Institute of Oceanology, Chinese Academy of Sciences
Article Information
- MA Yuexin, DU Xin, LIU Yubin, ZHANG Tao, WANG Yue, ZHANG Saisai
- Characterization of the bacterial communities associated with biofilters in two full-scale recirculating aquaculture systems
- Journal of Oceanology and Limnology, 39(3): 1143-1150
- http://dx.doi.org/10.1007/s00343-020-0120-8
Article History
- Received Mar. 10, 2020
- accepted in principle Apr. 23, 2020
- accepted for publication Jun. 15, 2020
2 Dalian Modern Agricultural Production Development Service Center, Dalian 116023, China;
3 Dalian Tianzheng Industry Company Limited, Dalian 116011, China
Recirculating aquaculture systems (RASs) have been widely applied in fish production (Sakami et al., 2012; Chen et al., 2013; Ruan et al., 2015; Huang et al., 2016; Rud et al., 2017) because RASs can minimize water requirements, discharge less of better treatable wastes and improve biosecurity (Itoi et al., 2006; Martins et al., 2010; Schreier et al., 2010; Rurangwa and Verdegem, 2015; Keuter et al., 2017). Moreover, RASs provide fish with optimal water quality through recycling thus offering the possibility to optimize feed utilization and to achieve high productions of healthy fish in controlled conditions (Rurangwa and Verdegem, 2015). In general, biofilter is an indispensable unit for RASs to maintain their water quality, which provides a fixed medium for microbial attachment and growth. Microorganisms within a biofilm formed on the surface of media play a major role in the operation of RASs. The high load of organic matter that resulted from uneaten feeds, dead bodies, and excreta of fish is converted by a diverse array of microorganisms. Nitrogen-containing organic compounds are mineralized into ammonia by heterotrophic ammonifying bacteria, with ammonia subsequently being oxidized into nitrate by autotrophic nitrifying bacteria in a biofilter. Heterotrophic bacteria within a biofilm are superior competitors for oxygen and space with autotrophic bacteria (Léonard et al., 2002; Itoi et al., 2006; Michaud et al., 2006), which are traditionally considered to hinder the effective operation of the system. Nevertheless, a moderate outer layer of heterotrophic bacteria in a biofilm may have a positive effect on nitrification by protecting nitrifying bacteria against flow detachment and grazing (Blancheton et al., 2013). Thus, it is essential to have a comprehensive understanding on the overall bacterial community structure in RASs for system management.
The advantage for puffer fish, Takifugu rubripes, cultured in a commercial RAS is that they can survive and maintain winter growth. Itoi et al. (2006) characterized the bacterial community in a laboratory scale RAS for puffer fish by clone library method. Only heterotrophic bacterial community associated with multistage biofiters was investigated in a fullscale RAS for puffer fish (Chen et al., 2013). Highthroughput sequencing is rapid and cost effective technology that allows for in-depth taxonomic characterization of bacterial community including uncultured bacteria (Metzker, 2010; Caporaso et al., 2012). In this study, two commercial RASs containing the same components were used for the culture of puffer fish with different sizes and densities. In addition, both systems had been operated for one and two years, respectively, before the experiment. The goal of this study was to characterize and compare bacterial communities in the biofilters of this two commercial RASs using Illumina Miseq sequencing.
2 MATERIAL AND METHOD 2.1 Description of experimental recirculating aquaculture systemTwo full-scale RASs (systems A and B) used in this study were operated at Dalian Tianzheng Industry Company Limited (Dalian, China). Before the experiment, systems A and B have been run for one and two years, respectively. Each RAS contained nine fish-rearing tanks with a water volume in each tank of approximately 50 m3, in which the puffer fish (A꞉ oneyear-old, mean size 210 g, 2 000 individuals per tank; B꞉ two-years-old, mean size 720 g, 1 000 individuals per tank) were reared and fed a natural diet (fresh fish, mainly Ammodytes personatus) twice a day (7꞉00 and 17꞉00) at a rate of 2%–3% body weight. Water temperature ranged from 20 ℃ to 21 ℃ and salinity from 24 to 25. The effluent from the tanks passed through a bowed screen and an air-floating tank to remove suspended and fine solids. The water circulation rate was 300 tons per hour. A submerged aerated biofilter (13.0 m in length, 3.3 m in width, and 2.0 m in depth) containing polyethylene elastic media treated metabolic wastes, which aerated by air flotation machine. An ultraviolet light chamber unit inactivated heterotrophic and coliform bacteria. Pure oxygen was injected into the contact unit using micropore diffusers, and treated water flowed back to the fish-rearing tanks by gravity (Lin et al., 2017).
