Institute of Oceanology, Chinese Academy of Sciences
Article Information
- CHEN Kang, WANG Weifeng, ZHU Weilin, CHEN Xiuli, WANG Huanling
- Morphological characteristics and genetic differentiation of Lutraria maxima in coast waters off southeast China
- Journal of Oceanology and Limnology, 39(4): 1387-1402
- http://dx.doi.org/10.1007/s00343-020-0172-9
Article History
- Received May. 8, 2020
- accepted in principle Jun. 24, 2020
- accepted for publication Oct. 5, 2020
2 Guangxi Key Laboratory of Aquatic Genetic Breeding and Healthy Aquaculture, Guangxi Academy of Fishery Sciences, Nanning 530021, China
The clam Lutraria maxima is a benthic bivalve with shell in faint yellow, and found in sandy habitats from the lower intertidal zone to 10-m depths. It is widely distributed in the coastal waters off the southern to southeastern China, and especially abundant in Beibu Gulf (Li et al., 2004). In recent years, this species has developed into one of the main varieties of shallow sea aquaculture especially in Guangxi and Guangdong provinces by natural seedling captivity (Pan and Su, 2007).
Morphological metric, also called the biological measurement method, is the most traditional and simple method for analyzing genetic variation. The variation of morphological traits among populations can be clarified by measuring the external morphological characteristics of the phenotypic traits. Multivariate analysis such as principal component and discriminant analysis can easily and intuitively represent the morphological differences among different populations, and clearly indicate the differentiation patterns of population structure (Frédérich et al., 2012). These methods have widely been used to distinguish different populations in some aquatic animals. Ruiz-Campos et al. (2003) study the morphological differences among different populations of wild trout (Salmonidae) in Mexico. Silva (2003) found the morphological differences between populations of sardines (Sardina pilchardus) from the northeast Atlantic Ocean and the western Mediterranean. As a kind of genetic markers, morphological markers are easily identified, simple and intuitive, but there are problems such as less number of markers, poor polymorphism, and subjective judgment of researchers. Additionally, they are often influenced by both environment and genetic factors (Murat and Aykut, 2015).
Microsatellites, also known as simple sequence repeats (SSR), or short tandem repeats (STR), refer to tandem repeat DNA sequences with 1-6 bases as a repeat core, and are distributed throughout the whole genome of an organism (Chistiakov et al., 2006). Due to high polymorphisms and convenient detection, microsatellite markers are relatively appropriate to estimate differentiation within and among populations, especially within narrow geographical range (Wolfus et al., 1997; Chareontawee et al., 2007). At present, microsatellite markers have been successfully applied to analyze genetic variation and population structure of mollusk. Genetic structure of six Sepia officinalis populations in Iberian Peninsula from the Atlantic to the Mediterranean was analyzed, and a higher degree of genetic variation in these populations was found with seven microsatellite markers (Pérez-Losada et al., 2002). It has been reported that 21 Mizuhopecten yessoensis populations from Japan (Hokkaido and Honshu) and Russia show high genetic diversity with 0.701 1-0.762 2 of average expected heterozygosity (Sato et al., 2005). Kelly and Rhymer (2005) analyzed the population genetic structure of Lampsilis cariosa in North America based on microsatellite markers, and pointed out that the biological and biogeographic history of the host fish had a potential impact on the population genetic structure of L. cariosa.
Previous researches about L. maxima focused mainly on nutrition (Pan et al., 2007), reproductive characteristics (Li et al., 2004), and artificial breeding (Su et al., 2009). However, there are few reports on the evaluation of L. maxima germplasm resources. Only a single study compared the genetic differences among three populations of L. siebaldii and one population of L. maxima using morphological features and random amplified polymorphic DNA (RAPD) techniques (Li et al., 2011a). Therefore, to identify microsatellite markers with high polymorphism based on transcriptome data, and analyze the genetic diversity and genetic variation of five populations of L. maxima, combined with morphological features.
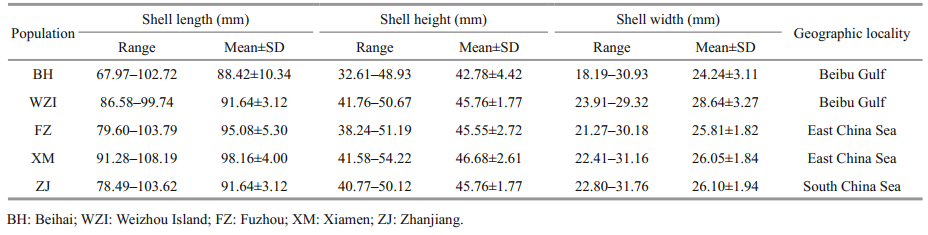
2 MATERIAL AND METHOD 2.1 Sample collectionLutraria maxima (Table 1) were sampled from five different populations along the east and south China coast, namely, Fuzhou (FZ, n=40) and Xiamen (XM, n=40) in Fujian Province, Zhanjiang (ZJ, n=40) in Guangdong Province, Weizhou Island (WZI, n=40) and Beihai (BH, n=40) in Guangxi Autonomous Region in summer 2018, and all samples were obtained from fishermen on local fishing wharfs (Fig. 1). Muscle tissues of each individual were preserved in 95% ethanol and stored at -80 ℃ for DNA extraction.
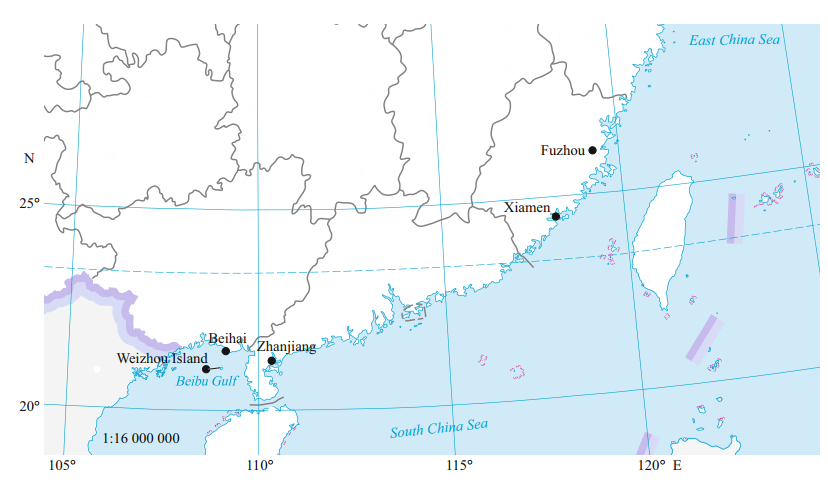
|
| Fig.1 Sampling locations of five L. maxima populations The map was adapted from the Ministry of Natural Resources, China (http://bzdt.ch.mnr.gov.cn/index.html), map review No. GS(2019)1671. Locations are labeled by dots (•), BH (Beihai): 21°48'N, 109°12'E; WZI (Weizhou Island): 21°04'N, 109°12'E; ZJ (Zhanjiang): 21°27'N, 110°37'E; XM (Xiamen): 24°48'N, 118°09'E; FZ (Fuzhou): 26°08'N, 119°30'E. |
This study was approved by the Institutional Animal Care and Use Committee (IACUC) of Huazhong Agricultural University, Wuhan, China, and conducted in accordance with the ethical standards and according to the regarding national and international guidelines.
