eigenvalue more than 1 (Buisson et al., 2008; Liu et al., 2019).
The response variable was linked to the additive predictors using a log-link function. The significant variables were selected by stepwise procedure. It started with a null model and added one predictive variable to the present model at each step (Luan et al., 2018; Zhang et al., 2020). Akaike Information Criterion (AIC) (Akaike, 1998) was used for variable selection, in which GAM with the lowest AIC value was regarded as the best model (Luan et al., 2018; Zhang et al., 2020). We extracted the percentage of variance explained, and checked the distribution of residuals for all models. Important variables were selected step by step in procedure starting with an empty model and adding a predictor variable to the current model in each step.
2.5 Model evaluationWe evaluated the performance of different GAMs in describing the spatial distribution of J. belangerii in Haizhou Bay. The abundance was log-transformed (ln(x+1)) to reduce the influence of extreme values and meet the requirement of error distribution in GAMs (Zhang et al., 2018a). Cross-validation was applied to assess the performance of GAMs (Guisan and Zimmermann, 2000; Mouquet et al., 2015; Yu et al., 2018). About 70% of the data subset was randomly selected for model development, and the remaining 30% were used to evaluate the performance of the habitat models (Smith, 1994; Zuur et al., 2007; Zhang et al., 2020). For each GAM, cross-validations were run for 100 times and linear regression between the observed and predicted abundance of J. belangerii was analyzed at each run (Xue et al., 2018, Zhang et al., 2018a). The predicted abundance distributions were mapped using ordinary kriging in AcrGIS. The performance of each GAM was evaluated according to the regression coefficient of determination (R2), the root mean square error (RMSE), the intercept, and the slope between the predicted and observed abundance data (Brennan et al., 2019). The model with the maximum R2, the minimum RMSE was selected as the optimal model (Brennan et al., 2019).
3 RESULTThe abundance of J. belangerii in the studied area ranged from 0 to 826.464 inds./km2, on average of 40.841±10.282 inds./km2. Inter-annual variation of the abundance was observed, from the highest (118.064±50.185 inds./km2) in 2016 to the lowest (4.354±3.500 inds./km2) in 2018.
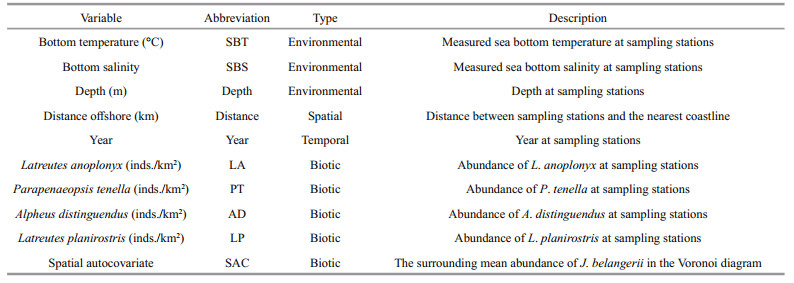
3.1 Diet compositionDecapoda and amphipoda were main prey groups of J. belangerii, with %IRI being 45.248% and 50.403%, respectively, followed by polychaeta (2.323%), and euphausiacea (1.798%). There were 19 prey species in total, of which Latreutes anoplonyx, Parapenaeopsis tenella, Alpheus distinguendus, and Latreutes planirostris dominated (Table 1). The total weight percentage in the diet was 55.336%. These four major prey species were selected as the biotic variables in habitat modeling.
Johnius belangerii mainly concentrated in the center and southern coastal waters of Haizhou Bay. Difference in spatial distribution of the four prey species was obvious (Fig. 3). L. anoplonyx distributed mainly in the southeastern and center area of Haizhou Bay. P. tenella and A. distinguendus gathered in the central and southern coastal waters, which is similar to the distribution of J. belangerii. However, L. planirostris amassed in the central and northern areas of Haizhou Bay, which was different from J. belangerii and other prey in the regard.
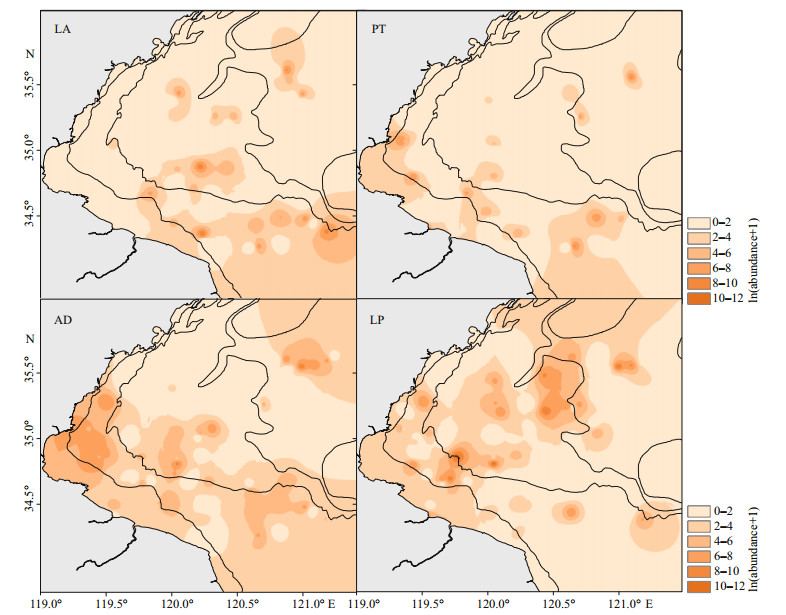
|
| Fig.3 The spatial distribution of four major prey species of J. belangerii during spring in 2011 and 2013–2018 in Haizhou Bay Ordinary kriging was applied to generate the maps. LA: Latreutes anoplonyx; PT: Parapenaeopsis tenella; AD: Alpheus distinguendus; LP: Latreutes planirostris. |
Johnius belangerii was found to mainly inhabit in the areas overlapped with its main prey species (Fig. 4). In addition, there was a significant correlation (R2=0.527, P < 0.01) between abundance of J. belangerii and its four major prey species. The fluctuation patterns in abundance between J. belangerii and the four prey species were similar from 2011 to 2018.