2.2 Sample collectionDuring the production of puffer fish, the biofilm samples were collected at days of 165 (A2) and 183 (A3), and 157 (B2) and 175 (B3) of the systems A and B running, respectively. There is an 8-day interval between the two systems start running and the sampling dates are the same. Three sampling sites were set every 30 curtain media, and 15 sampling sites in total. Three repetitions (each contained a mixture from five sampling sites) were pooled equally as one biofilm sample. The polyethylene elastic media samples were collected in 50-mL falcon tubes, stored in a portable cooler, transferred to the laboratory, and processed immediately. The media were gently rinsed and suspended in sterile seawater, then shaken vigorously with a vortex mixer for 20 min. Biofilm samples were obtained by the liquid phase centrifugation at 6 000 r/min for 15 min.
2.3 DNA extraction and PCR amplificationDNA was extracted from four biofilm samples using a soil DNA isolation kit (Tiangen Biochemical Technology Limited, Beijing, China) according to the manufacturer's protocols. The 16S rRNA gene V4–V5 region was amplified by polymerase chain reaction (PCR) with primers 515F (5'-GTGCCAGCMGCCGCGG-3') and 907R (5'-CCGTCAATTCMTTTRAGTTT-3') (Turner et al., 1999) using the PCR program described by Ma et al. (2019) except for annealing at 54 ℃. PCR reactions were performed in triplicate and the reaction volume was 20 μL containing 5 μL of 10× Extaq Buffer, 1.6 μL of 2.5 mmol/L dNTPs (each nucleotide), 0.8 μL of 5 μmol/L primer (each direction), 0.5 U of Extaq Polymerase, 0.3 μL of bovine serum albumin and 10 ng of template DNA.
2.4 Illumina MiSeq sequencingPurified amplicons were obtained, pooled, and paired-end sequenced (2×300) on an Illumina MiSeq platform following standard protocols (Sun et al., 2016) (Majorbio Bio-Pharm Technology Company Limited, Shanghai, China).
2.5 Bioinformatics analysisSequencing data were processed according to methods and protocols described by Sun et al. (2016) and Ma et al. (2019).
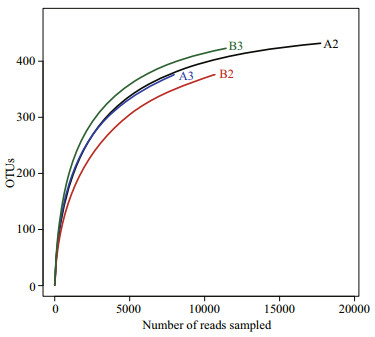
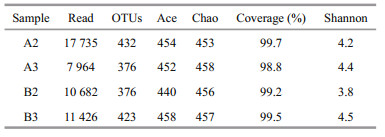
3 RESULT 3.1 Alpha diversity of the bacterial communityA total of 47 807 optimized reads with an average length of 394 bp were recovered from the four samples and clustered into 500 operational taxonomic units (OTUs). The numbers of OTUs in the A2, A3, B2, and B3 samples were 432, 376, 376, and 423, respectively. Rarefaction curves generated at the OTU level approached saturation (Fig. 1), indicating that the sequencing depth was sufficient to reflect the diversity of the bacterial community. Good's coverage estimations revealed that 98.8%−99.7% of the species were retrieved. The richness and diversity of the bacterial community were similar among four samples on the basis of the Chao, Ace, and Shannon indices (Table 1).