2.2 Morphometric analysisEleven shell characters were measured with Vernier calipers for each individual as previously described (Li et al., 2011a). To eliminate the influence of different shellfish sizes on the morphological metics, a previous method (Gardner and Thompson, 1999) was referred prior to analysis, and the values of 10 morphometric (SH, SW, DUA, HAR, WAR, HPR, WPR, WT, DTL, and DPM) indices except for shell length (SL) were log10-transformed before being divided by log10-transformed total shell length to correct for size dependent variation. Ten morphological proportion traits were analyzed using SPSS19.0 for discriminant analysis, principal component analysis (PCA), and cluster analysis. The principal components were extracted according to the variance-covariance matrix and determined as the principal components whose eigenvalues are > 1, and finally the dispersion point graph was drawn according to the load values of PCA1 and PCA2. The Euclidean distance coefficient was constructed by using the clustering method of intergroup connection (Yang et al., 2019).
2.3 Genetic diversity analysis based on microsatellite markers 2.3.1 Transcriptome sequencingMuscle tissues from five L. maxima (from BH population) were mixed in a pool, and frequently frozen in liquid nitrogen. Total RNAs were extracted from muscle tissues using the TRIzol regent according to the manufacturer's instructions. RNA-seq transcriptome library was constructed and sequenced with 150-bp paired-end reads using Illumina HiSeq 2500. Transcriptome assembly was performed with Trinity software (Grabherr et al., 2011).
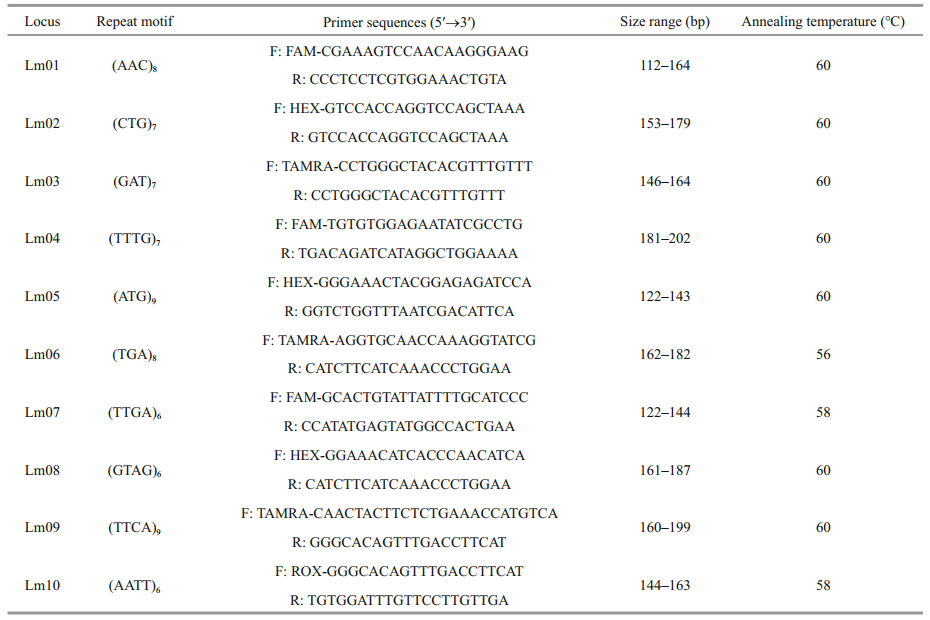
2.3.2 Identification of microsatellite markersMicrosatellite markers were identified in the transcriptome data using the MISA version 1.0 (http://pgrc.ipk-gatersleben.de/misa/misa.html) with the search criteria of repeat unit, single nucleotide repeat times ≥10, dinucleotide repeat times ≥6, three, four, five, six nucleotide repeat times ≥5, compound microsatellite marker site base interval ≤100 bp. Primer pairs for each microsatellite marker were designed using Primer 3 software (Untergasser et al., 2012).
2.3.3 DNA extraction and microsatellite marker genotypingGenomic DNA was extracted from muscle tissues of all 200 fresh specimens as per the protocol described by Li et al. (2006). A total of 50 primer pairs were designed and amplified in 10 randomly selected individuals to obtain the appropriate microsatellite primers. PCR was performed in a 10-μL reaction mixture that included 0.3 μmol/L each primer, 0.15 mmol/L each dNTP, 1-μL 1×PCR buffer, 0.15-μL Taq DNA polymerase, 0.5-μL template DNA (100 ng/μL). Thermal cycling conditions for each locus were as follows: 3 min at 94℃, followed by 32 cycles of 94 ℃ for 30 s, annealing temperature (Table 2) for 30 s, and 72 ℃ for 25 s, and a final extension of 72 ℃ for 10 min. The PCR products were examined by electrophoresis on 8% non-denaturing polyacrylamide gelelectrophoresis (PAGE) gels and silver stained as previously described (Zhang et al., 2013).
Finally, 10 highly polymorphic microsatellite marker primers (Table 2) were obtained and labeled in fluorescence. PCR amplification was performed in the five populations of L. maxima. The PCR products were detected by capillary electrophoresis using an ABI3730 sequencer (Applied Biosystems, USA), and genotypes were analyzed by Genemarker v1.91 software (Georgia Institute of Technology, Atlanta, Georgia, USA).
2.3.4 Statistical analysisTo analyze the genetic diversity, software Cervus v3.0 (Kalinowski et al., 2007) was used to estimate the number of alleles (Na) and effective alleles (Ne), expected (He) and observed (Ho) heterozygosity, Wright's F statistics (FIS) and polymorphism information content (PIC). GenAlEx 6.5 (Peakall and Smouse, 2012) was employed to calculate Nei's genetic distance (DA), and analysis of molecular variance (AMOVA). Gene flow was calculated through the FST between the two populations, and the calculation formula was as follows: gene flow (NM)=(1-FST)/(4FST) (Wright, 1965). The presence of null alleles was detected by MICRO-CHECKER (van Oosterhout et al., 2004), and frequency of the null alleles was estimated using Brookfield (1996) equation, r=(He-Ho)/(1+He). Software Structure 2.3.4 (Falush et al., 2003) was used for population genetic structure analysis, of which the parameter "Length of Burnin Period" was set to 10 000, "Number of MCMC Reps after Burnin" was set to 50 000, K was 1-7 and run 20 times for each value. Combined with the online tool, Structure Harvester (http://taylor0.biology.ucla.edu/structureHarvester/), the best K value was determined (Evanno et al., 2005). CLUMPP 1.1.2 (Jakobsson and Rosenberg, 2007) was used to repeatedly sample and analyze the structure Harvester's results. Distruct 1.1 (Rosenberg, 2004) was finally used to draw the structure graph of Structure. Using MEGA 7.0 (Kumar et al., 2016), the unrooted unweighted pair-population method with arithmetic means (UPGMA) dendrogram was used to further verify the genetic grouping in Structure based on the DA. Arlequin v3.0 (Excoffier et al., 2005) was employed to calculate pairwise FST values, and their significance by bootstrapping analysis (1 000 replicates) was tested for evaluating genetic differentiation among populations. Deviation from Hardy-Weinberg equilibrium (HWE) for each locus-population combination was assessed.