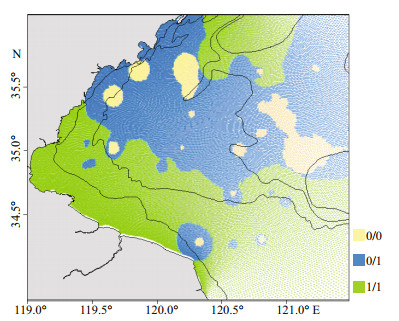
|
| Fig.4 The spatial location relationship between J. belangerii and its four main prey species Yellow (0/0): both J. belangerii and main prey species are absent; blue (0/1): J. belangerii is absent, but its main prey species are present; green (1/1): J. belangerii and its main prey species are all present. |
Nine potential explanatory variables were selected as candidate variables to explore the performance of GAMs (Table 2).
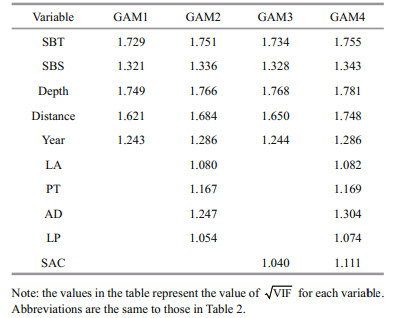
Overall, four GAMs with different combination of candidate variables were built, including GAM1 with only abiotic variables, GAM2 with abiotic variables and SAC, GAM3 with abiotic and biotic variables, and GAM4 with all abiotic, biotic variables and SAC. The degree of multicollinearity was measured by VIF (Table 3). The 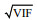
|
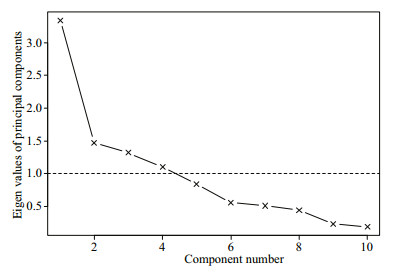
The PCA revealed four optimal principal components (Fig. 5) whose cumulative of variance was 0.72 (Table 4). The higher loading values in PC1 were contributed by abiotic variables with proportion of variance at 0.30, while those for biotic variables, year and biotic variables, and SAC variables were 0.16, 0.15, and 0.11, respectively (Table 4).
|
| Fig.5 The principal components selected in habitat modeling The crushed stone diagram shows the principal components (black line with × symbol), eigenvalue for 1 (dashed line). |
GAMs with different input variables exhibited different performance for model fitting. The explained deviances ranged from 19.40% for GAM1 with only abiotic variables to 55.60% for GAM5 with PCA based variables (Table 5). In addition, the highest determination coefficient (R2) was found in GAM5 (0.497), while the lowest R2 was in GAM1 (0.162). Overall, GAM5 with PCA based variables performed better than the other GAMs, which had the highest values of R2 and deviance explained.