|
| Fig.1 Rarefaction curves of operational taxonomic units clustered at 97% similarity among biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) |
|
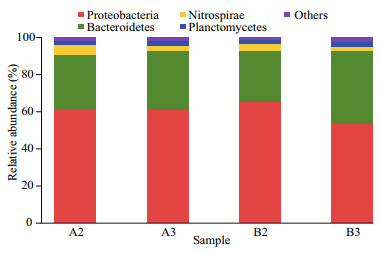
The 47 807 optimized reads were classified as 19 phyla, with Proteobacteria and Bacteroidetes making up >90% of the reads among four samples. The most abundant 16S rRNA gene sequences belonged to Proteobacteria, followed by those to Bacteroidetes (Fig. 2). Relative abundances for the16S rRNA gene sequences assigned to Nitrospirae and Planctomycetes were 2.4%–5.5% and 1.7%–2.8%, respectively (Fig. 2). Percentages of other phyla in four samples were less than 1% (Fig. 2).
|
| Fig.2 Taxonomic classification of bacterial reads retrieved from DNA amplicons from biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) into phylum level using the RDP Classifier Sequence abundance less than 1% at each sample was combined into 'others'. |
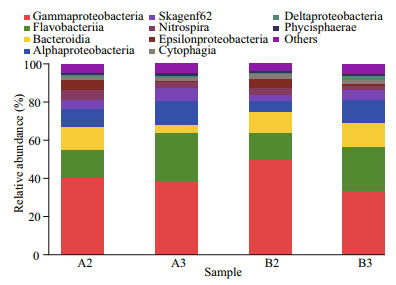
Thirty-seven classes were identified among four samples, and reads from Gammaproteobacteria and Flavobacteriia were abundant (Fig. 3). The relative abundances for sequences fell within the Bacteroidia in A2, B2, and B3 samples were 12%, 11%, and 12%, respectively, while in A3 sample 4.3% (Fig. 3). The percentages of sequences assigned to Alphaproteobacteria in A3 (12%) and B3 (12%) samples were higher than those in A2 (9.0%) and B2 (6.1%) samples (Fig. 3). The relative abundances of Nitrospira in A2, A3, B2, and B3 samples were 5.5%, 2.7%, 3.7%, and 2.4%, respectively (Fig. 3). The percentages of reads from Epsilonproteobacteria were higher in A2 and B2 samples (4.9% and 5.2%) than the other two samples (< 1%) (Fig. 3).
|
| Fig.3 Taxonomic classification of bacterial reads retrieved from DNA amplicons from biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) into class level using the RDP Classifier Sequence abundance less than 1% at each sample was combined into 'others'. |
One hundred and thirty-eight genera were identified in total and the sequences belonging to Colwellia were the most abundant among four samples (Fig. 4). Reads from Marinifilum were second most abundant in A2, B2, and B3 samples, followed by those belonging to Nitrospira (5.5%) and Arcobacter (4.9%); Arcobacter (5.2%) and Nitrospira (3.7%); and Lutibacter (5.1%) and Winogradskyella (3.3%) in A2, B2 and B3 samples, respectively (Fig. 4). However, sequences assigned to Oceanospirillum (6.7%) were the second most abundant in the A3 sample, followed by those belonging to Lutibacter (6.5%), Winogradskyella (5.7%), Pseudoalteromonas (5.3%), and Phaeobacter (3.6%) (Fig. 4). In addition, reads from Vibrio (0.54%, 0.73%, 0.52%, and 0.14%) and Coxiella (0.079%, 0.038%, 0, and 0.18%) were observed among A2, A3, B2, and B3 samples.

|
| Fig.4 Taxonomic classification of bacterial reads retrieved from DNA amplicons from biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) into genus level using the RDP Classifier Sequence abundance less than 1% at each sample was combined into 'others'. |
Regarding ammonia- and nitrite-oxidizing bacteria in A2, A3, B2, and B3 samples (Fig. 5), OTU 415 was identified as Nitrosomonas and OTU 113 was related to Nitrosococcus (Fig. 5), three OTUs (numbers 132, 145, and 239) were classified as Nitrospira, of which OTU 145 belonged to Candidatus Nitrospira salsa (Fig. 5).