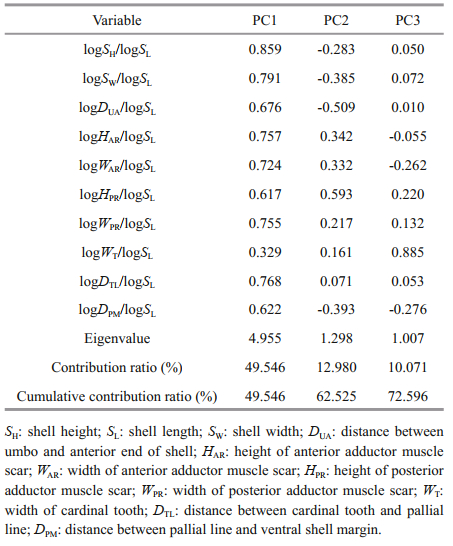
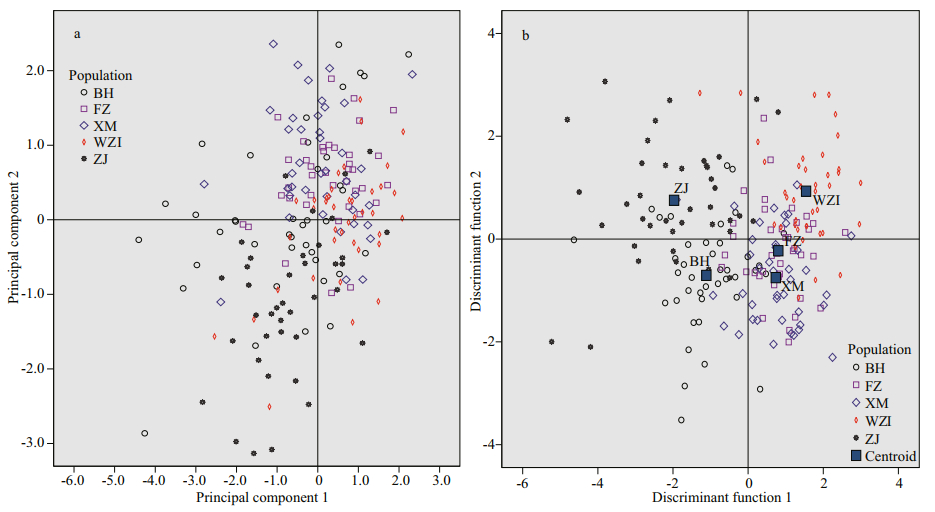
3 RESULT 3.1 Geometric morphometrics 3.1.1 Principal component analysisTo determine the principal component factors affecting the morphological differences of L. maxima in different geographical populations, principal component analysis of 10 morphological proportion traits was carried out, and 3 principal components (PC1-PC3) were obtained (Table 3). The first three principal components accounted for 49.546%, 12.980%, and 10.071% of the total variation, respectively, and the cumulative contribution rate was 72.596%. In the first principal component, the contribution rate of SH/SL and SW/SL was the largest, mainly reflecting differences of shell height and width. PC2 mainly showed the indicators of DUA/SL and HPR/SL, which mainly reflected the characteristics of the distance between umbo and anterior end of shell, and height of posterior adductor muscle scar. PC3 mainly reflected the width of cardinal tooth (WT/SL) and distance between pallial line and ventral shell margin (DPM/SL). The scatter diagram of the principal component PC1 and PC2 of the five populations showed that there were more interlaced overlapping regions between the FZ and XM populations, suggesting that the genetic relationship of the two populations was the closest (Fig. 2a).
|
|
| Fig.2 Distribution of PC1 and PC2 (a) and discriminant function analysis (b) for five populations of L. maxima based on morphological indicators |
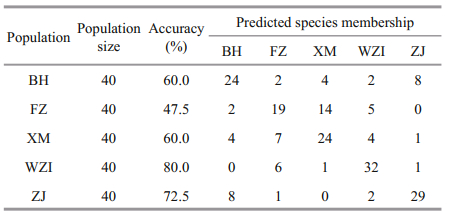
The results show that the comprehensive discrimination rate of the five populations was 64.0% (Table 4). Discriminant analysis of different populations showed that BH and ZJ populations were generally located on the left side of the scatter plot, and the other three populations were mostly distributed on the right side of the scatter plot. In addition, the XM and FZ populations clearly occupied a significant overlapping region (Fig. 2b). Geographically, the distance between the two populations was the closest, although some individuals were confused.
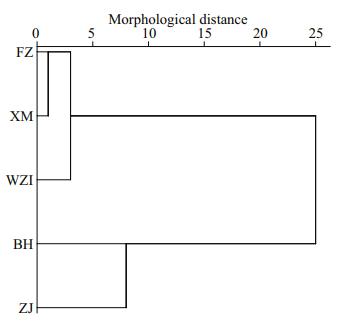
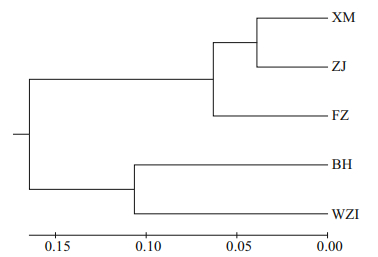
The Euclidean distance tree was obtained by the method of intergroup connection based on mean values of the 10 morphological proportion traits, and showed that these five populations of L. maxima were divided into two major branches (Fig. 3). The BH and ZJ populations were clustered first, and the XM, FZ and WZI populations were clustered into another one.
|
| Fig.3 Cluster analysis of five populations of L. maxima in the Euclidean distance based on morphological indicators |
To explore polymorphism in transcriptome-derived microsatellite makers and further understand the genetic diversity and structure of different populations, RNA-seq from muscle tissues was performed. In total, 63 047 transcripts were assembled using MISA software, and 2 561 microsatellite markers were found. Among these microsatellite markers, the most abundant motifs were mononucleotide (47.87%) repeats, followed by trinucleotide (26.63%), and dinucleotide (18.04%) repeat types. The tetranucleotide, pentanucleotide, and complex nucleotide repeat types were all less than 5%. A total of 50 primer pairs were designed, but only 10 pairs of primers were well amplified and highly polymorphic (Table 2).