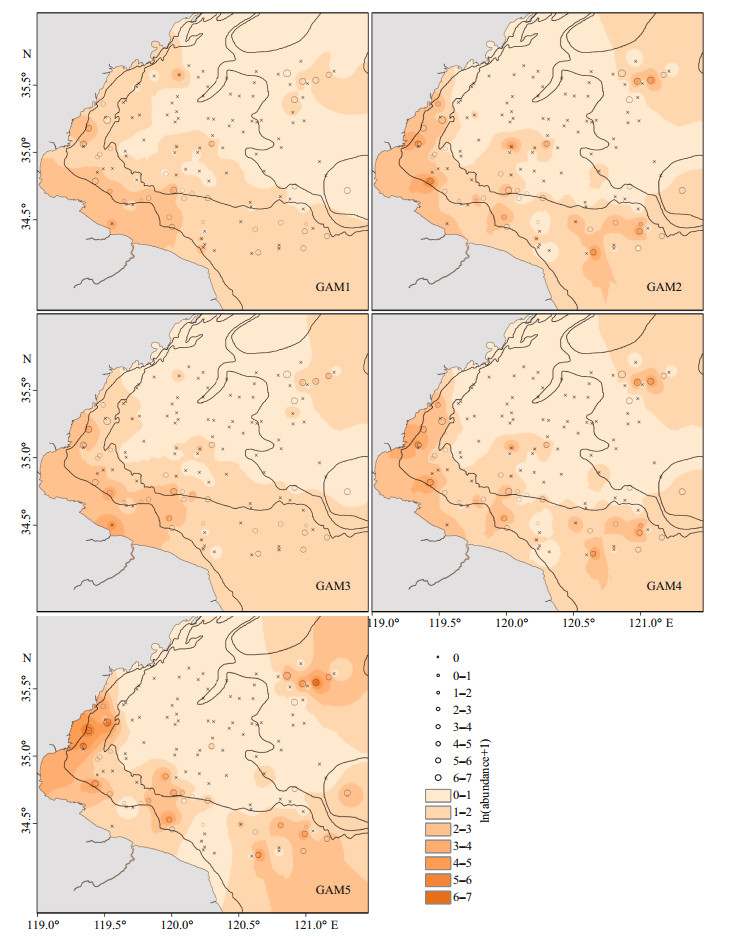
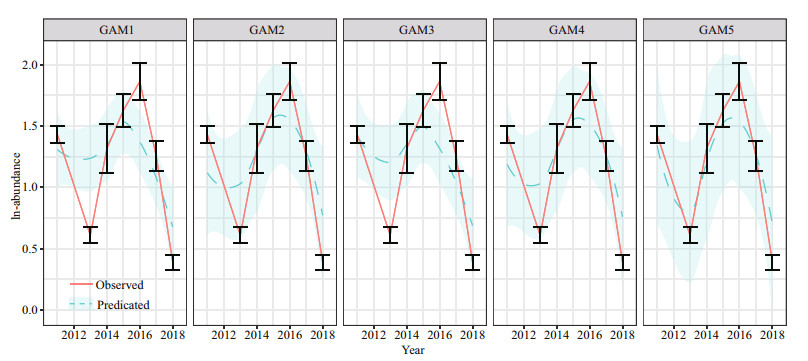
The predicted abundance of J. belangerii incorporating biotic variables and SAC into input variables had higher correlation with observed abundance, especially for GAM5 with PCA based variables (Fig. 6). The correlation coefficients between predicted and observed abundance of J. belangerii during spring from 2011 to 2018 in Haizhou Bay ranged from 0.445 (GAM1) to 0.808 (GAM5) (Fig. 7).
|
| Fig.6 Overlaid maps of observed (black circles) and predicted (color contours) abundance of J. belangerii based on five GAMs during spring in Haizhou Bay Predictions were made for each sampling location by ordinary kriging. |
|
| Fig.7 Predicated and observed mean abundance of J. belangerii during spring from 2011 to 2018 in Haizhou Bay based on five GAMs Shaded regions represent 95% confidence intervals. Error bars represent the mean observed abundance (±S.E.). |
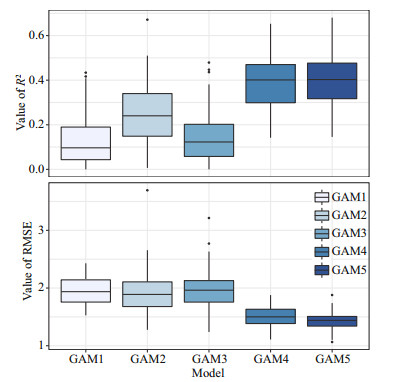
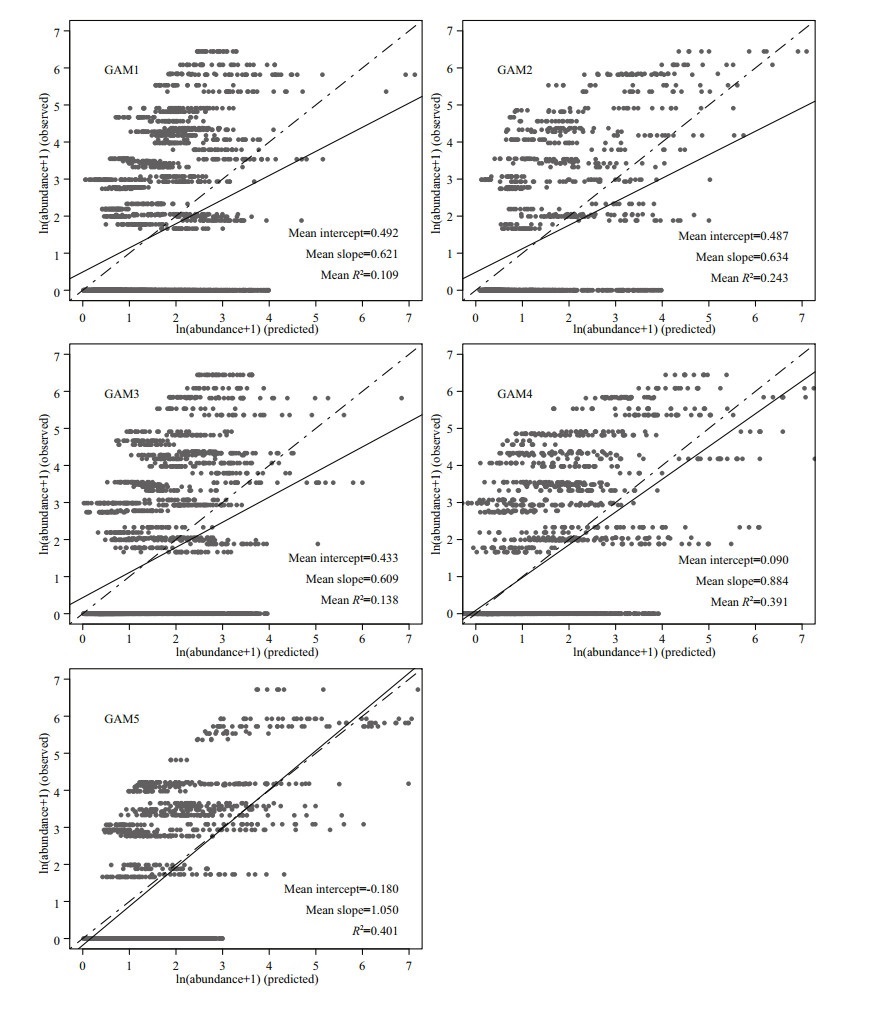
For cross-validation, R2 of all GAMs ranged from 0.01 to 0.68 and RMSE ranged from 1.06 to 3.69 (Fig. 8). Among the five GAMs, GAM5 showed the best performances than other models, with the highest R2 and lowest RMSE. The overall predictive performance of the five GAMs was evaluated by linear regression analyses between observed and predicted abundance of J. belangerii. Results reveal that GAM5 performed better than the other four GAMs, with the highest slope and R2 closest to 1. A tendency to underestimate the abundance of J. belangerii was observed in the other four GAMs (Fig. 9), and the degree of underestimation increased with higher prevalence.