|
| Fig.5 Relative abundances of ammonia- and nitriteoxidizing bacteria among biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) |
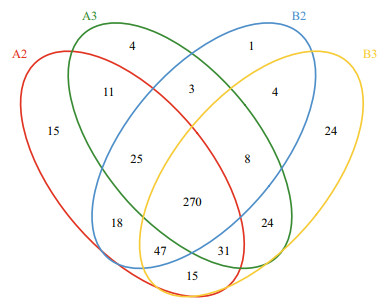
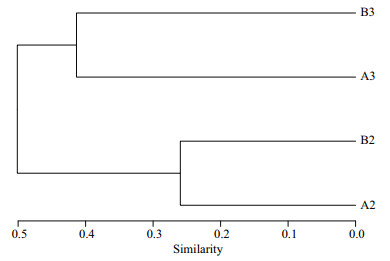
The Venn diagram (Fig. 6) indicated that 54% of OTUs and 70%–71% of OTUs were shared by four and two (A2 and A3, B2 and B3) biofilm samples. Each sample contained only a minority of unique OTUs (1–24). Variability among four samples was captured by cluster analysis (Fig. 7). The results from a cluster analysis of the OTUs profile show that the A2 and B2 samples formed one cluster and the A3 and B3 samples formed another.
|
| Fig.6 Venn diagram showing operational taxonomic units unique to and shared by biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) |
|
| Fig.7 Cluster analysis of operational taxonomic units profile for biofilm samples of biofiter 1 (days 165 (A2) and 183 (A3)) and biofiter 2 (days 157 (B2) and 175 (B3)) according to Bray-Curtis distance |
In the present study, a total of 19 different phyla were identified, while the culturable bacterial community in biofilms from biofilters in another commercial puffer fish RAS were only classified into four phyla (Proteobacteria, Bacteroidetes, Firmicutes, and Actinobacteria) (Chen et al., 2013). Among the total of reads detected, Proteobacteria and Bacteroidetes were dominant phyla, which are in line with previous studies (Michaud et al., 2009; Ruan et al., 2015; Huang et al., 2016). Both phyla play an important role in degradation of organic compounds in aquatic environments (Kirchman, 2002; Kersters et al., 2006). The most abundant class in Proteobacteria was Gammaproteobacteria, followed by Alphaproteobacteria. The dominant class in Bacteroidetes was Flavobacteria. These findings are similar to previous results from a full-scale marine RAS biofilter (Ruan et al., 2015). The relative abundance of phylum Nitrospirae (class Nitrospira) (2.4%–5.5%) was in the range of other marine RAS biofilters (Ruan et al., 2015; Huang et al., 2016). Sequences related to the Planctomycetales were detected in four samples. This could suggest the presence of anammox through which nitrogen was eliminated by combining ammonia and nitrite to form nitrogen gas (van de Graaf et al., 1995). Tal et al. (2006) first confirmed the presence of anammox Planctomycetales in the microbial biofilm from the denitrifying biofilters of a RAS for the gilthead seabream, Sparus aurata. Moreover, Planctomycetes is frequently observed in biofilters of laboratory scale and full-scale marine RASs (Itoi et al., 2006; Michaud et al., 2009; Ruan et al., 2015; Huang et al., 2016).
At the genus level, Colwellia, Marinifilum, Oceanospirillum, Lutibacter, Winogradskyella Nitrospira, Pseudoalteromonas, Arcobacter, and Phaeobacter were the top nine genera when data from four samples were considered together. They are different from those of other puffer fish RASs (Itoi et al., 2006; Chen et al., 2013). Using clone library method, bacteria in the aquarium (pebbles as medium) of puffer fish were composed predominantly by genera belonging to Algibacter, Flavobacterium, Muricauda, Psychroserpens, Lewinella, Caminicella, Lactococcus, and Marinobacter (Itoi et al., 2006). Using culturedependent technique, the abundant genera were Vibrio, Shewanella and Pseudoalteromonas in biofilters (plastic brush packing medium) in a commercial puffer fish RAS (Chen et al., 2013). The differences in bacterial genera between puffer fish RASs may relate to fish density, biofilter construction, media, time of biofilm formation except for various methods of bacterial community analysis. Colwellia, Arcobacter and Lutibacter have been identified as abundant genera in different sections of a biofilter in a RAS for full-scale tongue sole, Cynoglossus semilaevis (Ruan et al., 2015). The genus Oceanospirillum was the second most abundant in A3 sample, members of this genus are known to be distributed ubiquitously in marine environments (Satomi et al., 2002). This taxon can utilize amino acids or the salts of organic acids as carbon sources (Pot and Gillis, 2015). In addition, Pseudoalteromonas and Shewanella were also isolated from the intestine of puffer fish (Li et al., 2015), which might be the source of bacteria for the biofilters.