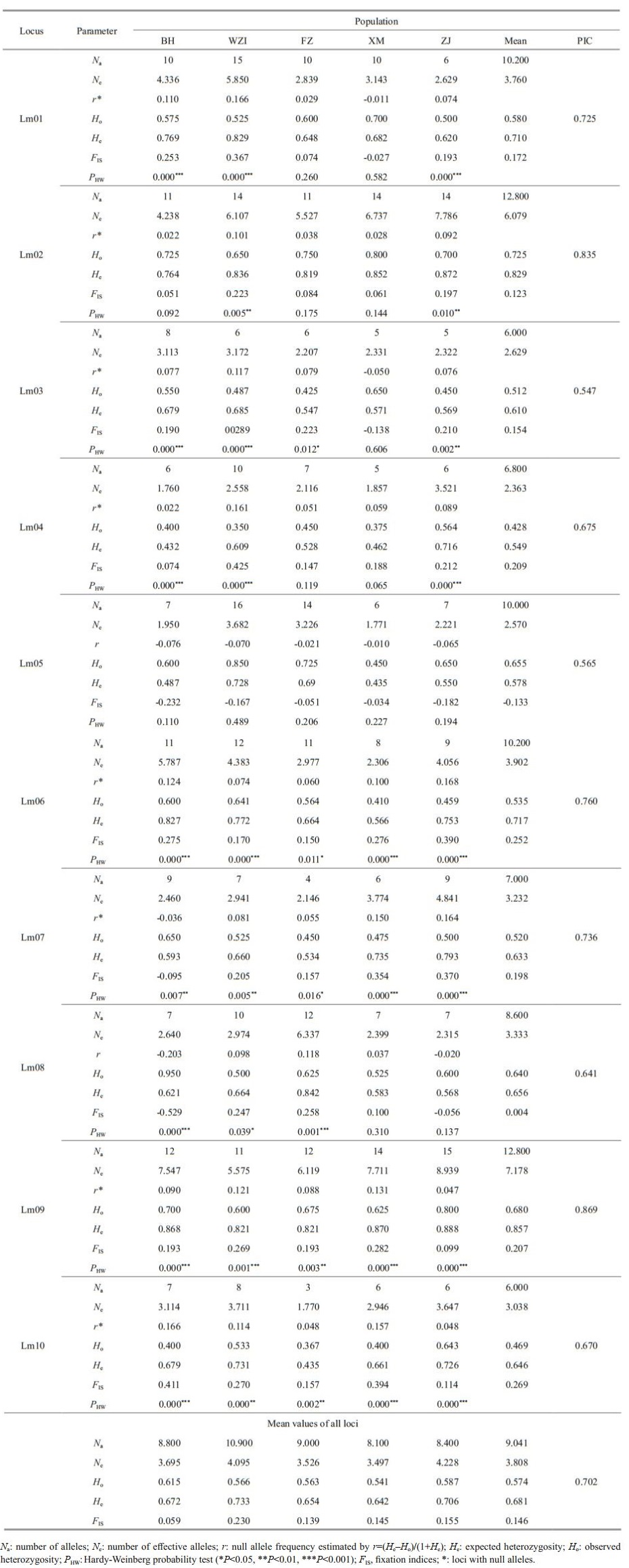
3.3 Genetic diversity and structure analysis 3.3.1 Polymorphisms of microsatellite lociThe average number of alleles in 10 microsatellite loci with polymorphism per locus was 9.041 (ranging from 6.000 to 12.800), PIC values were more than 0.5 at all loci (mean of 0.702). The expected heterozygosity (He) ranged from 0.549 to 0.857, and the mean observed heterozygosity (Ho) ranged from 0.428 to 0.725, indicating that these microsatellite loci had high polymorphism (Table 5). Hardy-Weinberg equilibrium (HWE) tests were performed and showed 35 tests with significant deviation from HWE in all five populations. The existence of null alleles was the main reason for deviation from HWE. Microchecker analysis showed that there were 8 microsatellite loci with null alleles (Table 5). The value of FIS was greater than 0 except that the Lm05 locus was negative, indicating that there was a loss of heterozygotes at these loci, which may also be another important reason for the deviation from HWE.
|
Fluorescent labeled microsatellite markers were synthesized and used to assess the genetic diversity of five geographical populations of L. maxima. Data for all parameters of genetic diversity for 200 individuals from the five populations are shown in Table 5. The average number of alleles (Na) per locus varied from 8.100 to 10.900, and the average number of effective alleles (Ne) per locus varied from 3.497 to 4.228. The five populations examined here exhibited high levels of heterozygosity. The expected heterozygosity (He) ranged from 0.642 to 0.733, which the highest mean He occurred in the WZI population, and the lowest mean He was in the XM population. The lowest mean observed heterozygosity (Ho) was in the XM population (0.541), whereas the highest in the BH population (0.615). At the species level of L. maxima, the average Na, Ne, Ho, and He at per locus were 9.041, 3.808, 0.574, and 0.681, respectively. In general, the diversity level of L. maxima population in China was relatively high.
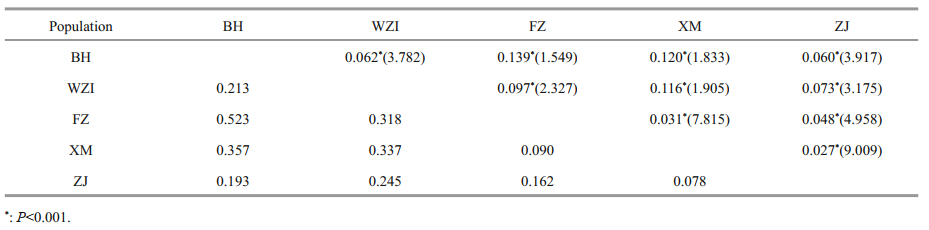
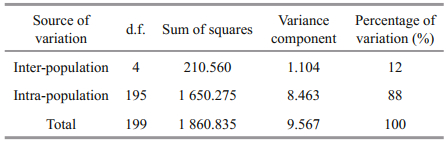
3.3.3 Genetic divergence and structureAnalysis of all pairwise FST statistics showed significant differences (P < 0.001) among the five populations. The highest divergence was between the BH and FZ populations (FST=0.139), while the lowest divergence was between the XM and ZJ populations (FST=0.027). The gene flow of five populations varied between 1.549 and 9.009 (Table 6). The analysis of molecular variance (AMOVA) showed that the genetic variation among and within populations was 12% and 88%, respectively (Table 7), indicating that genetic variation mainly came from within populations, but different populations still had significant genetic differentiation.
|
|
Structure 2.3.4 was used to execute the hypothetical K value of 1-7. The results were uploaded to StructureHarvester to determine the best K value, and K and ΔK line chart and K and LnP(D) scatter plot were showed (Fig. 4a & b). According to the trend analysis of K value, when K=2, ΔK had a maximum value, and followed by K=3. Based on the value of K=2, the population structure analysis showed that the five populations could be divided into two clusters, BH+WZI and FZ+XM+ZJ, at a certain degree of gene exchange between each population (Fig. 4c). When K=3, individuals from BH and WZI populations weresignificantly different from each other, and the other three populations were similar to some extents, in which XM and ZJ was the more similar (Fig. 4d).

|
| Fig.4 Genetic structure results of five populations of L. maxima based on ten microsatellite loci a. the relation between the likelihood L (K) values and ΔK values; b. the relation between the value of logarithmic likelihood function and the value of K; c. Bayesian clustering of the optimal grouping K=2 for five populations; d. Bayesian clustering of the optimal grouping K=3 for five populations. |
Based on the genetic distance (DA), two major branches were showed in the UPGMA tree (Fig. 5), which was consistent with the result of the Structure analysis. The two populations (BH, WZI) from Beibu Gulf were clustered into one branch, the other three populations (FZ, XM, and ZJ) from the southeastern coastal zones of China region were clustered in the middle of the tree, and among them, the XM and ZJ populations were the first to be gathered together.
|
| Fig.5 UPMGA dendrogram of L. maxima by Nei's genetic distance based on ten microsatellite loci |
Long-term geographical isolation and adaptability to different environments result in certain reproductive isolation among different geographical populations of the same species due to lack of genetic exchange, thus forming certain differences in their morphology, physiology, and genetics (Li et al., 2011b). To determine the management unit of a natural population, it is important to evaluate the diversity and structure of the population. Therefore, in the present study, L. maxima from five populations were collected and used to analyze the population structure and diversity based on both morphology and microsatellite makers.