|
| Fig.8 R2 and RMSE resulted from 100 times crossvalidations of five generalized additive models |
|
| Fig.9 The linear regression analysis (100 iterations of cross-validations) between observed and predicted abundance of J. belangerii with five generalized additive models (GAMs) Solid line: mean of 100 linear regressions; dashed line: unbiased relationship (1∶1) between observed and predicted abundance; grey dots: abundance from cross-validation. |
Habitat models can be used to identify key areas of protection to meet stakeholder's expectations (Cañadas et al., 2005). However, it is difficult to find a balance between high explanatory power and the elimination of random noise in the model (Li et al., 2015). In addition, for response variable, neglecting the effect of over-dispersion may lead to the selection of models to be more complex than necessary (Richards, 2008), where the model is not generalized outside of the sample used for calibration (Virgili et al., 2017). Therefore, selecting appropriate model and input data are critical in the habitat modelling, especially for species that are barely detected, with their high-density regions being difficult to be identified.
Johnius belangerii is one of the dominant fish species in the study area (Zhang et al., 2018b). It is a demersal fish and likes to inhabit shallow waters with warm temperate. Spring is the main spawning season and Haizhou Bay serves as the main spawning ground for J. belangerii. Based on the three environmental variables, i.e. SBT, SBS, and depth, Zhang et al. (2018b) studied the habitat suitability of J. belangerii during spring in Haizhou Bay. The study found that depth and SBT were the main factors affecting the distribution of J. belangerii. The optimal habitat of J. belangerii varied with life history stage. The distribution of suitable habitat was closely related to external environmental factors. J. belangerii has a patched spatial distribution pattern. In trawl surveys, more than 70% of sampling stations have the abundance value of zero, which has an obvious overdispersed dataset. Compared with other generalist feeders, J. belangerii has more specified feeding habits (Xiao et al., 2019), with a few major prey species that may have significant impacts on its distributions. In this study, we evaluated the effect of input variables on the performance of habitat models. Results show that GAM with PCA based variables (GAM5) exhibited the highest R2, lowest RMSE, and the slope closest to 1, which can be used to predict the spatial distribution of J. belangerii and avoid overestimating or underestimating the abundance.
The relationship between spatial distribution of fish and explanatory variables is usually nonlinear and complex (Zhao et al., 2014). PCA is one of the methods being used to reduce the dimension of variables, retain important information, and avoid collinearity between variables (Ahmadi-Nedushan et al., 2006; Buisson et al., 2008). In GAM5, important factors can be reflected in the PCs and were identified as new input variables. The PCA-GAM was proved to be able to reduce the multicollinearity between explanatory variables in GAM, and improve the performance of the species distribution model.
The predictive ability of habitat models could be limited by the explanatory variables collected from different sources (Dambach and Rödder, 2011). The spatial distribution of prey is one of the factors driving the distribution of predators through bottom-up effect (Bi et al., 2011; Zerbini et al., 2016). Understanding of the spatial relationship between fish and their prey can help to discover the hot spots of feeding and the spatial distribution of predators (Lasley-Rasher et al., 2015). The spatial overlap between predators and their prey species can improve the feeding efficiency and reduce the energy loss of foraging. The availability of prey species can also compensate for some unfavorable factors, such as the effects of temperature and salinity (Youcef et al., 2013). That is probably the reason why prey species is one of the important drivers for the distribution of J. belangerii. Lack consideration of predator-prey interactions may lead to biased predictions of species distributions.