In line with previous studies (Tal et al., 2003; Foesel et al., 2008; Ruan et al., 2015; Huang et al., 2016), it was demonstrated that representatives of the genus Nitrospira were identified as dominant nitrite-oxidizing bacteria (NOB) in biofilters of marine RASs. The relative abundance of genus Nitrospira across the four samples ranged from 2.4% to 5.5%, which included a marine nitrite-oxidizing Nitrospira species Candidatus Nitrospira salsa (Haaijer et al., 2013). In addition, the sequence number of the NOB was roughly an order of magnitude higher than that of ammonia-oxidizing bacteria, which included genera Nitrosomonas and Nitrosococcus, suggesting that the NOB may not be the limiting factor in nitrification performance. These findings indicate that Nitrosomonas and Nitrospira play an important role in removing ammonia and nitrite from puffer fish RASs. Similar results were observed in biofilters of other marine RASs (Ruan et al., 2015; Huang et al., 2016).
Several potential pathogens, such as Coxiella sp. and Vibrio sp., were detected in this study. Vibrio spp. have been isolated from biofilms of biofilters in another commercial pufferfish RAS (Chen et al., 2013). Coxiella sp. and Vibrio sp. have been observed associated with the biofilters packing medium in a RAS for sea bass, Dicentrarchus labrax (Michaud et al., 2009). Members of both genera have also been found in biofilters among several marine RASs (Huang et al., 2016). Despite the presence of potential pathogens, no diseased fish were observed during the period of this study. One of reasons might be that the relative abundances of the potential bacterial pathogens in biofilters of puffer fish RASs were low (Coxiella, 0–0.18%; Vibrio, 0.14%–0.73%).
The results obtained of this study suggest that both biofilters contain a large bacterial diversity, with similarities in the bacterial communities being observed between the two systems as shown in the cluster analysis. Although the puffer fish sizes and densities were different in both systems, the daily feeding dose for two-years-old puffer fish was slightly less than that for one-year-old, thus the loads of organic metabolic wastes and ammonia on both systems at the same sampling date were not much different. However, there were different bacterial communities at the two sampling dates, this might be explained by the bacterial decomposition of substrate, making the nutrient substances of bacteria different at different sampling dates.
5 CONCLUSIONProteobacteria and Bacteroidetes were the most abundant phyla in both biofilters that involved in the degradation of organic compounds. Nitrosomonas and Nitrospira were main ammonia- and nitriteoxidizing bacteria being responsible for ammonia and nitrite oxidation. Both biofilters harbored similar bacterial communities and dissimilar at the two different sampling dates. These results are useful for the management and stable operation of the puffer fish RASs.
6 DATA AVAILABILITY STATEMENTAll data generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Blancheton J P, Attramadal K L K, Michaud L, d'Orbcastel E R, Vadstein O. 2013. Insight into bacterial population in aquaculture systems and its implication. Aquacultural Engineering, 53: 30-39.
DOI:10.1016/j.aquaeng.2012.11.009 |
Caporaso J G, Lauber C L, Walters W A, Berg-Lyons D, Huntley J, Fierer N, Owens S M, Betley J, Fraser L, Bauer M, Gormley N, Gilbert J A, Smith G, Knight R. 2012. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. The ISME Journal, 6(8): 1 621-1 624.
DOI:10.1038/ismej.2012.8 |
Chen Z, Liu Y, Liu L Z, Wang X J, Liu Z P, Liu Y. 2013. Heterotrophic bacterial community structure of multistage biofilters in a commercial pufferfish Takifugu rubripes RAS. Advanced Materials Research, 726-731: 1 621-1 627.
DOI:10.4028/www.scientific.net/AMR.726-731.1621 |
Foesel B U, Gieseke A, Schwermer C, Stief P, Koch L, Cytryn E, De La Torré J R, Van Rijn J, Minz D, Drake H L, Schramm A. 2008. Nitrosomonas Nm143-like ammonia oxidizers and Nitrospira marina-like nitrite oxidizers dominate the nitrifier community in a marine aquaculture biofilm. FEMS Microbiology Ecology, 63(2): 192-204.