4.1 Microsatellite screening strategySince Tautz and Renz (1984) reported microsatellite sequences obtained from various organisms, microsatellite markers have become one of the widely used genetic markers (Zhao et al., 2019). We used high-throughput sequencing to screen transcriptome-derived microsatellite makers, which features fast, high efficiency, strong transferability, correlation with potential genes, and large amount of data, thus accelerating the efficiency of microsatellite maker development (Ma et al., 2014; Ueno et al., 2015). With the rapid development of sequencing technology and bioinformatics, more and more transcriptome-derived microsatellite makers have been developed for aquatic animals, such as Ictalurus punctatus (Serapion et al., 2004), Carassius auratus gibelio (Yue et al., 2004), and Argopecten irradians (Zhan et al., 2005), etc. In this study, 50 primers were designed and synthesized, and verified and analyzed in the individual genomic DNA of L. maxima. Among them, 28 pairs had amplification products and 10 microsatellite markers with high polymorphism were obtained, accounting for 35.7% of the total. Microsatellite marker polymorphisms are important to evaluate genetic diversity. Compared with the genome derived microsatellite markers, the transcriptome-derived microsatellite markers have low polymorphism due to stricter evolutionary restrictions (Ellis and Burke, 2007), but some studies show different results (Yue et al., 2004). Polymorphic information content (PIC) is an important indicator of genetic diversity level (Botstein et al., 1980), and all loci in five populations were highly polymorphic due to PIC > 0.5, suggesting higher polymorphisms and strong allelic variation in L. maxima based on transcriptome-derived microsatellite markers. Therefore, the 10 pairs of microsatellite primers obtained can be suitable for evaluating the genetic diversity and variation among populations.
4.2 Genetic diversity within populationHeterozygosity, also known as gene diversity, is generally considered the optimal parameter for assessing population variation at the genetic level (Nei et al., 1975). If the average heterozygosity of one population is higher than 0.5, it indicates that the population is not selected by high intensity and has rich genetic diversity, and vice versa (Li et al., 2010). Therefore, higher heterozygosity contributes to the richer genetic diversity of the species and the stronger adaptability to environments (Qin et al., 2013). In this study, the mean He and Ho values in five L. maxima populations range from 0.642 to 0.733 and 0.541 to 0.615, and the average He (0.681) was higher than Ho (0.574), indicating the loss of heterozygotes. The average effective allele number of five populations was high, suggesting that L. maxima has a high genetic variation, which is conducive to the protection of germplasm resources, whereas populations with a low degree of genetic differentiation should be paid attention to strengthen protection measures (Qin et al., 2013). The mean Ne value in the five L. maxima populations ranged from 3.497 to 4.228, further confirming the high genetic diversity of this species. High genetic diversity seems to be a common feature of marine bivalves, such as Pinctada maxima (Smith et al., 2003), Scapharca broughtonii (An and Park, 2005) and Mizuhopecten yessoensis (Sato et al., 2005).
Moreover, in all populations, FIS was positive, suggesting the presence of null alleles (Pemberton et al., 1995). Microchecker software detected eight out of 10 loci with null alleles. This phenomenon has been reported on other shellfish, such as Crassostrea gigas showing null alleles in 51.9% (41/79) ofmicrosatellite loci (Li et al., 2003a). In addition, null alleles lead to the loss of heterozygotes (Wanna et al., 2004), our results also showed that the five geographical populations deviated significantly from HWE at most of the detected loci. Selkoe and Toonen (2006) believe that loci with deviation from HWE would not have a significant impact on the subsequent analysis and should not be discarded. Some studies reveal that the loss of heterozygotes in aquatic animals is one factor to result in deviation from HWE (Evans et al., 2004). Therefore, it is speculated that there may be serious heterozygote deficiencies in L. maxima populations, and the mating between individuals is probably not random. There may be several possible explanations for the deviation from HWE expectations. Firstly, human activities, such as overfishing, the introgression of cultured stocks and other reasons will lead to genetic balance deviation of wild population, eventually, loss of diversity and population viability (So et al., 2006). Secondly, in recent years, the rapid development of coastal areas along the southeast of China has led to the degradation of habitats and the reduction of effective populations, which may lead to certain inbreeding among individuals. Thirdly, it has been found that marine bivalves with longer larval stages have the higher dispersibility (Arnaud et al., 2000), and the migration and diffusion of the larvae will probably promote the gene exchange among geographical populations, and thus the migration of animals will promote the gene exchange among different geographical populations. In addition, studies have shown that inbreeding, intra-population structure (Wahlund effect), and sampling bias are the causes of heterozygote deficiencies (Borsa et al., 1991; Gallardo et al., 1998; Wang et al., 2007; Cannas et al., 2012).
4.3 Genetic divergence and cluster among populationsPrincipal component analysis is an effective multivariate analysis method (Gibson et al., 1984; Voss et al., 1990). This study showed that the cumulative contribution rate of the first three principal components was 72.596%, which was lower than 85% (Eisenhour, 1999), indicating that different populations of L. maxima could not be completely correctly distinguished by morphological differences with several selected parameters, combined with discriminant analysis with 64% of the overall discriminant accuracy rate. The XM and FZ populations occupied the same overlap based on the geographic principal component analysis and the discriminant analysis, which indicated that the morphological characteristics between the two populations are hardly distinguishable.
Based on the morphological indicators, the Euclidean distance tree was constructed, showing that the BH and ZJ populations were first clustered together, and the XM, FZ, and WZI populations were clustered into one, which does not match the geographical locations of populations. BH and ZJ are separated by the Leizhou Peninsula and the Qiongzhou Strait, and WZI and BH are apart by a narrow sea, so there should be more genetic exchange between the WZI and BH populations. In fact, morphological differences are determined by both genetic and environmental factors, and sometimes environmental factors may be the dominant factors. Laudien et al. (2003) concluded that the morphological differences of Donax serra are determined by the characteristics of the geographical environment. Therefore, this study further explored relationships within populations and among populations based on microsatellite markers.
UPGMA analysis showed that the five populations were divided into two main clusters, Beibu Gulf group (Beihai and Weizhou Island) and the Southeast China Sea group (Zhanjiang, Xiamen, and Fuzhou), the reason may be that the current in Qiongzhou Strait mainly flows from east to west in all seasons, which definitely prevent the L. maxima from expanding eastwards. However, the ZJ population is geographically the closest to the BH and WZI populations, but the ZJ population is the first to cluster with the XM and FZ populations. Because the L. maxima has a planktonic larval stage (Li et al., 2003b), we speculate that in summer, China Coastal Current flows northward, and has been flowing from the South China Sea to the East China Sea (Ni et al., 2014), thus strengthening the connection between the seas. The Structure results show that when K=2, the ΔK value was the largest, which further corroborated the UPGMA dendrogram results. At the same time, we found that the two clusters had a low degree of mixing, and the different clusters were relatively concentrated in Bayesian clustering In general, the genetic structure of L. maxima was related to the geographical locations sampled, which the same water area promotes frequent gene exchanges, resulting in similar genetic structures.
Based on the pairwise FST values and the Nei's genetic distance, it was found that there was significant moderate genetic differentiation between the BH and WZI populations located in Beibu Gulf, and the FZ and XM populations located in East China Sea (FST=0.097-0.139). During the Quaternary glacial period, the sea level in the East China Sea fell, promoting the formation of a land bridge between the mainland and Taiwan of China, and separating the East China Sea from the South China Sea and the Pacific Ocean (Ni et al., 2014). And Beibu Gulf is a semi-closed bay, which probably promotes L. maxima population to form independent refuges in the two seas, and blocks the genetic exchange of species between Beibu Gulf group (BH and WZI) and Southeast China Sea group (FZ and XM). Additionally, a low level of genetic differentiation (FST=0.027- 0.048) was found among the FZ, XM, and ZJ populations located along the southeast coast of China with the distance of about 260-1 254 km. Guillemaud et al. (2003) believe that the difference of genetic differentiation is not obvious when the geographical distance is less than 60 km, but it is obvious when the geographical distance is 150-200 km, indicating that geographical distance may not be the main influencing factor. In the later period of the glacier, the global temperature rises, all marginal sea creatures recontact (Huang et al., 2015), thus the gene exchange results in the genetic differentiation of the species due to geographic isolation eliminated (Coope, 1979; Liu et al., 2011). This may be one reason for the formation of a cluster in the Southeast China Sea group (FZ, XM, and ZJ) and low genetic differentiation. Similarly, low levels of genetic differentiation are also detected in other marine bivalves, such as Crassostrea rhizophorae (Ignacio et al., 2000), Cerastoderma glaucum (Mariani et al., 2002) and Pecten jacobaeus (Ríos et al., 2002). Additionally, it has been reported that genetic variation of Chlamys farreri is closely related to currents (Zhan et al., 2009). The effect of ocean currents on the diffusion ability of plankton larvae is also significant due to the large geographic distance span of the East China Sea and the extremely weak movement ability of plankton species (Lee and Boulding, 2009). Therefore, genetic differentiation among the FZ, XM, and ZJ populations was probably affected by the warm sea current in southeast China (Lambeck et al., 2002) since the larvae of L. maxima are dispersed in a relatively large area during the planktonic life stage of about 12 days (Li et al., 2003b). Of course, the effect of human activities should be also considered.