Previous study revealed the necessity to incorporate SAC into habitat models via spatial methods when significant SAC was detected in the residuals (Wu et al., 2009). We found that the GAMs incorporating SAC showed substantial reductions in residual spatial autocorrelation (Supplementary Fig.S1). Through the addition of SAC in GAMs, an extra part of the variance was explained and more accurate estimates of regression coefficients were provided (Jewell et al., 2007; Wu et al., 2009). This study showed that GAMs that ignore SAC in ecological data might misestimate the spatial distribution of target species, which may result in improper management recommendations.
Information on the spatial distribution of target species is important for the management and monitoring of fishery resources. The development of marine protected areas (MPA) and ecosystem-based fishery management (EBFM) need to assess the temporal and spatial distribution of target species and their relationship with environmental factors, particularly in areas under severe climate changes. In addition, fisheries managers wish to obtain reliable species distribution map to identify essential habitats for species (Vinagre et al., 2006; Chang et al., 2012). However, EBFM is often hampered by the lack of information on habitats and distribution of important species (Pikitch et al., 2004). Understanding the spatiotemporal dynamics of species distribution and its relationship with environmental variables is critical for the implementation of EBFM and the habitat assessment (Liu et al., 2019). Neglecting the effects of biotic variables and SAC may result in biased habitat models. This study showed that PCA-based GAM incorporating SAC of target species could improve the performance of habitat model, which can better serve management strategies and identify Essential Fish Habitats (EFH) (Xue et al., 2018).
The lack of field observations of prey species and spatial autocorrelation of predator could reduce the accuracy of SDMs (Wu et al., 2009; Xue et al., 2018). Based on our findings and previous study (Xue et al., 2018), the habitat models of marine organisms should monitor both predators and their important prey species. Further investigation of the interaction between predators and prey is needed in future. The differences in the growth and dynamics of both predators and prey should also be fully taken into account to improve the accuracy of habitat models, which will help to provide more support for the conservation and sustainable utilization of fishery resources. In addition, the impact of climate change on marine life in small-scale sea areas has received increasing attention from researchers (Bates et al., 2018). The spatial-temporal distribution of target species over different life history stage under climate change should also be considered in future work.
5 DATA AVAILABILITY STATEMENTThe data generated or analyzed in this study are available from the corresponding author on reasonable request.
ACKNOWLEDGMENT: We are grateful to many colleagues and graduate students for assistance in sample collection and analyses in laboratory and in the field. We also thank two anonymous reviewers for their valuable and constructive comments on this paper.
Ahmadi-Nedushan B, St-Hilaire A, Bérubé M, Robichaud É, Thiémonge N, Bobée B. 2006. A review of statistical methods for the evaluation of aquatic habitat suitability for instream flow assessment. River Research and Applications, 22(5): 503-523.
DOI:10.1002/rra.918 |
Akaike H. 1998. Information theory and an extension of the maximum likelihood principle. In: Parzen E, Tanabe K, Kitagawa G eds. Selected Papers of Hirotugu Akaike. Springer, New York, https://doi.org/10.1007/978-1-4612-1694-0_15.
|
Arrizabalaga H, Dufour F, Kell L, Merino G, Ibaibarriaga L, Chust G, Irigoien X, Santiago J, Murua H, Fraile I, Chifflet M, Goikoetxea N, Sagarminaga Y, Aumont O, Bopp L, Herrera M, Fromentin J M, Bonhomeau S. 2015. Global habitat preferences of commercially valuable tuna. Deep Sea Research Part II: Topical Studies in Oceanography, 113: 102-112.
DOI:10.1016/j.dsr2.2014.07.001 |
Bailey H, Thompson P M. 2009. Using marine mammal habitat modelling to identify priority conservation zones within a marine protected area. Marine Ecology Progress Series, 378: 279-287.
DOI:10.3354/meps07887 |
Bates A E, Helmuth B, Burrows M T, Duncan M I, Garrabou J, Guy-Haim T, Lima F, Queiros A M, Seabra R, Marsh R, Belmaker J, Bensoussan N, Dong Y W, Mazaris A D, Smale D, Wahl M, Rilov G. 2018. Biologists ignore ocean weather at their peril. Nature, 560(7718): 299-301.
DOI:10.1038/d41586-018-05869-5 |
Bi H S, Peterson W T, Lamb J, Casillas E. 2011. Copepods and salmon: characterizing the spatial distribution of juvenile salmon along the Washington and Oregon coast, USA. Fisheries Oceanography, 20(2): 125-138.
DOI:10.1111/j.1365-2419.2011.00573.x |
Brennan C E, Maps F, Gentleman W C, Plourde S, Lavoie D, Chassé J, Lehoux C, Krumhansl K A, Johnson C L. 2019. How transport shapes copepod distributions in relation to whale feeding habitat: demonstration of a new modelling framework. Progress in Oceanography, 171: 1-21.
DOI:10.1016/j.pocean.2018.12.005 |
Buisson L, Blanc L, Grenouillet G. 2008. Modelling stream fish species distribution in a river network: the relative effects of temperature versus physical factors. Ecology of Freshwater Fish, 17(2): 244-257.
DOI:10.1111/j.1600-0633.2007.00276.x |
Cañadas A, Sagarminaga R, De Stephanis R, Urquiola E, Hammond P S. 2005. Habitat preference modelling as a conservation tool: proposals for marine protected areas for cetaceans in southern Spanish waters. Aquatic Conservation: Marine and Freshwater Ecosystems, 15(5): 495-521.