DOI:10.1111/j.1574-6941.2007.00418.x |
Haaijer S C M, Ji K, Van Niftrik L, Hoischen A, Speth D, Jetten M S M, Damsté J S S, Op den Camp H J M. 2013. A novel marine nitrite-oxidizing Nitrospira species from Dutch coastal North Sea water. Frontiers in Microbiology, 4: 60.
DOI:10.3389/fmicb.2013.00060 |
Huang Z T, Wan R, Song X F, Liu Y, Hallerman E, Dong D P, Zhai J M, Zhang H S, Sun L Y. 2016. Metagenomic analysis shows diverse, distinct bacterial communities in biofilters among different marine recirculating aquaculture systems. Aquaculture International, 24(5): 1 393-1 408.
DOI:10.1007/s10499-016-9997-9 |
Itoi S, Niki A, Sugita H. 2006. Changes in microbial communities associated with the conditioning of filter material in recirculating aquaculture systems of the pufferfish Takifugu rubripes. Aquaculture, 256(1-4): 287-295.
DOI:10.1016/j.aquaculture.2006.02.037 |
Kersters K, De Vos P, Gillis M, Swings J, Vandamme P, Stackebrandt E. 2006. Introduction to the Proteobacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K H, Stackebrandt E eds. The Prokaryotes: A Handbook on the Biology of Bacteria. Springer, New York. p. 3-37. Chapter
|
Keuter S, Beth S, Quantz G, Schulz C, Spieck E. 2017. Longterm monitoring of nitrification and nitrifying communities during biofilter activation of two marine recirculation aquaculture systems (RAS). International Journal of Aquaculture and Fishery Sciences, 3(3): 51-61.
DOI:10.17352/2455-8400.000029 |
Kirchman D L. 2002. The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiology Ecology, 39(2): 91-100.
DOI:10.1111/j.1574-6941.2002.tb00910.x |
Léonard N, Guiraud J P, Gasset E, Cailleres J P, Blancheton J P. 2002. Bacteria and nutrients-nitrogen and carbon-in a recirculating system for sea bass production. Aquacultural Engineering, 26(2): 111-127.
|
Li Y Y, Zhang T, Zhang C Y, Zhu Y, Ding J F, Ma Y X. 2015. Bacterial diversity in the intestine of young farmed puffer fish Takifugu rubripes. Chinese Journal of Oceanology and Limnology, 33(4): 913-918.
DOI:10.1007/s00343-015-4219-2 |
Lin Z L, Wang H, Yu C Y, Lv F H, Liu H M, Zhang T. 2017. Commercial production of tiger puffer (Takifugu rubripes) in winter using a recirculating aquaculture system. Journal of Ocean University of China, 16(1): 107-113.
DOI:10.1007/s11802-017-2802-1 |
Ma Y X, Li M, Sun J X, Hao Z L, Liang J, Zhao X W. 2019. Characterization of bacterial community associated with four organs of the Yesso scallop (Patinopecten yessoensis) by high-throughput sequencing. Journal of Ocean University of China, 18(2): 493-500.
DOI:10.1007/s11802-019-3791-z |
Martins C I M, Eding E H, Verdegem M C J, Heinsbroek L T N, Schneider O, Blancheton J P, d'Orbcastel E R, Verreth J A J. 2010. New developments in recirculating aquaculture systems in Europe: a perspective on environmental sustainability. Aquacultural Engineering, 43(3): 83-93.
DOI:10.1016/j.aquaeng.2010.09.002 |
Metzker M L. 2010. Sequencing technologies-the next generation. Nature Reviews Genetics, 11(1): 31-46.
DOI:10.1038/nrg2626 |
Michaud L, Blancheton J P, Bruni V, Piedrahita R. 2006. Effect of particulate organic carbon on heterotrophic bacterial populations and nitrification efficiency in biological filters. Aquacultural Engineering, 34(3): 224-233.