AMOVA revealed that the genetic variation among populations was 12%, indicating that each population had significant genetic differentiation. Studies have shown that geographic differences may lead to differences in genetic structure and genetic diversity of marine shellfish populations (Kong et al., 2007). Some studies have also shown that environmental stress has a large impact on the population genetic structure of shellfish (Geist and Kuehn, 2005). Environmental differences in water temperature and salinity, etc. between Beibu Gulf group and Southeast China Sea group might also be one of the causes of genetic differentiation.
5 CONCLUSIONThis study was the first attempt to elucidate the genetic characteristics of L. maxima populations in China. Ten polymorphic transcriptome-derived microsatellite markers combined with morphology were used to assess the genetic diversity and genetic differentiation among wild L. maxima populations in China. The genetic diversity level of the L. maxima populations in the southeastern coast of China remained relatively high, and moderate genetic differentiation has occurred. The five L. maxima populations exhibited high genetic diversity as indicated by the high Ne, He, and Ho, indicating the germplasm resources were in good condition. Moderate levels of genetic differentiation were detected between BH and WZI populations located in the Beibu Gulf, and FZ, XM, and ZJ populations located in the East China Sea, which gain support also by UPGMA tree and the Structure analysis. However, with the continuous development and utilization of fishery resources, the scope of fishing is gradually expanding, and the fishery resources of wild species show a trend of gradual decline. Therefore, it is necessary to protect them against further decline in genetic diversity. Our results suggested that populations from the two regions of L. maxima should be considered as separate units in fishery management, which will provide certain references for establishment of protection strategy of L. maxima.
6 DATA AVAILABILITY STATEMENTData are available on request from the authors.
7 CONFLICT OF INTERESTThe authors declare no conflict of interest to each other.
An H Y, Park J Y. 2005. Ten new highly polymorphic microsatellite loci in the blood clam Scapharca broughtonii. Molecular Ecology Notes, 5(4): 896-898.
DOI:10.1111/j.1471-8286.2005.01104.x |
Arnaud S, Monteforte M, Galtier N, Bonhomme F, Blanc F. 2000. Population structure and genetic variability of pearl oyster Pinctada mazatlanica along Pacific coasts from Mexico to Panama. Conservation Genetics, 1(4): 299-308.
DOI:10.1023/a:1011575722481 |
Borsa P, Zainuri M, Delay B. 1991. Heterozygote deficiency and population structure in the bivalve Ruditapes decussatus. Heredity, 66(1): 1-8.
DOI:10.1038/hdy.1991.1 |
Botstein D, White R L, Skolnick M, Davis R W. 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32(3): 314-331.
|
Brookfield J F Y. 1996. A simple new method for estimating null allele frequency from heterozygote deficiency. Molecular Ecology, 5(3): 453-455.
DOI:10.1111/j.1365-294X.1996.tb00336.x |
Cannas R, Sacco F, Follesa M C, Sabatini A, Arculeo M, Lo Brutto S, Maggio T, Deiana A M, Cau A. 2012. Genetic variability of the blue and red shrimp Aristeus antennatus in the Western Mediterranean Sea inferred by DNA microsatellite loci. Marine Ecology, 33(3): 350-363.
DOI:10.1111/j.1439-0485.2011.00504.x |
Chareontawee K, Poompuang S, Na-Nakorn U, Kamonrat W. 2007. Genetic diversity of hatchery stocks of giant freshwater prawn (Macrobrachium rosenbergii) in Thailand. Aquaculture, 271(1-4): 121-129.
DOI:10.1016/j.aquaculture.2007.07.001 |
Chistiakov D A, Hellemans B, Volckaert F A M. 2006. Microsatellites and their genomic distribution, evolution, function and applications: a review with special reference to fish genetics. Aquaculture, 255(1-4): 1-29.
DOI:10.1016/j.aquaculture.2005.11.031 |
Coope G R. 1979. Late Cenozoic Fossil Coleoptera: evolution, biogeography, and ecology. Annual Review of Ecology and Systematics, 10(1): 247-267.
DOI:10.1146/annurev.es.10.110179.001335 |
Eisenhour D J. 1999. Systematics of Macrhybopsis tetranema (Cypriniformes: Cyprinidae). Copeia, 1999(4): 969-980.
DOI:10.2307/1447972 |
Ellis J R, Burke J M. 2007. EST-SSRs as a resource for population genetic analyses. Heredity, 99(2): 125-132.
DOI:10.1038/sj.hdy.6801001 |
Evanno G, Regnaut S, Goudet J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology, 14(8): 2611-2620.
DOI:10.1111/j.1365-294X.2005.02553.x |
Evans B, Bartlett J, Sweijd N, Cook P, Elliott N G. 2004. Loss of genetic variation at microsatellite loci in hatchery produced abalone in Australia (Haliotis rubra) and South Africa (Haliotis midae). Aquaculture, 233(1-4): 109-127.
DOI:10.1016/j.aquaculture.2003.09.037 |
Excoffier L, Laval G, Schneider S. 2005. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online, 1: 47-50.
|
Falush D, Stephens M, Pritchard J K. 2003. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics, 164(4): 1567-1587.
DOI:10.1093/genetics/164.4.1567 |
Frédérich B, Liu S Y V, Dai C F. 2012. Morphological and genetic divergences in a coral reef Damselfish, Pomacentrus coelestis. Evolutionary Biology, 39(3): 359-370.
DOI:10.1007/s11692-011-9158-z |
Gallardo M H, Peñaloza L, Clasing E. 1998. Gene flow and allozymic population structure in the clam Venus antiqua (King of Broderip), (Bivalvia, Veneriidae) from Southern Chile. Journal of Experimental Marine Biology and Ecology, 230(2): 193-205.
DOI:10.1016/s0022-0981(98)00069-0 |
Gardner J P A, Thompson R J. 1999. High levels of shared allozyme polymorphism among strongly differentiated congeneric clams of the genus Astarte (Bivalvia: Mollusca). Heredity, 82(1): 89-99.
DOI:10.1038/sj.hdy.6884590 |
Geist J, Kuehn R. 2005. Genetic diversity and differentiation of central European freshwater pearl mussel (Margaritifera margaritifera L.) populations: implications forconservation and management. Molecular Ecology, 14(2): 425-439.
DOI:10.1111/j.1365-294x.2004.02420.x |
Gibson A R, Baker A J, Moeed A. 1984. Morphometric variation in introduced populations of the common myna (Acridotheres tristis): an application of the jackknife to principal component analysis. Systematic Biology, 33(4): 408-421.