DOI:10.1002/aqc.689 |
Chang Y J, Sun C L, Chen Y, Yeh S Z, Dinardo G. 2012. Habitat suitability analysis and identification of potential fishing grounds for swordfish, Xiphias gladius, in the South Atlantic Ocean. International Journal of Remote Sensing, 33(23): 7 523-7 541.
DOI:10.1080/01431161.2012.685980 |
Cormon X, Loots C, Vaz S, Vermard Y, Marchal P. 2014. Spatial interactions between saithe (Pollachius virens) and hake (Merluccius merluccius) in the North Sea. ICES Journal of Marine Science, 71(6): 1 342-1 355.
DOI:10.1093/icesjms/fsu120 |
Dambach J, Rödder D. 2011. Applications and future challenges in marine species distribution modeling. Aquatic Conservation: Marine and Freshwater Ecosystems, 21(1): 92-100.
DOI:10.1002/aqc.1160 |
Dolan T E, Patrick W S, Link J S. 2016. Delineating the continuum of marine ecosystem-based management: a US fisheries reference point perspective. ICES Journal of Marine Science, 73(4): 1 042-1 050.
DOI:10.1093/icesjms/fsv242 |
Giannoulaki M, Iglesias M, Tugores M P, Bonanno A, Patti B, Felice A D, Leonori I, Bigot J L, Tičina V, Pyrounaki M M, Tsagarakis K, Machias A, Somarakis S, Schismenou E, Quinci E, Basilone G, Cuttitta A, Campanella F, Miquel J, Oñate D, Valavanis V. 2013. Characterizing the potential habitat of European anchovy Engraulis encrasicolus in the Mediterranean Sea, at different life stages. Fisheries Oceanography, 22(2): 69-89.
DOI:10.1111/fog.12005 |
Gormley A M, Forsyth D M, Griffioen P, Lindeman M, Ramsey D S L, Scroggie M P, Woodford L. 2011. Using presence-only and presence-absence data to estimate the current and potential distributions of established invasive species. Journal of Applied Ecology, 48(1): 25-34.
DOI:10.1111/j.1365-2664.2010.01911.x |
Guedes A P P, Araújo F G, Pessanha A L M, Milagre R R. 2015. Partitioning of the feeding niche along spatial, seasonal and size dimensions by the fish community in a tropical Bay in Southeastern Brazil. Marine Ecology, 36(1): 38-56.
DOI:10.1111/maec.12115 |
Guisan A, Edwards Jr T C, Hastie T. 2002. Generalized linear and generalized additive models in studies of species distributions: setting the scene. Ecological Modelling, 157(2-3): 89-100.
DOI:10.1016/S0304-3800(02)00204-1 |
Guisan A, Thuiller W. 2005. Predicting species distribution: offering more than simple habitat models. Ecology Letters, 8(9): 993-1.
DOI:10.1111/j.1461-0248.2005.00792.x |
Guisan A, Zimmermann N E. 2000. Predictive habitat distribution models in ecology. Ecological Modelling, 135(2-3): 147-186.
DOI:10.1016/s0304-3800(00)00354-9 |
Hastie T, Tibshirani R. 1986. Generalized additive models. Statistical Science, 1(3): 297-310.
DOI:10.1214/ss/1177013604 |
Jewell K J, Arcese P, Gergel S E. 2007. Robust predictions of species distribution: spatial habitat models for a brood parasite. Biological Conservation, 140(3-4): 259-272.
DOI:10.1016/j.biocon.2007.08.017 |
Johnson A F, Jenkins S R, Hiddink J G, Hinz H. 2013. Linking temperate demersal fish species to habitat: scales, patterns and future directions. Fish and Fisheries, 14(3): 256-280.
DOI:10.1111/j.1467-2979.2012.00466.x |
Jordaan A, Chen Y, Townsend D W, Sherman S. 2010. Identification of ecological structure and species relationships along an oceanographic gradient in the Gulf of Maine using multivariate analysis with bootstrapping. Canadian Journal of Fisheries and Aquatic Sciences, 67(4): 701-719.
DOI:10.1139/F10-010 |
Kabacoff R. 2015. R in Action: Data Analysis and Graphics with R. 2nd ed. Manning, New York.
|
Kitchens L L, Rooker J R. 2014. Habitat associations of dolphinfish larvae in the Gulf of Mexico. Fisheries Oceanography, 23(6): 460-471.
DOI:10.1111/fog.12081 |
Lasley-Rasher R S, Brady D C, Smith B E, Jumars P A. 2015. It takes guts to locate elusive crustacean prey. Marine Ecology Progress Series, 538: 1-12.
DOI:10.3354/meps11481 |
Lee A H, Wang K, Scott J A, Yau K K W, Mclachlan G J. 2006. Multi-level zero-inflated Poisson regression modelling of correlated count data with excess zeros. Statistical Methods in Medical Research, 15(1): 47-61.
DOI:10.1191/0962280206sm429oa |
Li B, Cao J, Chang J H, Wilson C, Chen Y. 2015. Evaluation of effectiveness of fixed-station sampling for monitoring American lobster settlement. North American Journal of Fisheries Management, 35(5): 942-957.