DOI:10.1016/j.aquaeng.2005.07.005 |
Michaud L, Lo Giudice A, Troussellier M, Smedile F, Bruni V, Blancheton J P. 2009. Phylogenetic characterization of the heterotrophic bacterial communities inhabiting a marine recirculating aquaculture system. Journal of Applied Microbiology, 107(6): 1 935-1 946.
DOI:10.1111/j.1365-2672.2009.04378.x |
Pot B, Gillis M. 2015. Oceanospirillum. John Wiley and Sons, Inc., New York.
DOI:10.1002/9781118960608.gbm01196
|
Ruan Y J, Guo X S, Ye Z Y, Liu Y, Zhu S M. 2015. Bacterial community analysis of different sections of a biofilter in a full-scale marine recirculating aquaculture system. North American Journal of Aquaculture, 77(3): 318-326.
DOI:10.1080/15222055.2015.1017128 |
Rud I, Kolarevic J, Holan A B, Berget I, Calabrese S, Terjesen B F. 2017. Deep-sequencing of the bacterial microbiota in commercial-scale recirculating and semi-closed aquaculture systems for Atlantic salmon post-smolt production. Aquacultural Engineering, 78: 50-62.
DOI:10.1016/j.aquaeng.2016.10.003 |
Rurangwa E, Verdegem M C J. 2015. Microorganisms in recirculating aquaculture systems and their management. Reviews in Aquaculture, 7(2): 117-130.
DOI:10.1111/raq.12057 |
Sakami T, Andoh T, Morita T, Yamamoto Y. 2012. Phylogenetic diversity of ammonia-oxidizing archaea and bacteria in biofilters of recirculating aquaculture systems. Marine Genomics, 7(9): 27-31.
DOI:10.1016/j.margen.2012.04.006 |
Satomi M, Kimura B, Hamada T, Harayama S, Fujii T. 2002. Phylogenetic study of the genus Oceanospirillum based on 16S rRNA and gyrB genes: emended description of the genus Oceanospirillum, description of Pseudospirillum gen. nov., Oceanobacter gen. nov. and Terasakiella gen. nov. and transfer of Oceanospirillum jannaschii and Pseudomonas stanieri to Marinobacterium as Marinobacterium jannaschii comb. nov. and Marinobacterium stanieri comb. no. International Journal of Systematic and Evolutionary Microbiology, 52(3): 739-747.
DOI:10.1099/ijs.0.01427-0 |
Schreier H J, Mirzoyan N, Saito K. 2010. Microbial diversity of biological filters in recirculating aquaculture systems. Current Opinion in Biotechnology, 21(3): 318-325.
DOI:10.1016/jxopbio.2010.03.011 |
Sun X Y, Liu J C, Li M, Zhao X W, Liang J, Sun P H, Ma Y X. 2016. Characterization of bacterial communities associating with larval development of Yesso scallop (Patinopecten yessoensis Jay, 1857) by high-throughput sequencing. Journal of Ocean University of China, 15(6): 1 067-1 072.
DOI:10.1007/s11802-016-3092-8 |
Tal Y, Watts J E M, Schreier H J. 2006. Anaerobic ammonium-oxidizing (anammox) bacteria and associated activity in fixed-film biofilters of a marine recirculating aquaculture system. Applied and Environmental Microbiology, 72(4): 2 896-2 904.
DOI:10.1128/AEM.72.4.2896-2904.2006 |
Tal Y, Watts J E M, Schreier S B, Sowers K R, Schreier H J. 2003. Characterization of the microbial community and nitrogen transformation processes associated with moving bed bioreactors in a closed recirculated mariculture system. Aquaculture, 215(1-4): 187-202.
DOI:10.1016/S0044-8486(02)00372-1 |
Turner S N, Pryer K M, Miao V P W, Palmer J D. 1999. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. The Journal of Eukaryotic Microbiology, 46(4): 327-338.
DOI:10.1111/j.1550-7408.1999.tb04612.x |
van de Graaf AA, Mulder A, de Bruijn P, Jetten M S, Robertson L A, Kuenen J G. 1995. Anaerobic oxidation of ammonium is a biologically mediated process. Applied and Environmental Microbiology, 61(4): 1 246-1 251.
|
 2021, Vol. 39
2021, Vol. 39