DOI:10.1093/sysbio/33.4.408 |
Grabherr M G, Haas B J, Yassour M, Levin J Z, Thompson D A, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q D, Chen Z H, Mauceli E, Hacohen N, Gnirke A, Rhind N, Di Palma F, Birren B W, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A. 2011. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnology, 29(7): 644-652.
DOI:10.1038/nbt.1883 |
Guillemaud T, Mieuzet L, Simon J C. 2003. Spatial and temporal genetic variability in French populations of the peach-potato aphid, Myzus persicae. Heredity, 91(2): 143-152.
DOI:10.1038/sj.hdy.6800292 |
Huang D W, Licuanan W Y, Hoeksema B W, Chen C A, Ang P O, Huang H, Lane D J W, Vo S T, Waheed Z, Affendi Y A, Yeemin T, Chou L M. 2015. Extraordinary diversity of reef corals in the South China Sea. Marine Biodiversity, 45(2): 157-168.
DOI:10.1007/s12526-014-0236-1 |
Ignacio B L, Absher T M, Lazoski C, Solé-Cava A M. 2000. Genetic evidence of the presence of two species of Crassostrea (Bivalvia: Ostreidae) on the coast of Brazil. Marine Biology, 136(6): 987-991.
DOI:10.1007/s002270000252 |
Jakobsson M, Rosenberg N A. 2007. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics, 23(14): 1801-1806.
DOI:10.1093/bioinformatics/btm233 |
Kalinowski S T, Taper M L, Marshall T C. 2007. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Molecular Ecology, 16(5): 1099-1106.
DOI:10.1111/j.1365-294x.2007.03089.x |
Kelly M W, Rhymer J M. 2005. Population genetic structure of a rare unionid (Lampsilis cariosa) in a recently glaciated landscape. Conservation Genetics, 6(5): 789-802.
DOI:10.1007/s10592-005-9037-1 |
Kong L F, Li Q, Qiu Z X. 2007. Genetic and morphological differentiation in the clam Coelomactra antiquata (Bivalvia: Veneroida) along the coast of China. Journal of Experimental Marine Biology and Ecology, 343(1): 110-117.
DOI:10.1016/j.jembe.2006.12.003 |
Kumar S, Stecher G, Tamura K. 2016. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33(7): 1870-1874.
DOI:10.1093/molbev/msw054 |
Lambeck K, Esat T M, Potter E K. 2002. Links between climate and sea levels for the past three million years. Nature, 419(6903): 199-206.
DOI:10.1038/nature01089 |
Laudien J, Flint N S, van der Bank F H, Brey T. 2003. Genetic and morphological variation in four populations of the surf clam Donax serra (Röding) from southern African sandy beaches. Biochemical Systematics and Ecology, 31(7): 751-772.
DOI:10.1016/s0305-1978(02)00252-1 |
Lee H J, Boulding E G. 2009. Spatial and temporal population genetic structure of four northeastern Pacific littorinid gastropods: the effect of mode of larval development on variation at one mitochondrial and two nuclear DNA markers. Molecular Ecology, 18(10): 2165-2184.
DOI:10.1111/j.1365-294X.2009.04169.x |
Li B, He J F, Ou X L, Su X J, Pan Y, Luo F G, Zheng H F, Qin Z B. 2011a. Morphological and genetic diversity analysis among three populations of clam Lutraria siebaldii in coastal Guangxi and Guangdong provinces. Journal of Dalian Ocean University, 26(5): 414-421.
|
Li G, Hubert S, Bucklin K, Ribes V, Hedgecock D. 2003a. Characterization of 79 microsatellite DNA markers in the Pacific oyster Crassostrea gigas. Molecular Ecology Notes, 3(2): 228-232.
DOI:10.1046/j.1471-8286.2003.00406.x |
Li Q Z, Chen R F, Tong W P, Jiang Y. 2004. Influence of salinity upon the embryogenesis of Lutraria maxima Janas. Journal of Guangxi Academy of Sciences, 20(1): 33-34.
(in Chinese with English abstract) |
Li Q Z, Tong W P, Su Q, Liu W N. 2003b. The morphogenesis of the Lutraria maxima's embryo, larvae and spat. Guangxi Sciences, 10(4): 296-299.
|
Li Q, Yu H, Yu R. 2006. Genetic variability assessed by microsatellites in cultured populations of the Pacific oyster (Crassostrea gigas) in China. Aquaculture, 254(1-4): 95-102.
|
Li S F, Zhao Y, Fan W J, Cai W Q, Xu Y F. 2011b. Possible genetic reproductive isolation between two tilapiine genera and species: Oreochromis niloticus and Sarotherodon melanotheron. Zoological Research, 32(5): 521-527.
DOI:10.3724/SP.J.1141.2011.05521 |
Li Z M, Xie L, Ye F L, Chen G L. 2010. SSR analysis of three species from primary parent and their first generation of Litopenaeus vannamei. Agricultural Science and Technology, 11(3): 57-61.
|
Liu J, Li Q, Kong L F, Zheng X D. 2011. Cryptic diversity in the pen shell Atrina pectinata (Bivalvia: Pinnidae): high divergence and hybridization revealed by molecular and morphological data. Molecular Ecology, 20(20): 4332-4345.
DOI:10.1111/j.1365-294x.2011.05275.x |
Ma H Y, Jiang W, Liu P, Feng N N, Ma Q Q, Ma C Y, Li S J, Liu Y X, Qiao Z G, Ma L B. 2014. Identification of transcriptome-derived microsatellite markers and their association with the growth performance of the mud crab (Scylla paramamosain). PLoS One, 9(2): e89134.
DOI:10.1371/journal.pone.0089134 |
Mariani S, Ketmaier V, de Matthaeis E. 2002. Genetic structuring and gene flow in Cerastoderma glaucum (Bivalvia: Cardiidae): evidence from allozyme variation at different geographic scales. Marine Biology, 140(4): 687-697.
DOI:10.1007/s00227-001-0753-x |
Murat T, Aykut K. 2015. Genetic and morphologic diversity of Pseudophoxinus (Cyprinidae): implication forconservation in Anatolia. Environmental Biology of Fishes, 98(2): 571-583.
DOI:10.1007/s10641-014-0291-2 |
Nei M, Maruyama T, Chakraborty R. 1975. The bottleneck effect and genetic variability in populations. Evolution, 29(1): 1-10.
DOI:10.1111/j.1558-5646.1975.tb00807.x |
Ni G, Li Q, Kong L F, Yu H. 2014. Comparative phylogeography in marginal seas of the northwestern Pacific. Molecular Ecology, 23(3): 534-548.
DOI:10.1111/mec.12620 |
Pan Y, Qin X M, Pan H P. 2007. Analysis and evaluation of nutritive composition in edible part of Lutraria maxima. Journal of Guangdong Ocean University, 27(3): 78-81.
(in Chinese with English abstract) |
Pan Y, Su Y P. 2007. The karyotype of Lutraria maxima Jonas. Marine Sciences, 31(9): 87-90.
|
Peakall R, Smouse P E. 2012. GenAlEx 6.5:genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics, 28(19): 2537-2539.
DOI:10.1093/bioinformatics/bts460 |
Pemberton J M, Slate J, Bancroft D R, Barrett J A. 1995. Nonamplifying alleles at microsatellite loci: a caution for parentage and population studies. Molecular Ecology, 4(2): 249-252.