DOI:10.1080/02755947.2015.1074961 |
Li M, Zhang C L, Xu B D, Xue Y, Ren Y P. 2017. Evaluating the approaches of habitat suitability modelling for whitespotted conger (Conger myriaster). Fisheries Research, 195: 230-237.
DOI:10.1016/j.fishres.2017.07.024 |
Liu X X, Wang J, Zhang Y L, Yu H M, Xu B D, Zhang C L, Ren Y P, Xue Y. 2019. Comparison between two GAMs in quantifying the spatial distribution of Hexagrammos otakii in Haizhou Bay, China. Fisheries Research, 218: 209-217.
DOI:10.1016/j.fishres.2019.05.019 |
Luan J, Zhang C L, Xu B D, Xue Y, Ren Y P. 2018. Modelling the spatial distribution of three Portunidae crabs in Haizhou Bay, China. PLoS One, 13(11): e0207457.
DOI:10.1371/journal.pone.0207457 |
Martin T G, Wintle B A, Rhodes J R, Kuhnert P M, Field S A, Low-Choy S J, Tyre A J, Possingham H P. 2005. Zero tolerance ecology: improving ecological inference by modelling the source of zero observations. Ecology Letters, 8(11): 1 235-1 246.
DOI:10.1111/j.1461-0248.2005.00826.x |
Marzloff M P, Melbourne-Thomas J, Hamon K G, Hoshino E, Jennings S, Van Putten I E, Pecl G T. 2016. Modelling marine community responses to climate-driven species redistribution to guide monitoring and adaptive ecosystem-based management. Global Change Biology, 22(7): 2 462-2 474.
DOI:10.1111/gcb.13285 |
Mouquet N, Lagadeuc Y, Devictor V, Doyen L, Duputié A, Eveillard D, Faure D, Garnier E, Gimenez O, Huneman P, Jabot F, Jarne P, Joly D, Julliard R, Kéfi S, Kergoat G J, Lavorel S, Le Gall L, Meslin L, Morand S, Morin X, Morlon H, Pinay G, Pradel R, Schurr F M, Thuiller W, Loreau M. 2015. REVIEW: predictive ecology in a changing world. Journal of Applied Ecology, 52(5): 1 293-1 310.
DOI:10.1111/1365-2664.12482 |
Muramatsu T, Hayama Y, Kutsukake K, Maeda K, Matsumoto T, Kudo H, Fujiwara K, Usami N. 2018. Application of weighted Voronoi diagrams to analyze nucleation sites of multicrystalline silicon ingots. Journal of Crystal Growth, 499: 62-66.
DOI:10.1016/j.jcrysgro.2018.07.028 |
Murase H, Kitakado T, Hakamada T, Matsuoka K, Nishiwaki S, Naganobu M. 2013. Spatial distribution of Antarctic minke whales (Balaenoptera bonaerensis) in relation to spatial distributions of krill in the Ross Sea, Antarctica. Fisheries Oceanography, 22(3): 154-173.
DOI:10.1111/fog.12011 |
Paquin M M, Buckley T W, Hibpshman R E, Canino M F. 2014. DNA-based identification methods of prey fish from stomach contents of 12 species of eastern north pacific groundfish. Deep Sea Research Part I Oceanographic Research Papers, 85: 110-117.
DOI:10.1016/j.dsr.2013.12.002 |
Peck M A, Arvanitidis C, Butenschön M, Canu D M, Chatzinikolaou E, Cucco A, Domenici P, Fernandes J A, Gasche L, Huebert K B, Hufnagl M, Jones M C, Kempf A, Keyl F, Maar M, Mahévas S, Marchal P, Nicolas D, Pinnegar J K, Rivot E, Rochette S, Sell A F, Sinerchia M, Solidoro C, Somerfield P J, Teal L R, Travers-Trolet M, van de Wolfshaar K E. 2018. Projecting changes in the distribution and productivity of living marine resources: a critical review of the suite of modelling approaches used in the large European project VECTORS. Estuarine, Coastal and Shelf Science, 201: 40-55.
DOI:10.1016/j.ecss.2016.05.019 |
Pikitch E K, Santora C, Babcock E A, Bakun A, Bonfil R, Conover D O, Dayton P, Doukakis P, Fluharty D, Heneman B, Houde E D, Link J, Livingston P A, Mangel M, McAllister M K, Pope J, Sainsbury K J. 2004. Ecosystem-based fishery management. Science, 305(5682): 346-347.
DOI:10.1126/science.1098222 |
Richards S A. 2008. Dealing with overdispersed count data in applied ecology. Journal of Applied Ecology, 45(1): 218-227.
DOI:10.1111/j.1365-2664.2007.01377.x |
Sagarese S R, Frisk M G, Cerrato R M, Sosebee K A, Musick J A, Rago P J. 2014. Application of generalized additive models to examine ontogenetic and seasonal distributions of spiny dogfish (Squalus acanthias) in the Northeast (US) shelf large marine ecosystem. Canadian Journal of Fisheries and Aquatic Sciences, 71(6): 847-877.
DOI:10.1139/cjfas-2013-0342 |
Schmiing M, Afonso P, Tempera F, Santos R S. 2013. Predictive habitat modelling of reef fishes with contrasting trophic ecologies. Marine Ecology Progress Series, 474: 201-216.