DOI:10.1111/j.1365-294x.1995.tb00214.x |
Pérez-Losada M, Guerra A, Carvalho G R, Sanjuan A, Shaw P W. 2002. Extensive population subdivision of the cuttlefish Sepia officinalis (Mollusca: Cephalopoda) around the Iberian Peninsula indicated by microsatellite DNA variation. Heredity, 89(6): 417-424.
DOI:10.1038/sj.hdy.6800160 |
Qin Y, Shi G, Sun Y. 2013. Evaluation of genetic diversity in Pampus argenteus using SSR markers. Genetics and Molecular Research, 12(4): 5833-5841.
DOI:10.4238/2013.November.22.10 |
Ríos C, Sanz S, Saavedra C, Peña J B. 2002. Allozyme variation in populations of scallops, Pecten jacobaeus (L.) and P. maximus (L.) (Bivalvia: Pectinidae), across the Almeria-Oran front. Journal of Experimental Marine Biology and Ecology, 267(2): 223-244.
DOI:10.1016/s0022-0981(01)00371-9 |
Rosenberg N A. 2004. DISTRUCT: a program for the graphical display of population structure. Molecular Ecology Notes, 4(1): 137-138.
DOI:10.1046/j.1471-8286.2003.00566.x |
Ruiz-Campos G, Camarena-Rosales F, Varela-Romero A, Sánchez-González S, De La Rosa-Vélez J. 2003. Morphometric variation of wild trout populations from northwestern Mexico (Pisces: Salmonidae). Reviews in Fish Biology and Fisheries, 13(1): 91-110.
DOI:10.1023/a:1026367113735 |
Sato M, Kawamata K, Zaslavskaya N, Nakamura A, Ohta T, Nishikiori T, Brykov V, Nagashima K. 2005. Development of microsatellite markers for Japanese scallop (Mizuhopecten yessoensis) and their application to a population genetic study. Marine Biotechnology, 7(6): 713-728.
DOI:10.1007/s10126-004-0127-8 |
Selkoe K A, Toonen R J. 2006. Microsatellites for ecologists: a practical guide to using and evaluating microsatellite markers. Ecology Letters, 9(5): 615-629.
DOI:10.1111/j.1461-0248.2006.00889.x |
Serapion J, Kucuktas H, Feng J N, Liu Z J. 2004. Bioinformatic mining of type i microsatellites from expressed sequence tags of channel catfish (Ictalurus punctatus). Marine Biotechnology, 6(4): 364-377.
DOI:10.1007/s10126-003-0039-z |
Silva A. 2003. Morphometric variation among sardine (Sardina pilchardus) populations from the northeastern Atlanticand the western Mediterranean. ICES Journal of Marine Science, 60(6): 1352-1360.
DOI:10.1016/s1054-3139(03)00141-3 |
Smith C, Benzie J A H, Wilson K J. 2003. Isolation and characterization of eight microsatellite loci from silver-lipped pearl oyster Pinctada maxima. Molecular Ecology Notes, 3(1): 125-127.
DOI:10.1046/j.1471-8286.2003.00377.x |
So N, Maes G E, Volckaert F A M. 2006. High genetic diversity in cryptic populations of the migratory sutchi catfish Pangasianodon hypophthalmus in the Mekong River. Heredity, 96(2): 166-174.
DOI:10.1038/sj.hdy.6800781 |
Su Q, Tong W P, Li Q Z, Yang J L, Chen R F, Jiang Y, Cai D J. 2009. The technologe for factory breeding of Lutraria (Psammophila) maxima Jonas. Guangxi Sciences, 16(3): 342-345.
(in Chinese with English abstract) |
Tautz D, Renz M. 1984. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Research, 12(10): 4127-4138.
DOI:10.1093/nar/12.10.4127 |
Ueno S, Wen Y F, Tsumura Y. 2015. Development of EST-SSR markers for Taxus cuspidata from publicly available transcriptome sequences. Biochemical Systematics and Ecology, 63: 20-26.
DOI:10.1016/j.bse.2015.09.016 |
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth B C, Remm M, Rozen S G. 2012. Primer3-new capabilities and interfaces. Nucleic Acids Research, 40(15): e115.
DOI:10.1093/nar/gks596 |
Van Oosterhout C, Hutchinson W F, Wills D P M, Shipley P. 2004. Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes, 4(3): 535-538.
DOI:10.1111/j.1471-8286.2004.00684.x |
Voss R S, Marcus L F, Escalante P P. 1990. Morphological evolution in Muroid rodents I. Conservative patterns of craniometric covariance and their ontogenetic basis in the neotropical genus Zygodontomys. Evolution, 44(6): 1568-1587.
DOI:10.1111/j.1558-5646.1990.tb03847.x |
Wang C Y, Yu X M, Tong J G. 2007. Microsatellite diversity and population genetic structure of redfin culter (Culter erythropterus) in fragmented lakes of the Yangtze River. Hydrobiologia, 586(1): 321-329.
DOI:10.1007/s10750-007-0702-x |
Wanna W, Rolland J L, Bonhomme F, Phongdara A. 2004. Population genetic structure of Penaeus merguiensis in Thailand based on nuclear DNA variation. Journal of Experimental Marine Biology and Ecology, 311(1): 63-78.
DOI:10.1016/j.jembe.2004.04.018 |
Wolfus G M, Garcia D K, Alcivar-Warren A. 1997. Application of the microsatellite technique for analyzing genetic diversity in shrimp breeding programs. Aquaculture, 152(1-4): 35-47.
DOI:10.1016/s0044-8486(96)01527-x |
Wright S. 1965. The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution, 19(3): 395-420.
DOI:10.1111/j.1558-5646.1965.tb01731.x |
Yang Q, Zhao F, Song C, Zhang T, Miao Z B, Zhuang P. 2019. Analysis of morphological variations among four different geographic populations of Coilia mystus in the Yangtze River Estuary and its adjacent waters. Marine Fisheries, 41(3): 294-303.
(in Chinese with English abstract) |
Yue G H, Ho M Y, Orban L, Komen J. 2004. Microsatellites within genes and ESTs of common carp and their applicability in silver crucian carp. Aquaculture, 234(1-4): 85-98.
DOI:10.1016/j.aquaculture.2003.12.021 |
Zhan A B, Bao Z M, Wang X L, Hu J J. 2005. Microsatellite markers derived from bay scallop Argopecten irradians expressed sequence tags. Fisheries Science, 71(6): 1341-1346.
DOI:10.1111/j.1444-2906.2005.01100.x |
Zhan A B, Hu J J, H X L, Zhou Z C, Hui M, Wang S, Peng W, Wang M L, Bao Z M. 2009. Fine-scale population genetic structure of Zhikong scallop (Chlamys farreri): do local marine currents drive geographical differentiation?. Marine Biotechnology, 11(2): 223-235.
DOI:10.1007/s10126-008-9138-1 |
Zhang L W, Li A Q, Wang X F, Xu J T, Zhang G Q, Su J G, Qi J M, Guan C Y. 2013. Genetic diversity of Kenaf (Hibiscus cannabinus) evaluated by Inter-Simple Sequence Repeat(ISSR). Biochemical Genetics, 51(9-10): 800-810.
DOI:10.1007/s10528-013-9608-7 |
Zhao Y Y, Zhu X C, Li Z, Xu W B, Dong J, Wei H, Li Y D, Li X D. 2019. Genetic diversity and structure of Chinese grass shrimp, Palaemonetes sinensis, inferred from transcriptome-derived microsatellite markers. BMC Genetics, 20: 75.
DOI:10.1186/s12863-019-0779-z |
 2021, Vol. 39
2021, Vol. 39