DOI:10.3354/meps10099 |
Segurado P, Araújo M B. 2004. An evaluation of methods for modelling species distributions. Journal of Biogeography, 31(10): 1 555-1 568.
DOI:10.1111/j.1365-2699.2004.01076.x |
Smith P. 1994. Autocorrelation in the logistic regression modelling of species distributions. Global Ecology and Biogeography Letters, 4: 47-61.
|
Vezza P, Muñoz-Mas R, Martinez-Capel F, Mouton A. 2015. Random forests to evaluate biotic interactions in fish distribution models. Environmental Modelling & Software, 67: 173-183.
DOI:10.1016/j.envsoft.2015.01.005 |
Vinagre C, Fonseca V, Cabral H, Costa M J. 2006. Habitat suitability index models for the juvenile soles, Solea solea and Solea senegalensis, in the Tagus estuary: defining variables for species management. Fisheries Research, 82(1-3): 140-149.
DOI:10.1016/j.fishres.2006.07.011 |
Virgili A, Racine M, Authier M, Monestiez P, Ridoux V. 2017. Comparison of habitat models for scarcely detected species. Ecological Modelling, 346: 88-98.
DOI:10.1016/j.ecolmodel.2016.12.013 |
Wood S N. 2006. Generalized Additive Models: An Introduction with R. Chapman & Hall/CRC, Boca Raton, Florida.
|
Wu D Q, Liu J, Zhang G S, Ding W J, Wang W, Wang R Q. 2009. Incorporating spatial autocorrelation into cellular automata model: an application to the dynamics of Chinese tamarisk (Tamarix chinensis Lour.).. Ecological Modelling, 220(24): 3 490-3 498.
DOI:10.1016/j.ecolmodel.2009.03.008 |
Xiao Z T, Wu J Y, Xu B D, Zhang C L, Ren Y P, Xue Y. 2019. Uniqueness measure based on the weighted trophic field overlap of species in the food web. Ecological Indicators, 101: 640-646.
DOI:10.1016/j.ecolind.2019.01.042 |
Xu B D, Zhang C L, Xue Y, Ren Y P, Chen Y. 2015. Optimization of sampling effort for a fishery-independent survey with multiple goals. Environmental Monitoring and Assessment, 187(5): 252.
DOI:10.1007/s10661-015-4483-9 |
Xue Y, Tanaka K, Yu H M, Chen Y, Guan L S, Li Z G, Yu H Q, Xu B D, Ren Y P, Wan R. 2018. Using a new framework of two-phase generalized additive models to incorporate prey abundance in spatial distribution models of juvenile slender lizardfish in Haizhou Bay, China. Marine Biology Research, 14(5): 508-523.
DOI:10.1080/17451000.2018.1447673 |
Youcef W A, Lambert Y, Audet C. 2013. Spatial distribution of Greenland halibut Reinhardtius hippoglossoides in relation to abundance and hypoxia in the estuary and Gulf of St.Lawrence. Fisheries Oceanography, 22(1): 41-60.
DOI:10.1111/fog.12004 |
Yu W, Guo A, Zhang Y, Chen X J, Qian W G, Li Y S. 2018. Climate-induced habitat suitability variations of chub mackerel Scomber japonicus in the East China Sea. Fisheries Research, 207: 63-73.
DOI:10.1016/j.fishres.2018.06.007 |
Zerbini A N, Friday N A, Palacios D M, Waite J M, Ressler P H, Rone B K, Moore S E, Clapham P J. 2016. Baleen whale abundance and distribution in relation to environmental variables and prey density in the Eastern Bering Sea. Deep Sea Research Part II: Topical Studies in Oceanography, 134: 312-330.
DOI:10.1016/j.dsr2.2015.11.002 |
Zhang C L, Chen Y, Xu B D, Xue Y, Ren Y P. 2018a. Comparing the prediction of joint species distribution models with respect to characteristics of sampling data. Ecography, 41(11): 1 876-1 887.
DOI:10.1111/ecog.03571 |
Zhang Y L, Xue Y, Yu H M, Yu H Q, Zhang C L, Xu B D, Ren Y P. 2018b. Study on the habitat suitability of Johnius belangerii during spring in the Haizhou Bay, China. Acta Oceanologica Sinica, 40(6): 83-91.
(in Chinese with English abstract) DOI:10.3969/j.issn.0253-4193.2018.06.008 |
Zhang Y L, Yu H M, Yu H Q, Xu B D, Zhang C L, Ren Y P, Xue Y, Xu L L. 2020. Optimization of environmental variables in habitat suitability modeling for mantis shrimp Oratosquilla oratoria in the Haizhou Bay and adjacent waters. Acta Oceanologica Sinica, 39(6): 36-47.
DOI:10.1007/s13131-020-1546-8 |
Zhao J, Cao J, Tian S Q, Chen Y, Zhang S Y, Wang Z H, Zhou X J. 2014. A comparison between two GAM models in quantifying relationships of environmental variables with fish richness and diversity indices. Aquatic Ecology, 48(3): 297-312.
DOI:10.1007/s10452-014-9484-1 |
Zuur A, Ieno E N, Smith G M. 2007. Analyzing Ecological Data. Springer, New York.
|
 2021, Vol. 39
2021, Vol. 39